

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138133.3 - phase: 0 /pseudo
(430 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
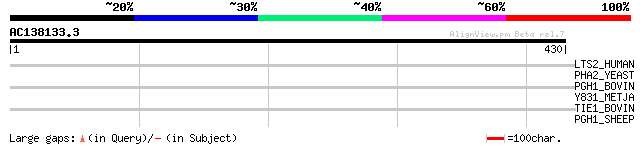
Score E
Sequences producing significant alignments: (bits) Value
LTS2_HUMAN (Q9NRM7) Serine/threonine protein kinase LATS2 (EC 2.... 33 1.4
PHA2_YEAST (P32452) Prephenate dehydratase (EC 4.2.1.51) (PDT) 31 7.0
PGH1_BOVIN (O62664) Prostaglandin G/H synthase 1 (EC 1.14.99.1) ... 31 7.0
Y831_METJA (Q58241) Hypothetical protein MJ0831 30 9.2
TIE1_BOVIN (Q06805) Tyrosine-protein kinase receptor Tie-1 precu... 30 9.2
PGH1_SHEEP (P05979) Prostaglandin G/H synthase 1 precursor (EC 1... 30 9.2
>LTS2_HUMAN (Q9NRM7) Serine/threonine protein kinase LATS2 (EC
2.7.1.37) (Large tumor suppressor homolog 2)
(Serine/threonine kinase kpm) (Kinase phosphorylated
during mitosis protein) (Warts-like kinase)
Length = 1088
Score = 33.1 bits (74), Expect = 1.4
Identities = 26/103 (25%), Positives = 45/103 (43%), Gaps = 17/103 (16%)
Query: 316 DTWRVADGYVLWYARVSHPQILPPIPRDLPRPANEEQIIAEQWQRY-----EARSSPDTY 370
D W V G +L+ V P L P P E Q+ W+ + + SP+
Sbjct: 895 DWWSV--GVILFEMLVGQPPFLAPTP-------TETQLKVINWENTLHIPAQVKLSPEAR 945
Query: 371 DMVSGVVAYADAQLGQEEVMSMTPQQWFEAM---THMREQIAP 410
D+++ + AD +LG+ + +F A+ + +R+Q AP
Sbjct: 946 DLITKLCCSADHRLGRNGADDLKAHPFFSAIDFSSDIRKQPAP 988
>PHA2_YEAST (P32452) Prephenate dehydratase (EC 4.2.1.51) (PDT)
Length = 368
Score = 30.8 bits (68), Expect = 7.0
Identities = 39/165 (23%), Positives = 67/165 (39%), Gaps = 35/165 (21%)
Query: 252 IMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDLPPPSIVQMF--------VDFGTHT 303
+ G + Y H + L+Q+ + +DV LP SI Q F +D+
Sbjct: 43 LFLGPKGTYSH--QAALQQF-------QSTSDVEYLPAASIPQCFNQLENDTSIDYSVVP 93
Query: 304 LKADARGEQAGEDTWRVADGYVLWYARVSHPQILPPIPRDLPRPANEEQIIAEQWQRYEA 363
L+ G+ V Y L R+ + P P D R + ++IAEQ+
Sbjct: 94 LENSTNGQ--------VVFSYDLLRDRMIKKALSLPAPADTNRITPDIEVIAEQY----- 140
Query: 364 RSSPDTYDMVSGV-VAYADAQLG--QEEVMSMTPQQWFEAMTHMR 405
P T+ ++S + + A LG +E ++ PQ W + ++R
Sbjct: 141 --VPITHCLISPIQLPNGIASLGNFEEVIIHSHPQVWGQVECYLR 183
>PGH1_BOVIN (O62664) Prostaglandin G/H synthase 1 (EC 1.14.99.1)
(Cyclooxygenase-1) (COX-1) (Prostaglandin-endoperoxide
synthase 1) (Prostaglandin H2 synthase 1) (PGH synthase
1) (PGHS-1) (PHS 1) (Fragment)
Length = 259
Score = 30.8 bits (68), Expect = 7.0
Identities = 15/34 (44%), Positives = 22/34 (64%), Gaps = 3/34 (8%)
Query: 317 TWRVADGYVLW--YARVSH-PQILPPIPRDLPRP 347
T+ VA Y+ W ++ VS+ +ILP +PRD P P
Sbjct: 9 TYNVAHDYISWESFSNVSYYTRILPSVPRDCPTP 42
>Y831_METJA (Q58241) Hypothetical protein MJ0831
Length = 432
Score = 30.4 bits (67), Expect = 9.2
Identities = 10/28 (35%), Positives = 18/28 (63%)
Query: 256 VRRVYRHLPERVLRQYGYVQTIPRHPTD 283
+ +V + + E++L YGY++ I H TD
Sbjct: 8 IEKVTKAIKEKILNHYGYIRVITHHDTD 35
>TIE1_BOVIN (Q06805) Tyrosine-protein kinase receptor Tie-1
precursor (EC 2.7.1.112)
Length = 1136
Score = 30.4 bits (67), Expect = 9.2
Identities = 20/50 (40%), Positives = 24/50 (48%)
Query: 329 ARVSHPQILPPIPRDLPRPANEEQIIAEQWQRYEARSSPDTYDMVSGVVA 378
ARV P PP PR L A + I WQR EA + P + +V VA
Sbjct: 633 ARVLLPPSGPPAPRHLHAQALSDSEIQLMWQRPEAAAGPISKYIVEVQVA 682
>PGH1_SHEEP (P05979) Prostaglandin G/H synthase 1 precursor (EC
1.14.99.1) (Cyclooxygenase -1) (COX-1)
(Prostaglandin-endoperoxide synthase 1) (Prostaglandin
H2 synthase 1) (PGH synthase 1) (PGHS-1) (PHS 1)
Length = 600
Score = 30.4 bits (67), Expect = 9.2
Identities = 14/34 (41%), Positives = 22/34 (64%), Gaps = 3/34 (8%)
Query: 317 TWRVADGYVLW--YARVSH-PQILPPIPRDLPRP 347
T+ +A Y+ W ++ VS+ +ILP +PRD P P
Sbjct: 129 TYNIAHDYISWESFSNVSYYTRILPSVPRDCPTP 162
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.140 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,830,491
Number of Sequences: 164201
Number of extensions: 2429447
Number of successful extensions: 6355
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 6352
Number of HSP's gapped (non-prelim): 7
length of query: 430
length of database: 59,974,054
effective HSP length: 113
effective length of query: 317
effective length of database: 41,419,341
effective search space: 13129931097
effective search space used: 13129931097
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC138133.3


