

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.19 - phase: 0
(881 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
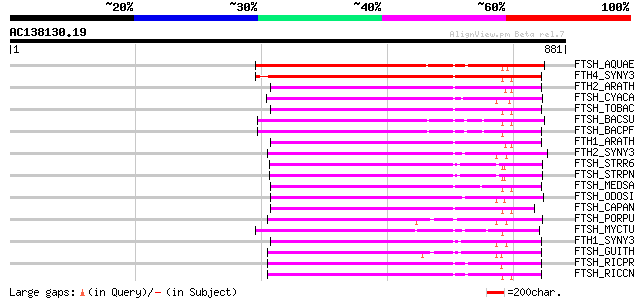
Score E
Sequences producing significant alignments: (bits) Value
FTSH_AQUAE (O67077) Cell division protein ftsH homolog (EC 3.4.2... 341 4e-93
FTH4_SYNY3 (P72991) Cell division protein ftsH homolog 4 (EC 3.4... 338 4e-92
FTH2_ARATH (Q9FH02) Cell division protein ftsH homolog 2, chloro... 313 1e-84
FTSH_CYACA (O19922) Cell division protein ftsH homolog (EC 3.4.2... 313 1e-84
FTSH_TOBAC (O82150) Cell division protein ftsH homolog, chloropl... 313 2e-84
FTSH_BACSU (P37476) Cell division protein ftsH homolog (EC 3.4.2... 313 2e-84
FTSH_BACPF (P94304) Cell division protein ftsH homolog (EC 3.4.2... 310 8e-84
FTH1_ARATH (Q39102) Cell division protein ftsH homolog 1, chloro... 309 2e-83
FTH2_SYNY3 (P73179) Cell division protein ftsH homolog 2 (EC 3.4... 308 3e-83
FTSH_STRR6 (P59652) Cell division protein ftsH homolog (EC 3.4.2... 307 9e-83
FTSH_STRPN (O69076) Cell division protein ftsH homolog (EC 3.4.2... 307 9e-83
FTSH_MEDSA (Q9BAE0) Cell division protein ftsH homolog, chloropl... 307 9e-83
FTSH_ODOSI (P49825) Cell division protein ftsH homolog (EC 3.4.2... 305 3e-82
FTSH_CAPAN (Q39444) Cell division protein ftsH homolog, chloropl... 305 3e-82
FTSH_PORPU (P51327) Cell division protein ftsH homolog (EC 3.4.2... 305 5e-82
FTSH_MYCTU (P96942) Cell division protein ftsH homolog (EC 3.4.2... 303 2e-81
FTH1_SYNY3 (Q55700) Cell division protein ftsH homolog 1 (EC 3.4... 302 2e-81
FTSH_GUITH (O78516) Cell division protein ftsH homolog (EC 3.4.2... 302 3e-81
FTSH_RICPR (Q9ZEA2) Cell division protein ftsH homolog (EC 3.4.2... 301 7e-81
FTSH_RICCN (Q92JJ9) Cell division protein ftsH homolog (EC 3.4.2... 299 2e-80
>FTSH_AQUAE (O67077) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 634
Score = 341 bits (875), Expect = 4e-93
Identities = 210/482 (43%), Positives = 303/482 (62%), Gaps = 31/482 (6%)
Query: 391 MKSGARVRRAQN---RRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRG 447
M G V RA N R Y+E V F DVAG+ +++ E++EI+++ +++ G
Sbjct: 125 MSGGGNVNRAFNFGKSRAKVYIEEKPKVTFKDVAGIEEVKEEVKEIIEYLKDPVKFQKLG 184
Query: 448 VKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAK 507
+ P G+LL G PGVGKTLLAKA+AGEA V F S+S S FVE++VGVGA+RVR L++ AK
Sbjct: 185 GRPPKGVLLYGEPGVGKTLLAKAIAGEAHVPFISVSGSDFVEMFVGVGAARVRDLFETAK 244
Query: 508 ENAPSVVFIDELDAVGRKRGLIK-GSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRP 566
++AP ++FIDE+DAVGR RG I G G ER+ TLNQLLV +DGF+ +I IA+TNRP
Sbjct: 245 KHAPCIIFIDEIDAVGRARGAIPVGGGHDEREQTLNQLLVEMDGFDTSDGIIVIAATNRP 304
Query: 567 DILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAE 626
DILDPAL+RPGRFDR+IFIPKP GR EILKVHAR K +A+DVD E VA T G GA+
Sbjct: 305 DILDPALLRPGRFDRQIFIPKPDVRGRYEILKVHARNKKLAKDVDLEFVARATPGFTGAD 364
Query: 627 LANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKERS---KEKWEQVAINEAAM 683
L N++ AA+ R + E++ +++ +A G L+RK + KEK E++AI+EA
Sbjct: 365 LENLLNEAALLAARKGKEEITMEEIEEALDRITMG-LERKGMTISPKEK-EKIAIHEAGH 422
Query: 684 AVAAMNLPNFDNIEYITIAPRAGRELGYVRTM-LESINFNDGMLTRQSLFDHITVQLAPR 742
A+ + + D + I+I PR G LG + + +E + D ++ L++ I V L R
Sbjct: 423 ALMGLVSDDDDKVHKISIIPR-GMALGVTQQLPIEDKHIYD----KKDLYNKILVLLGGR 477
Query: 743 AADEMWFGKDQLSTIWAETADNAR-VAARMYMIGGLSDK-----YRGVSNFWV------- 789
AA+E++FGKD ++T A +A RM + G+SDK R V+N ++
Sbjct: 478 AAEEVFFGKDGITTGAENDLQRATDLAYRMVSMWGMSDKVGPIAIRRVANPFLGGMTTAV 537
Query: 790 ---TDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQL 846
D + EID E +I+ YE+AK I+++ K + +V +L+ K+T+T E+ V + +L
Sbjct: 538 DTSPDLLREIDEEVKRIITEQYEKAKAIVEEYKEPLKAVVKKLLEKETITCEEFVEVFKL 597
Query: 847 HG 848
+G
Sbjct: 598 YG 599
>FTH4_SYNY3 (P72991) Cell division protein ftsH homolog 4 (EC
3.4.24.-)
Length = 616
Score = 338 bits (867), Expect = 4e-92
Identities = 201/475 (42%), Positives = 292/475 (61%), Gaps = 32/475 (6%)
Query: 390 FMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVK 449
F KS ARV+ +E V F DVAG+ + +LEL E+V F + + + G K
Sbjct: 143 FGKSKARVQ----------MEPQTQVTFGDVAGIEQAKLELTEVVDFLKNADRFTELGAK 192
Query: 450 IPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKEN 509
IP G+LL GPPG GKTLLAKAVAGEAGV FFSIS S+FVE++VGVGASRVR L+++AK N
Sbjct: 193 IPKGVLLVGPPGTGKTLLAKAVAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFEQAKAN 252
Query: 510 APSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDIL 569
AP +VFIDE+DAVGR+RG G G ER+ TLNQLL +DGFEG +I +A+TNRPD+L
Sbjct: 253 APCIVFIDEIDAVGRQRGAGLGGGNDEREQTLNQLLTEMDGFEGNTGIIIVAATNRPDVL 312
Query: 570 DPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELAN 629
D AL+RPGRFDR++ + +P + GR EIL VHAR K +++DVD + +A T G GA+L+N
Sbjct: 313 DSALMRPGRFDRQVVVDRPDYAGRREILNVHARGKTLSQDVDLDKIARRTPGFTGADLSN 372
Query: 630 IVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAA 687
++ AAI R + TE+S D++ A G ++K R S+++ VA +EA A+
Sbjct: 373 LLNEAAILAARRNLTEISMDEVNDAIDRVLAGP-EKKNRVMSEKRKTLVAYHEAGHALVG 431
Query: 688 MNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEM 747
+P++D ++ I+I PR GR G G+ +R L + + V L R A+E+
Sbjct: 432 ALMPDYDPVQKISIIPR-GRAGGLTWFTPSEDRMESGLYSRSYLQNQMAVALGGRIAEEI 490
Query: 748 WFGKDQLSTIWAETADN-ARVAARMYMIGGLSDKY-------RGVSNFWVTDRINE---- 795
FG+++++T + ARVA +M G+SD+ +G F D ++
Sbjct: 491 IFGEEEVTTGASNDLQQVARVARQMVTRFGMSDRLGPVALGRQGGGVFLGRDIASDRDFS 550
Query: 796 ------IDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
ID E ++++ Y+RAK++L +N+ ++D L LV K+T+ E++ L+
Sbjct: 551 DETAAAIDEEVSQLVDQAYQRAKQVLVENRGILDQLAEILVEKETVDSEELQTLL 605
>FTH2_ARATH (Q9FH02) Cell division protein ftsH homolog 2,
chloroplast precursor (EC 3.4.24.-)
Length = 704
Score = 313 bits (803), Expect = 1e-84
Identities = 188/449 (41%), Positives = 266/449 (58%), Gaps = 22/449 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG + +LEL+E+V F + + Y G KIP G LL GPPG GKTLLA+AVAGE
Sbjct: 247 VTFGDVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGE 306
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AGV FFS +AS+FVE++VGVGASRVR L+++AK AP +VFIDE+DAVGR+RG G G
Sbjct: 307 AGVPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRGAGMGGGN 366
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ T+NQLL +DGF G VI +A+TNRPD+LD AL+RPGRFDR++ + +P GR+
Sbjct: 367 DEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRV 426
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+ILKVH+R K I +DVDYE VA T G GA+L N++ AAI R E+S D++ A
Sbjct: 427 QILKVHSRGKAIGKDVDYEKVARRTPGFTGADLQNLMNEAAILAARRELKEISKDEISDA 486
Query: 655 AQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYV 712
+ G ++K S+EK VA +EA A+ +P +D + I+I PR G+ G
Sbjct: 487 LERIIAGP-EKKNAVVSEEKKRLVAYHEAGHALVGALMPEYDPVAKISIIPR-GQAGGLT 544
Query: 713 RTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNARVAARM 771
G+ +R L + + V L R A+E+ FG + ++T + +RVA +M
Sbjct: 545 FFAPSEERLESGLYSRSYLENQMAVALGGRVAEEVIFGDENVTTGASNDFMQVSRVARQM 604
Query: 772 YMIGGLSDKYRGVS------NFWVTDRINE-----------IDLEAMKILNLCYERAKEI 814
G S K V+ N ++ ++ +D E +++ Y RAKEI
Sbjct: 605 VERFGFSKKIGQVAVGGAGGNPFLGQSMSSQKDYSMATADVVDAEVRELVEKAYVRAKEI 664
Query: 815 LQQNKTLMDTLVNELVVKKTLTKEDIVRL 843
+ ++ L L+ K+T+ E+ + L
Sbjct: 665 ITTQIDILHKLAQLLIEKETVDGEEFMSL 693
>FTSH_CYACA (O19922) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 614
Score = 313 bits (802), Expect = 1e-84
Identities = 185/455 (40%), Positives = 268/455 (58%), Gaps = 21/455 (4%)
Query: 408 YLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLL 467
++E + F DVAG+ + + EL+EIV F + G IP G+LL GPPG GKTLL
Sbjct: 161 HMEAKTGIVFEDVAGIEEAKEELQEIVAFLKDSRKFTNVGATIPKGVLLVGPPGTGKTLL 220
Query: 468 AKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRG 527
AKA+AGEA FFSIS S+FVE++VGVGASRVR L+++AKE AP +VFIDE+DAVGR+RG
Sbjct: 221 AKAIAGEASAPFFSISGSEFVEMFVGVGASRVRDLFKKAKEKAPCIVFIDEIDAVGRQRG 280
Query: 528 LIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPK 587
+ G G ER+ TLNQLL +DGF G VI +A+TNR D+LD AL+RPGRFDR+I +
Sbjct: 281 VGIGGGNDEREQTLNQLLTEMDGFSGDTGVIVVAATNRIDVLDSALLRPGRFDRQIMVSL 340
Query: 588 PGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVS 647
P GR+ ILKVH++KK I +DV E++A T G GA+LAN++ AAI +R + E++
Sbjct: 341 PNINGRLAILKVHSKKKKIHKDVLLEVIARRTPGFSGADLANLLNEAAILTVRRGKVEIT 400
Query: 648 TDDLLQAAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGR 707
++ + G+ +A +EA AVAA LP+ D ++ +T+ PR R
Sbjct: 401 MKEIEDSIDKIIAGLEGSPLADSRIKRLIAYHEAGHAVAATFLPHHDPVQKVTLIPR--R 458
Query: 708 ELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARV 767
+ + L N + ++++ + I LA RA +E+ FG +++ A
Sbjct: 459 QAKGLTWFLP--NDDQFLVSKSQILSKIIAALAGRAMEEIVFGLPEVTIGAANDIKQVTF 516
Query: 768 AAR-------MYMIGGLSDKYRGVSNFWVTD----------RINEIDLEAMKILNLCYER 810
AR M +G + + F D + ++DLE IL CY +
Sbjct: 517 MARQMVTKFGMSKVGPICLENSSSEVFIGRDLMGRHELSEEMVAKVDLEVRSILKDCYIQ 576
Query: 811 AKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQ 845
A+ IL QN+ L+D +VNELV K+T+ ++ +R+V+
Sbjct: 577 ARTILSQNRKLIDRVVNELVEKETIEAKEFMRIVE 611
>FTSH_TOBAC (O82150) Cell division protein ftsH homolog, chloroplast
precursor (EC 3.4.24.-) (DS9)
Length = 714
Score = 313 bits (801), Expect = 2e-84
Identities = 186/449 (41%), Positives = 268/449 (59%), Gaps = 22/449 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG + +LEL+E+V F + + Y G KIP G LL GPPG GKTLLA+AVAGE
Sbjct: 250 VTFADVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGE 309
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AGV FFS +AS+FVE++VGVGASRVR L+++AK AP +VFIDE+DAVGR+RG G G
Sbjct: 310 AGVPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRGAGMGGGN 369
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ T+NQLL +DGF G VI +A+TNRPD+LD AL+RPGRFDR++ + +P GRI
Sbjct: 370 DEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRI 429
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+IL+VH+R K + +DVD+E +A T G GA+L N++ AAI R E+S D++ A
Sbjct: 430 KILQVHSRGKALTKDVDFEKIARRTPGYTGADLQNLMNEAAILAARRELKEISKDEISDA 489
Query: 655 AQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYV 712
+ G ++K S EK + VA +EA A+ +P +D + I+I PR G+ G
Sbjct: 490 LERIIAGP-EKKNAVVSDEKKKLVAYHEAGHALVGALMPEYDPVAKISIIPR-GQAGGLT 547
Query: 713 RTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNARVAARM 771
G+ +R L + + V L R A+E+ FG+D ++T + +RVA +M
Sbjct: 548 FFAPSEERLESGLYSRSYLENQMAVALGERVAEEVIFGQDNVTTGASNDFMQVSRVARQM 607
Query: 772 YMIGGLSDKY------RGVSNFWVTDRINE-----------IDLEAMKILNLCYERAKEI 814
G S K G N ++ +++ +D E +++ YERA EI
Sbjct: 608 VERLGFSKKIGQVAIGGGGGNPFLGQQMSTQKDYSMATADVVDAEVRELVERAYERATEI 667
Query: 815 LQQNKTLMDTLVNELVVKKTLTKEDIVRL 843
+ + ++ L L+ K+T+ E+ + L
Sbjct: 668 ITTHIDILHKLAQLLIEKETVDGEEFMSL 696
>FTSH_BACSU (P37476) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 637
Score = 313 bits (801), Expect = 2e-84
Identities = 200/476 (42%), Positives = 271/476 (56%), Gaps = 29/476 (6%)
Query: 394 GARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGG 453
G+RV + Y E VKF DVAG + + EL E+V+F + G +IP G
Sbjct: 137 GSRVMNFGKSKAKLYTEEKKRVKFKDVAGADEEKQELVEVVEFLKDPRKFAELGARIPKG 196
Query: 454 ILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSV 513
+LL GPPG GKTLLAKA AGEAGV FFSIS S FVE++VGVGASRVR L++ AK+NAP +
Sbjct: 197 VLLVGPPGTGKTLLAKACAGEAGVPFFSISGSDFVEMFVGVGASRVRDLFENAKKNAPCL 256
Query: 514 VFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPAL 573
+FIDE+DAVGR+RG G G ER+ TLNQLLV +DGF +I IA+TNR DILDPAL
Sbjct: 257 IFIDEIDAVGRQRGAGLGGGHDEREQTLNQLLVEMDGFSANEGIIIIAATNRADILDPAL 316
Query: 574 VRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEV 633
+RPGRFDR+I + +P IGR +LKVHAR KP+ E V+ + +A T G GA+L N++
Sbjct: 317 LRPGRFDRQITVDRPDVIGREAVLKVHARNKPLDETVNLKSIAMRTPGFSGADLENLLNE 376
Query: 634 AAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLP 691
AA+ R ++ ++ D+ +A G +K R SK++ VA +E V + L
Sbjct: 377 AALVAARQNKKKIDARDIDEATDRVIAGPA-KKSRVISKKERNIVAYHEGGHTVIGLVLD 435
Query: 692 NFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGK 751
D + +TI PR G+ GY + + T+ L D I L R A+E+ FG
Sbjct: 436 EADMVHKVTIVPR-GQAGGYAVMLPREDRY---FQTKPELLDKIVGLLGGRVAEEIIFG- 490
Query: 752 DQLSTIWAETADNA-RVAARMYMIGGLSDKY--------RGVSNFWVTDRIN-------- 794
++ST A +A RM G+S+K +G F D N
Sbjct: 491 -EVSTGAHNDFQRATNIARRMVTEFGMSEKLGPLQFGQSQGGQVFLGRDFNNEQNYSDQI 549
Query: 795 --EIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQLHG 848
EID E +I+ CYERAK+IL +N+ ++ + L+ +TL E I L+ HG
Sbjct: 550 AYEIDQEIQRIIKECYERAKQILTENRDKLELIAQTLLKVETLDAEQIKHLID-HG 604
>FTSH_BACPF (P94304) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 679
Score = 310 bits (795), Expect = 8e-84
Identities = 199/472 (42%), Positives = 273/472 (57%), Gaps = 28/472 (5%)
Query: 394 GARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGG 453
G+RV + E KF DVAG + + EL E+V+F + G +IP G
Sbjct: 142 GSRVMNFGKSKAKMVNEDKKKAKFKDVAGADEEKQELVEVVEFLKDPRKFSAIGARIPKG 201
Query: 454 ILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSV 513
+LL GPPG GKTLLA+AVAGEAGV FFSIS S FVE++VGVGASRVR L++ AK+NAP +
Sbjct: 202 VLLVGPPGTGKTLLARAVAGEAGVPFFSISGSDFVEMFVGVGASRVRDLFENAKKNAPCI 261
Query: 514 VFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPAL 573
+FIDE+DAVGR+RG G G ER+ TLNQLLV +DGF +I IA+TNR DILDPAL
Sbjct: 262 IFIDEIDAVGRQRGAGLGGGHDEREQTLNQLLVEMDGFSANEGIIIIAATNRADILDPAL 321
Query: 574 VRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEV 633
+RPGRFDR+I + +P GR E+LKVHAR KP+ +DV+ + +A+ T G GA+L N++
Sbjct: 322 LRPGRFDRQIQVNRPDVNGREEVLKVHARNKPLNDDVNLKTIATRTPGFSGADLENLLNE 381
Query: 634 AAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLP 691
AA+ R T++S + +A G +K R S ++ + VA +EA V + L
Sbjct: 382 AALVAARHDHTKISMIHIEEAIDRVIAGPA-KKSRVISPKEKKIVAWHEAGHTVVGVKLE 440
Query: 692 NFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGK 751
N D + +TI PR G GY + + + +T+ L D I L R A+E+ FG
Sbjct: 441 NADMVHKVTIVPR-GMAGGYAVMLPKEDRY---FMTQPELLDKIIGLLGGRVAEEVTFG- 495
Query: 752 DQLSTIWAETADNAR-VAARMYMIGGLSDKY----------------RGVSNFW-VTDRI 793
++ST A +A +M G+S+K R + N +D I
Sbjct: 496 -EVSTGAHNDFQRATGIARKMVTEYGMSEKLGPMQFISGSGGQVFLGRDIQNEQNYSDAI 554
Query: 794 -NEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
+EIDLE +I+ CY R K+IL +NK +D + L+ +TL E I LV
Sbjct: 555 AHEIDLEVQRIIKECYARCKQILLENKDSLDLVAKTLLDMETLDAEQIKSLV 606
>FTH1_ARATH (Q39102) Cell division protein ftsH homolog 1,
chloroplast precursor (EC 3.4.24.-)
Length = 716
Score = 309 bits (792), Expect = 2e-83
Identities = 183/449 (40%), Positives = 268/449 (58%), Gaps = 22/449 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG + +LEL+E+V F + + Y G KIP G LL GPPG GKTLLA+AVAGE
Sbjct: 259 VSFADVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGE 318
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AGV FFS +AS+FVE++VGVGASRVR L+++AK AP +VFIDE+DAVGR+RG G G
Sbjct: 319 AGVPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRGAGMGGGN 378
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ T+NQLL +DGF G VI +A+TNRPD+LD AL+RPGRFDR++ + +P GR+
Sbjct: 379 DEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRV 438
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+IL+VH+R K + +DVD++ VA T G GA+L N++ AAI R E+S D++ A
Sbjct: 439 KILQVHSRGKALGKDVDFDKVARRTPGFTGADLQNLMNEAAILAARRELKEISKDEISDA 498
Query: 655 AQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYV 712
+ G ++K S+EK VA +EA A+ +P +D + I+I PR G+ G
Sbjct: 499 LERIIAGP-EKKNAVVSEEKKRLVAYHEAGHALVGALMPEYDPVAKISIIPR-GQAGGLT 556
Query: 713 RTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNARVAARM 771
G+ +R L + + V L R A+E+ FG + ++T + +RVA +M
Sbjct: 557 FFAPSEERLESGLYSRSYLENQMAVALGGRVAEEVIFGDENVTTGASNDFMQVSRVARQM 616
Query: 772 YMIGGLSDKYRGVS------NFWVTDRINE-----------IDLEAMKILNLCYERAKEI 814
G S K V+ N ++ +++ +D E +++ Y+RA EI
Sbjct: 617 IERFGFSKKIGQVAVGGPGGNPFMGQQMSSQKDYSMATADIVDAEVRELVEKAYKRATEI 676
Query: 815 LQQNKTLMDTLVNELVVKKTLTKEDIVRL 843
+ + ++ L L+ K+T+ E+ + L
Sbjct: 677 ITTHIDILHKLAQLLIEKETVDGEEFMSL 705
>FTH2_SYNY3 (P73179) Cell division protein ftsH homolog 2 (EC
3.4.24.-)
Length = 665
Score = 308 bits (790), Expect = 3e-83
Identities = 183/463 (39%), Positives = 265/463 (56%), Gaps = 21/463 (4%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + EL+E+V F E + G KIP G+LL GPPG GKTLLA
Sbjct: 202 MEAKTGVGFDDVAGIDEAKEELQEVVTFLKQPEKFTAIGAKIPRGVLLIGPPGTGKTLLA 261
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AKENAP +VFIDE+DAVGR+RG+
Sbjct: 262 KAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENAPCLVFIDEIDAVGRQRGV 321
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG +I IA+TNRPD+LD AL+RPGRFDR++ + P
Sbjct: 322 GYGGGNDEREQTLNQLLTEMDGFEGNSGIIVIAATNRPDVLDLALLRPGRFDRQVTVDYP 381
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
GR IL +HA+ K + E+V +A T G GA+LAN++ AAI R + ++
Sbjct: 382 DVQGRELILAIHAQNKKLHEEVQLAAIARRTPGFTGADLANVLNEAAIFTARRRKEAITM 441
Query: 649 DDLLQAAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRE 708
++ A GM + +A +E A+ P D +E +T+ PR G+
Sbjct: 442 AEVNDAIDRVVAGMEGTPLVDSKSKRLIAYHEVGHALIGTLCPGHDPVEKVTLIPR-GQA 500
Query: 709 LGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVA 768
G + + ++TR + I L R A+E+ FG D+++T +
Sbjct: 501 QGLTWFTPDE---DQSLMTRNQMIARIAGLLGGRVAEEVIFGDDEVTTGAGNDIEKITYL 557
Query: 769 AR-------MYMIGGLSDKYRGVSNF----------WVTDRINEIDLEAMKILNLCYERA 811
AR M +G ++ + G NF + D ID E I+ ++RA
Sbjct: 558 ARQMVTKLGMSSLGLVALEEEGDRNFSGGDWGKRSEYSEDIAARIDREIQAIVTAAHQRA 617
Query: 812 KEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQLHGHAKPIP 854
I+++N+ LMD LV+ L+ ++T+ E +LV+ + ++ P
Sbjct: 618 TRIIEENRNLMDLLVDALIDQETIEGEHFRQLVESYQQSQKQP 660
>FTSH_STRR6 (P59652) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 652
Score = 307 bits (786), Expect = 9e-83
Identities = 185/450 (41%), Positives = 266/450 (59%), Gaps = 24/450 (5%)
Query: 413 VDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
+ V+F+DVAG + + EL E+V+F + + + G +IP G+LL GPPG GKTLLAKAVA
Sbjct: 182 IKVRFSDVAGAEEEKQELVEVVEFLKDPKRFTKLGARIPAGVLLEGPPGTGKTLLAKAVA 241
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
GEAGV FFSIS S FVE++VGVGASRVRSL+++AK+ AP+++FIDE+DAVGR+RG+ G
Sbjct: 242 GEAGVPFFSISGSDFVEMFVGVGASRVRSLFEDAKKAAPAIIFIDEIDAVGRQRGVGLGG 301
Query: 533 GGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIG 592
G ER+ TLNQLL+ +DGFEG +I IA+TNR D+LDPAL+RPGRFDRK+ + +P G
Sbjct: 302 GNDEREQTLNQLLIEMDGFEGNEGIIVIAATNRSDVLDPALLRPGRFDRKVLVGRPDVKG 361
Query: 593 RIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLL 652
R ILKVHA+ KP+AEDVD ++VA T G VGA+L N++ AA+ R +++ + D+
Sbjct: 362 REAILKVHAKNKPLAEDVDLKLVAQQTPGFVGADLENVLNEAALVAARRNKSIIDASDID 421
Query: 653 QAAQMEERGMLDR-KERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGY 711
+A G + K S+++ E VA +EA + + L N + +TI PR GR GY
Sbjct: 422 EAEDRVIAGPSKKDKTVSQKERELVAYHEAGHTIVGLVLSNARVVHKVTIVPR-GRAGGY 480
Query: 712 VRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQ--LSTIWAETADNARVAA 769
M+ + +L+++ + + + + R A+E+ F S + + AR
Sbjct: 481 ---MIALPKEDQMLLSKEDMKEQLAGLMGGRVAEEIIFNVQTTGASNDFEQATQMARAMV 537
Query: 770 RMYMIGGLSDK-----YRG---------VSNFWVTDRINEIDLEAMKILNLCYERAKEIL 815
Y G+S+K Y G EID E +LN +A EI+
Sbjct: 538 TEY---GMSEKLGPVQYEGNHAMLGAQSPQKSISEQTAYEIDEEVRSLLNEARNKAAEII 594
Query: 816 QQNKTLMDTLVNELVVKKTLTKEDIVRLVQ 845
Q N+ + L+ +TL I L +
Sbjct: 595 QSNRETHKLIAEALLKYETLDSTQIKALYE 624
>FTSH_STRPN (O69076) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 652
Score = 307 bits (786), Expect = 9e-83
Identities = 185/450 (41%), Positives = 266/450 (59%), Gaps = 24/450 (5%)
Query: 413 VDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
+ V+F+DVAG + + EL E+V+F + + + G +IP G+LL GPPG GKTLLAKAVA
Sbjct: 182 IKVRFSDVAGAEEEKQELVEVVEFLKDPKRFTKLGARIPAGVLLEGPPGTGKTLLAKAVA 241
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
GEAGV FFSIS S FVE++VGVGASRVRSL+++AK+ AP+++FIDE+DAVGR+RG+ G
Sbjct: 242 GEAGVPFFSISGSDFVEMFVGVGASRVRSLFEDAKKAAPAIIFIDEIDAVGRQRGVGLGG 301
Query: 533 GGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIG 592
G ER+ TLNQLL+ +DGFEG +I IA+TNR D+LDPAL+RPGRFDRK+ + +P G
Sbjct: 302 GNDEREQTLNQLLIEMDGFEGNEGIIVIAATNRSDVLDPALLRPGRFDRKVLVGRPDVKG 361
Query: 593 RIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLL 652
R ILKVHA+ KP+AEDVD ++VA T G VGA+L N++ AA+ R +++ + D+
Sbjct: 362 REAILKVHAKNKPLAEDVDLKLVAQQTPGFVGADLENVLNEAALVAARRNKSIIDASDID 421
Query: 653 QAAQMEERGMLDR-KERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGY 711
+A G + K S+++ E VA +EA + + L N + +TI PR GR GY
Sbjct: 422 EAEDRVIAGPSKKDKTVSQKERELVAYHEAGHTIVGLVLSNARVVHKVTIVPR-GRAGGY 480
Query: 712 VRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQ--LSTIWAETADNARVAA 769
M+ + +L+++ + + + + R A+E+ F S + + AR
Sbjct: 481 ---MIALPKEDQMLLSKEDMKEQLAGLMGGRVAEEIIFNVQTTGASNDFEQATQMARAMV 537
Query: 770 RMYMIGGLSDK-----YRG---------VSNFWVTDRINEIDLEAMKILNLCYERAKEIL 815
Y G+S+K Y G EID E +LN +A EI+
Sbjct: 538 TEY---GMSEKLGPVQYEGNHAMLGAQSPQKSISEQTAYEIDEEVRSLLNEARNKAAEII 594
Query: 816 QQNKTLMDTLVNELVVKKTLTKEDIVRLVQ 845
Q N+ + L+ +TL I L +
Sbjct: 595 QSNRETHKLIAEALLKYETLDSTQIKALYE 624
>FTSH_MEDSA (Q9BAE0) Cell division protein ftsH homolog, chloroplast
precursor (EC 3.4.24.-)
Length = 706
Score = 307 bits (786), Expect = 9e-83
Identities = 182/449 (40%), Positives = 269/449 (59%), Gaps = 23/449 (5%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG + +LEL+E+V F + + Y G KIP G LL GPPG GKTLLA+AVAGE
Sbjct: 250 VTFADVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGE 309
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AG FFS +AS+FVE++VGVGASRVR L+++AK AP +VFIDE+DAVGR+RG G G
Sbjct: 310 AGTPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRGAGLGGGN 369
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ T+NQLL +DGF G VI +A+TNRPD+LD AL+RPGRFDR++ + +P GR+
Sbjct: 370 DEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRV 429
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+IL+VH+R K +A+DVD++ +A T G G +L N++ AAI R E+S D++ A
Sbjct: 430 KILQVHSRGKALAKDVDFDKIARRTPGFTGVDLQNLMNEAAILAARRDLKEISKDEIADA 489
Query: 655 AQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYV 712
+ G ++K S+EK + VA +EA A+ +P +D + I+I PR G+ G
Sbjct: 490 LERIIAGP-EKKNAVVSEEKKKLVAYHEAGHALVGALMPEYDPVAKISIIPR-GQAGGLT 547
Query: 713 RTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNARVAARM 771
G+ +R L + + V L R A+E+ FG+D ++T + +RVA +M
Sbjct: 548 FFAPSEERLESGLYSRSYLENQMAVALGGRVAEEV-FGQDNVTTGASNDFMQVSRVARQM 606
Query: 772 YMIGGLSDKY------RGVSNFWVTDRINE-----------IDLEAMKILNLCYERAKEI 814
G S K G N ++ +++ +D E ++++ YERA +I
Sbjct: 607 VERFGFSKKIGQVAIGGGGGNPFLGQQMSSQKDYSMATADIVDKEVRELVDKAYERATQI 666
Query: 815 LQQNKTLMDTLVNELVVKKTLTKEDIVRL 843
+ + ++ L L+ K+T+ E+ + L
Sbjct: 667 INTHIDILHKLAQLLIEKETVDGEEFMSL 695
>FTSH_ODOSI (P49825) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 644
Score = 305 bits (781), Expect = 3e-82
Identities = 178/451 (39%), Positives = 264/451 (58%), Gaps = 22/451 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F D+AG+ + + E EEIV F + Y G KIP GILL GPPG GKTLLAKA+A E
Sbjct: 183 VSFKDIAGIDEAKTEFEEIVSFLKEPDKYTIVGAKIPKGILLVGPPGTGKTLLAKAIANE 242
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
A V FFS++ S+FVE+++G+GA+RVR L+++A ENAP +VFIDE+DAVGR+RG G G
Sbjct: 243 ADVPFFSVAGSEFVEMFIGIGAARVRDLFKKASENAPCIVFIDEIDAVGRERGAGVGGGN 302
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ TLNQLL +DGF+ VI + +TNR DILD AL+RPGRFDR++ + P +GR+
Sbjct: 303 DEREQTLNQLLTEMDGFKENKGVIVVGATNRADILDAALLRPGRFDRQVTVNLPDRLGRV 362
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
ILKVHAR KP+ EDV +A+ T G GA+LAN++ AAI R ++ ++ +++ +A
Sbjct: 363 GILKVHARNKPLGEDVSLVQLANRTPGFSGADLANLLNEAAILATRYKKSSITKNEVNEA 422
Query: 655 AQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRT 714
A G+ + +A +E A+ L + D +E IT+ PR G + G
Sbjct: 423 ADRIIGGIAGAPMEDTKNKRLIAYHEVGHAITGSVLKSHDEVEKITLTPRGGAK-GLTWF 481
Query: 715 MLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAAR---- 770
E + +L+R +L I L RAA+++ FG+ +++T + AR
Sbjct: 482 TPEE---DQSLLSRSALLARIITTLGGRAAEQVIFGEPEVTTGASSDLQQVTNLARQMVT 538
Query: 771 ---MYMIG--GLSDKYRG---------VSNFWVTDRINEIDLEAMKILNLCYERAKEILQ 816
M IG L D+ G + + + + ID E KI+ CYE+A EI+
Sbjct: 539 RFGMSNIGPLALEDESTGQVFLGGNMASGSEYAENIADRIDDEVRKIITYCYEKAIEIVL 598
Query: 817 QNKTLMDTLVNELVVKKTLTKEDIVRLVQLH 847
N+ ++D +V +L+ K+T+ ++ L+ +
Sbjct: 599 DNRVVIDLIVEKLLDKETMDGDEFRELLSTY 629
>FTSH_CAPAN (Q39444) Cell division protein ftsH homolog, chloroplast
precursor (EC 3.4.24.-) (Fragment)
Length = 662
Score = 305 bits (781), Expect = 3e-82
Identities = 181/438 (41%), Positives = 261/438 (59%), Gaps = 22/438 (5%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG + +LEL+E+V F + + Y G KIP G LL GPPG GKTLLA+AVAGE
Sbjct: 227 VTFADVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGE 286
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AGV FFS +AS+FVE++VGVGASRVR L++ AK AP +VFIDE+DAVGR+RG G G
Sbjct: 287 AGVPFFSCAASEFVELFVGVGASRVRHLFENAKSKAPCIVFIDEIDAVGRQRGAGLGGGN 346
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ T+NQLL +DGF G VI +A+TNRPD+LD AL+RPG+FDR++ + +P GR+
Sbjct: 347 DEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGKFDRQVTVDRPDVAGRV 406
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
IL+VH+R K +A+DVD++ +A T G GA+L N++ AAI R E+S D++ A
Sbjct: 407 RILQVHSRGKALAKDVDFDKIARRTPGFTGADLQNLMNEAAILAARRDLKEISKDEISDA 466
Query: 655 AQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYV 712
+ G ++K S EK + VA +EA A+ +P +D + I+I PR G+ G
Sbjct: 467 LERIIAGP-EKKNAVVSDEKKKLVAYHEAGHALVGALMPEYDPVAKISIIPR-GQAGGLT 524
Query: 713 RTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNARVAARM 771
G+ +R L + + V L R A+E+ FG+D ++T + +RVA +M
Sbjct: 525 FFAPSEERLESGLYSRSYLENQMAVALGGRVAEEVIFGEDNVTTGASNDFMQVSRVARQM 584
Query: 772 YMIGGLSDKY------RGVSNFWVTDRINE-----------IDLEAMKILNLCYERAKEI 814
G S K G N ++ +++ +D E +++ YERAK+I
Sbjct: 585 VERLGFSKKIGQVAIGGGGGNPFLGQQMSTQKDYSMATADVVDSEVRELVEKAYERAKQI 644
Query: 815 LQQNKTLMDTLVNELVVK 832
+ + ++ L L+ K
Sbjct: 645 ITTHIDILHKLAQLLIEK 662
>FTSH_PORPU (P51327) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 628
Score = 305 bits (780), Expect = 5e-82
Identities = 185/460 (40%), Positives = 267/460 (57%), Gaps = 33/460 (7%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + E +E+V F E + G KIP G+LL GPPG GKTLLA
Sbjct: 164 MEAKTGVVFNDVAGVEEAKEEFQEVVTFLKQPESFTAVGAKIPKGVLLVGPPGTGKTLLA 223
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AK+NAP +VFIDE+DAVGR+RG
Sbjct: 224 KAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKDNAPCIVFIDEIDAVGRQRGT 283
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG VI IA+TNR DILD AL+RPGRFDR++ + P
Sbjct: 284 GVGGGNDEREQTLNQLLTEMDGFEGNTGVIVIAATNRADILDSALLRPGRFDRQVSVDVP 343
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSR----- 643
F GR+ IL+VHA+ K + V E +A T G GA+LAN++ AAI R +
Sbjct: 344 DFRGRLAILEVHAKNKKMESKVSLETIARRTPGFSGADLANLLNEAAILTARRRKSAMTM 403
Query: 644 TEVSTDDLLQAAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAP 703
+E+ T A +E ++D K + +A +E A+ L + D ++ +T+ P
Sbjct: 404 SEIDTSIDRVVAGLEGTPLIDSKSK-----RLIAYHEVGHAIIGSLLEHHDPVQKVTLIP 458
Query: 704 RAGRELGYVRTMLESINFND-GMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETA 762
R G R + +D +++R + I L RAA+E+ FG +++T +
Sbjct: 459 R-----GQARGLTWFTPSDDQSLISRSQILARIVGALGGRAAEEIIFGDAEVTTGASNDL 513
Query: 763 DNARVAAR-------MYMIGGLSDKYRGVSNF----------WVTDRINEIDLEAMKILN 805
AR M IG LS + +G F + + ID + +I++
Sbjct: 514 QQVTSMARQMVTRFGMSKIGPLSLESQGSDPFLGRGMGGGSEYSDEVATNIDKQVREIVS 573
Query: 806 LCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQ 845
CY+ AK+I++ N+ +MD LV+ L+ K+T+ + +V+
Sbjct: 574 ECYKEAKKIVKDNRVVMDRLVDLLIEKETIEGNEFRHIVK 613
>FTSH_MYCTU (P96942) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 760
Score = 303 bits (775), Expect = 2e-81
Identities = 188/470 (40%), Positives = 273/470 (58%), Gaps = 26/470 (5%)
Query: 391 MKSGARVRRAQNRRLPQYLERGVD-VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVK 449
M+ GAR+ + + L + + F DVAG+ + EL EI F + Y+ G K
Sbjct: 135 MQGGARMGFGFGKSRAKQLSKDMPKTTFADVAGVDEAVEELYEIKDFLQNPSRYQALGAK 194
Query: 450 IPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKEN 509
IP G+LL GPPG GKTLLA+AVAGEAGV FF+IS S FVE++VGVGASRVR L+++AK+N
Sbjct: 195 IPKGVLLYGPPGTGKTLLARAVAGEAGVPFFTISGSDFVEMFVGVGASRVRDLFEQAKQN 254
Query: 510 APSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDIL 569
+P ++F+DE+DAVGR+RG G G ER+ TLNQLLV +DGF R VI IA+TNRPDIL
Sbjct: 255 SPCIIFVDEIDAVGRQRGAGLGGGHDEREQTLNQLLVEMDGFGDRAGVILIAATNRPDIL 314
Query: 570 DPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELAN 629
DPAL+RPGRFDR+I + P GR +L+VH++ KP+A D D + +A T GM GA+LAN
Sbjct: 315 DPALLRPGRFDRQIPVSNPDLAGRRAVLRVHSKGKPMAADADLDGLAKRTVGMTGADLAN 374
Query: 630 IVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAA 687
++ AA+ R++ T V T L+ A G RK R S+++ + A +E +AA
Sbjct: 375 VINEAALLTARENGT-VITGPALEEAVDRVIGGPRRKGRIISEQEKKITAYHEGGHTLAA 433
Query: 688 MNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEM 747
+P+ + I +TI R GR G+ + E + G+ TR + + + RAA+E+
Sbjct: 434 WAMPDIEPIYKVTILAR-GRTGGHAVAVPEE---DKGLRTRSEMIAQLVFAMGGRAAEEL 489
Query: 748 WFGKDQLSTIWAETADNARVAARMYMIGGLSDKY-----------------RGVSNFWVT 790
F + + ++ ++A M G+S K G +
Sbjct: 490 VFREPTTGAV-SDIEQATKIARSMVTEFGMSSKLGAVKYGSEHGDPFLGRTMGTQPDYSH 548
Query: 791 DRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDI 840
+ EID E K++ + A EIL + + ++DTL EL+ K+TL + ++
Sbjct: 549 EVAREIDEEVRKLIEAAHTEAWEILTEYRDVLDTLAGELLEKETLHRPEL 598
>FTH1_SYNY3 (Q55700) Cell division protein ftsH homolog 1 (EC
3.4.24.-)
Length = 627
Score = 302 bits (774), Expect = 2e-81
Identities = 178/447 (39%), Positives = 262/447 (57%), Gaps = 21/447 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG+ + + EL+E+V F E + G KIP G+LL GPPG GKTLLAKA+AGE
Sbjct: 169 VMFDDVAGIDEAKEELQEVVTFLKQPERFTAVGAKIPKGVLLVGPPGTGKTLLAKAIAGE 228
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AGV FFSIS S+FVE++VGVGASRVR L+++AKENAP ++FIDE+DAVGR+RG G G
Sbjct: 229 AGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENAPCLIFIDEIDAVGRQRGAGIGGGN 288
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ TLNQLL +DGFEG +I IA+TNRPD+LD AL+RPGRFDR++ + P + GR
Sbjct: 289 DEREQTLNQLLTEMDGFEGNTGIIIIAATNRPDVLDSALMRPGRFDRQVMVDAPDYSGRK 348
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
EIL+VHAR K +A +V + +A T G GA+LAN++ AAI R ++ ++ ++ A
Sbjct: 349 EILEVHARNKKLAPEVSIDSIARRTPGFSGADLANLLNEAAILTARRRKSAITLLEIDDA 408
Query: 655 AQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRT 714
GM + +A +E A+ L + D ++ +T+ PR G+ G
Sbjct: 409 VDRVVAGMEGTPLVDSKSKRLIAYHEVGHAIVGTLLKDHDPVQKVTLIPR-GQAQGLT-- 465
Query: 715 MLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAAR---- 770
+ N G+ T+ L I + RAA+E FG D+++T AR
Sbjct: 466 -WFTPNEEQGLTTKAQLMARIAGAMGGRAAEEEVFGDDEVTTGAGGDLQQVTEMARQMVT 524
Query: 771 ---MYMIGGLSDKYRGVSNF----------WVTDRINEIDLEAMKILNLCYERAKEILQQ 817
M +G +S + G F + + ID + ++ ++ A++I+Q+
Sbjct: 525 RFGMSNLGPISLESSGGEVFLGGGLMNRSEYSEEVATRIDAQVRQLAEQGHQMARKIVQE 584
Query: 818 NKTLMDTLVNELVVKKTLTKEDIVRLV 844
+ ++D LV+ L+ K+T+ E+ ++V
Sbjct: 585 QREVVDRLVDLLIEKETIDGEEFRQIV 611
>FTSH_GUITH (O78516) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 631
Score = 302 bits (773), Expect = 3e-81
Identities = 179/458 (39%), Positives = 268/458 (58%), Gaps = 31/458 (6%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + E EE+V F E + G KIP G+LL GPPG GKTLLA
Sbjct: 164 MEAKTGVTFNDVAGVDEAKEEFEEVVSFLKKPERFTAVGAKIPKGVLLVGPPGTGKTLLA 223
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AKEN+P +VFIDE+DAVGR+RG
Sbjct: 224 KAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENSPCIVFIDEIDAVGRQRGT 283
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG +I IA+TNR D+LD AL+RPGRFDR++ + P
Sbjct: 284 GIGGGNDEREQTLNQLLTEMDGFEGNTGIIIIAATNRVDVLDAALLRPGRFDRQVTVDVP 343
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
GR+EIL VHAR K + + E++A T G GA+LAN++ AAI R + +++
Sbjct: 344 DVKGRLEILNVHARNKKLDLSISLELIAKRTPGFSGADLANLLNEAAILTARRRKKQITI 403
Query: 649 DDLLQA-----AQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAP 703
++ + A ME + ++D K + +A +E A+ L + D ++ +T+ P
Sbjct: 404 SEIDASIDRVIAGMEGKALVDSKTK-----RLIAYHEVGHAIIGTLLKHHDPVQKVTLVP 458
Query: 704 RAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETAD 763
R G+ G + + + +++R + I L RAA+E+ FG +++T
Sbjct: 459 R-GQAKGLT---WFTPSEDQSLISRSQILARIMGALGGRAAEEVVFGLPEVTTGAGNDLQ 514
Query: 764 NARVAAR-------MYMIGGLS----------DKYRGVSNFWVTDRINEIDLEAMKILNL 806
AR M IG LS + G S+ + D + ID++ I+
Sbjct: 515 QVTSMARQMVTRFGMSNIGPLSLESQNSDPFLGRTMGSSSQYSEDIASRIDMQVRAIIQH 574
Query: 807 CYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
C+ +I++ N+ ++D LV+ L+ K+T+ ++ ++V
Sbjct: 575 CHTETVQIIKDNRVVIDKLVDLLIEKETIDGDEFRQIV 612
>FTSH_RICPR (Q9ZEA2) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 637
Score = 301 bits (770), Expect = 7e-81
Identities = 186/451 (41%), Positives = 264/451 (58%), Gaps = 20/451 (4%)
Query: 410 ERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAK 469
++G + F DVAG+ + + EL EIV F +++ G KIP G LL GPPG GKTLLAK
Sbjct: 147 DKGPKITFKDVAGIDEAKEELTEIVDFLRDPSKFQKLGGKIPKGCLLIGPPGTGKTLLAK 206
Query: 470 AVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLI 529
A+AGEA V FFSIS S FVE++VGVGASRVR ++++ K NAP ++FIDE+DAVGR RG+
Sbjct: 207 AIAGEANVPFFSISGSDFVEMFVGVGASRVRDMFEQGKRNAPCIIFIDEIDAVGRHRGIG 266
Query: 530 KGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPG 589
G G ER+ TLNQ+LV +DGFE V+ IA+TNRPD+LD AL+RPGRFDR+I + P
Sbjct: 267 MGGGNDEREQTLNQMLVEMDGFEANEGVVIIAATNRPDVLDRALLRPGRFDRQIAVANPD 326
Query: 590 FIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTD 649
GR +ILKVH +K V I+A T G GAELAN+V AA+ R + EV
Sbjct: 327 INGREQILKVHLKKIKYNSTVLARIIARGTPGFSGAELANLVNEAALIAARLGKKEVDMH 386
Query: 650 DLLQAAQMEERGMLDRKERSKEKWEQV-AINEAAMAVAAMNLPNFDNIEYITIAPRAGRE 708
D+ +A G++ R EK +++ A +E A+ + P I TI PR G
Sbjct: 387 DMEEAKDKVLMGVVRRSIAMSEKEKRLTAYHEGGHALVGLYCPAASPIHKATIIPR-GNA 445
Query: 709 LGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNA-RV 767
LG V+ + E+ ++ R+ + I V +A R A+E+ FG++++++ + A +
Sbjct: 446 LGMVQRLPETDEYSQ---NREQMESSIAVYMAGRVAEEIIFGRNKVTSGASSDIKGATNI 502
Query: 768 AARMYMIGGLS-------------DKYRGVSNFWVTDRINE-IDLEAMKILNLCYERAKE 813
A M GLS D Y S+ +++ E ID E +I+ YE AK+
Sbjct: 503 ARAMVTKAGLSDLIGPIFHGSNSDDMYGRQSSNEISEATAELIDAEVKRIITQGYEFAKD 562
Query: 814 ILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
IL ++ + TL N L+ +TL+ + I L+
Sbjct: 563 ILTKHIDQLHTLANALIEYETLSGQQIKNLL 593
>FTSH_RICCN (Q92JJ9) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 637
Score = 299 bits (765), Expect = 2e-80
Identities = 187/451 (41%), Positives = 263/451 (57%), Gaps = 20/451 (4%)
Query: 410 ERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAK 469
++G + F DVAG+ + + EL EIV F +++ G KIP G LL GPPG GKTLLAK
Sbjct: 147 DKGPKITFKDVAGIDEAKEELTEIVDFLRDPSKFQKLGGKIPKGCLLIGPPGTGKTLLAK 206
Query: 470 AVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLI 529
A+AGEA V FFSIS S FVE++VGVGASRVR ++++ K NAP ++FIDE+DAVGR RG+
Sbjct: 207 AIAGEANVPFFSISGSDFVEMFVGVGASRVRDMFEQGKRNAPCIIFIDEIDAVGRHRGIG 266
Query: 530 KGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPG 589
G G ER+ TLNQ+LV +DGFE V+ IA+TNRPD+LD AL+RPGRFDR+I + P
Sbjct: 267 MGGGNDEREQTLNQMLVEMDGFEANEGVVIIAATNRPDVLDRALLRPGRFDRQIAVANPD 326
Query: 590 FIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTD 649
GR +ILKVH +K V I+A T G GAELAN+V AA+ R + EV
Sbjct: 327 INGREQILKVHLKKIKYNSTVLARIIARGTPGFSGAELANLVNEAALIAARLGKKEVDMH 386
Query: 650 DLLQAAQMEERGMLDRKERSKEKWEQV-AINEAAMAVAAMNLPNFDNIEYITIAPRAGRE 708
D+ +A G+ R EK +++ A +E A+ + P I TI PR G
Sbjct: 387 DMEEAKDKVLMGVARRSIAMSEKEKRLTAYHEGGHALVGLYCPAASPIHKATIIPR-GNA 445
Query: 709 LGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNA-RV 767
LG V+ + E+ ++ R+ + I V +A R A+E+ FG++++++ + A +
Sbjct: 446 LGMVQRLPETDEYSQ---NREQMESSIAVYMAGRVAEEIIFGRNKVTSGASSDIKGATNI 502
Query: 768 AARMYMIGGLSDK----YRGVS--NFWVTDRINE--------IDLEAMKILNLCYERAKE 813
A M GLSD + G S + + NE ID E +I+ YE AK+
Sbjct: 503 ARAMVTKAGLSDLIGPIFHGSSGDDMYGRQPNNETSEATAELIDAEVKRIITQGYEFAKD 562
Query: 814 ILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
IL ++ + TL N L+ +TL+ + I L+
Sbjct: 563 ILTKHIDQLHTLANALIEYETLSGQQIKNLL 593
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.134 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 98,054,778
Number of Sequences: 164201
Number of extensions: 4186708
Number of successful extensions: 25323
Number of sequences better than 10.0: 1106
Number of HSP's better than 10.0 without gapping: 547
Number of HSP's successfully gapped in prelim test: 580
Number of HSP's that attempted gapping in prelim test: 20758
Number of HSP's gapped (non-prelim): 3153
length of query: 881
length of database: 59,974,054
effective HSP length: 119
effective length of query: 762
effective length of database: 40,434,135
effective search space: 30810810870
effective search space used: 30810810870
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC138130.19


