

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138012.9 - phase: 0
(672 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
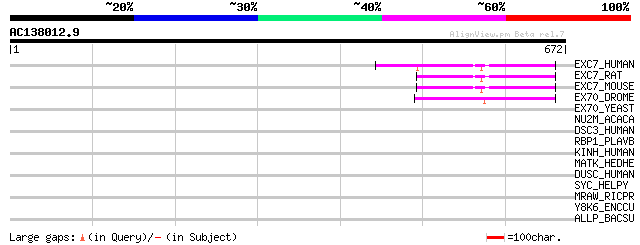
Score E
Sequences producing significant alignments: (bits) Value
EXC7_HUMAN (Q9UPT5) Exocyst complex component 7 (Exocyst complex... 73 3e-12
EXC7_RAT (O54922) Exocyst complex component 7 (Exocyst complex c... 70 1e-11
EXC7_MOUSE (O35250) Exocyst complex component 7 (Exocyst complex... 69 5e-11
EX70_DROME (Q9VSJ8) 70 kDa exocyst complex protein 47 2e-04
EX70_YEAST (P19658) 70 kDa exocyst complex protein 43 0.003
NU2M_ACACA (Q37376) NADH-ubiquinone oxidoreductase chain 2 (EC 1... 34 1.1
DSC3_HUMAN (Q14574) Desmocollin 3 precursor (Desmocollin 4) (HT-CP) 33 1.9
RBP1_PLAVB (Q00798) Reticulocyte binding protein 1 precursor 33 3.2
KINH_HUMAN (P33176) Kinesin heavy chain (Ubiquitous kinesin heav... 33 3.2
MATK_HEDHE (Q8WH60) Maturase K (Intron maturase) 32 4.2
DUSC_HUMAN (Q9UNI6) Dual specificity protein phosphatase 12 (EC ... 32 5.4
SYC_HELPY (P41259) Cysteinyl-tRNA synthetase (EC 6.1.1.16) (Cyst... 32 7.1
MRAW_RICPR (Q9ZCY2) S-adenosyl-methyltransferase mraW (EC 2.1.1.-) 32 7.1
Y8K6_ENCCU (Q8SUI0) Hypothetical protein ECU08_2060 31 9.3
ALLP_BACSU (P94575) Probable allantoin permease (Allantoin trans... 31 9.3
>EXC7_HUMAN (Q9UPT5) Exocyst complex component 7 (Exocyst complex
component Exo70)
Length = 735
Score = 72.8 bits (177), Expect = 3e-12
Identities = 59/239 (24%), Positives = 115/239 (47%), Gaps = 25/239 (10%)
Query: 444 GQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEASSFFLKQME-----QIMRML 498
G H++T + ++ + +L + SS F K++ +++ L
Sbjct: 497 GTVHELTSNAILFLQQLLDFQETAGAMLASQETSSSATSYSSEFSKRLLSTYICKVLGNL 556
Query: 499 QRKLIVKSENCKDRALRHIFMLNNRSHI-EAMNKFS--RLETIFGNDWFQNNKAKIQQNL 555
Q L+ KS+ +D AL IF+ NN ++I +++ K +L + ++ + I+Q +
Sbjct: 557 QLNLLSKSKVYEDPALSAIFLHNNYNYILKSLEKSELIQLVAVTQKTAERSYREHIEQQI 616
Query: 556 DLYKRSAWDEVMDFL------------KLDNNESITKELLKEKIHLFNNRFEAICRVQSA 603
Y+RS W +V D++ KL + E ++++KE+ FN+ E +C++Q A
Sbjct: 617 QTYQRS-WLKVTDYIAEKNLPVFQPGVKLRDKE---RQIIKERFKGFNDGLEELCKIQKA 672
Query: 604 WFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGI-LGNQAYKYIKYGMIEIQDLLNHLF 661
W I ++ R I + I+ YG F+ + + KYIKYG+ ++ D+++ LF
Sbjct: 673 WAIPDTEQRDRIRQAQKTIVKETYGAFLQKFGSVPFTKNPEKYIKYGVEQVGDMIDRLF 731
>EXC7_RAT (O54922) Exocyst complex component 7 (Exocyst complex
component Exo70) (rExo70)
Length = 653
Score = 70.5 bits (171), Expect = 1e-11
Identities = 51/185 (27%), Positives = 98/185 (52%), Gaps = 20/185 (10%)
Query: 493 QIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHI-EAMNKFS--RLETIFGNDWFQNNKA 549
+++ LQ L+ KS+ +D AL IF+ NN ++I +++ K +L + ++ +
Sbjct: 469 KVLGNLQLNLLSKSKVYEDPALSAIFLHNNYNYILKSLEKSELIQLVAVTQKTAERSYRE 528
Query: 550 KIQQNLDLYKRSAWDEVMDFL------------KLDNNESITKELLKEKIHLFNNRFEAI 597
I+Q + Y+RS W +V D++ KL + E ++++KE+ FN+ E +
Sbjct: 529 HIEQQIQTYQRS-WLKVTDYIAEKNLPVFQPGVKLRDKE---RQMIKERFKGFNDGLEEL 584
Query: 598 CRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGI-LGNQAYKYIKYGMIEIQDL 656
C++Q AW I ++ R +I + +I+ YG F+ R + KYIKY + ++ D+
Sbjct: 585 CKIQKAWAIPDTEQRDKIRQAQKSIVKETYGAFLHRYSSVPFTKNPEKYIKYRVEQVGDM 644
Query: 657 LNHLF 661
++ LF
Sbjct: 645 IDRLF 649
>EXC7_MOUSE (O35250) Exocyst complex component 7 (Exocyst complex
component Exo70)
Length = 697
Score = 68.6 bits (166), Expect = 5e-11
Identities = 50/185 (27%), Positives = 97/185 (52%), Gaps = 20/185 (10%)
Query: 493 QIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHI-EAMNKFS--RLETIFGNDWFQNNKA 549
+++ LQ L+ KS+ +D AL IF+ NN ++I +++ K +L + ++ +
Sbjct: 513 KVLGNLQLNLLSKSKVYEDPALSAIFLHNNYNYILKSLEKSELIQLVAVTQKTAERSYRE 572
Query: 550 KIQQNLDLYKRSAWDEVMDFL------------KLDNNESITKELLKEKIHLFNNRFEAI 597
I+Q + Y+RS W +V D++ KL + E ++++KE+ FN+ E +
Sbjct: 573 HIEQQIQTYQRS-WLKVTDYIAEKNLPVFQPGVKLRDKE---RQMIKERFKGFNDGLEEL 628
Query: 598 CRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGI-LGNQAYKYIKYGMIEIQDL 656
C++Q W I ++ R +I + +I+ YG F+ R + KYIKY + ++ D+
Sbjct: 629 CKIQKVWAIPDTEQRDKIRQAQKDIVKETYGAFLHRYGSVPFTKNPEKYIKYRVEQVGDM 688
Query: 657 LNHLF 661
++ LF
Sbjct: 689 IDRLF 693
>EX70_DROME (Q9VSJ8) 70 kDa exocyst complex protein
Length = 693
Score = 47.0 bits (110), Expect = 2e-04
Identities = 42/183 (22%), Positives = 81/183 (43%), Gaps = 12/183 (6%)
Query: 491 MEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHI-EAMNKFSRLETI-FGNDWFQNNK 548
+++ + L ++ K E D+A +H+F LNN +I +++ + + ++ + +++
Sbjct: 507 IKKALAELNLSIMNKCEQYNDQATKHLFRLNNIHYILKSLQRSNLIDLVTLAEPECEHSY 566
Query: 549 AKIQQNLDLYKRSAWDEVM-DFLKLD--------NNESITKELLKEKIHLFNNRFEAICR 599
++ + L + W +++ LD + + +LKE+ FN FE C+
Sbjct: 567 MEMIRELKASYQKTWSKMLVGIYSLDELPKPVAGKVKDKDRSVLKERFSNFNKDFEEACK 626
Query: 600 VQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGI-LGNQAYKYIKYGMIEIQDLLN 658
+Q I LR I +LP Y F G+ KY+KY EI +L+
Sbjct: 627 IQRGISIPDVILREGIKRDNVEHILPIYNRFYEIYSGVHFSKNPDKYVKYRQHEINAMLS 686
Query: 659 HLF 661
LF
Sbjct: 687 KLF 689
>EX70_YEAST (P19658) 70 kDa exocyst complex protein
Length = 623
Score = 42.7 bits (99), Expect = 0.003
Identities = 37/149 (24%), Positives = 70/149 (46%), Gaps = 8/149 (5%)
Query: 518 FMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEV----MDFLKLD 573
F+L N + +E + + S L + + + ++++ Y S W ++ MD + +D
Sbjct: 475 FILMNLTLVEQIVEKSELNLMLAGEG-HSRLERLKKRYISYMVSDWRDLTANLMDSVFID 533
Query: 574 NN--ESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFV 631
++ +S KE +KEK FN FE + + + L+ + S + ++++P Y F
Sbjct: 534 SSGKKSKDKEQIKEKFRKFNEGFEDLVSKTKQYKLSDPSLKVTLKSEIISLVMPMYERFY 593
Query: 632 GRLHGILGNQAYKYIKYGMIEIQDLLNHL 660
R N K+IKY E+ +LN L
Sbjct: 594 SRYKDSFKNPR-KHIKYTPDELTTVLNQL 621
>NU2M_ACACA (Q37376) NADH-ubiquinone oxidoreductase chain 2 (EC
1.6.5.3)
Length = 527
Score = 34.3 bits (77), Expect = 1.1
Identities = 50/244 (20%), Positives = 95/244 (38%), Gaps = 41/244 (16%)
Query: 1 MIPVIIRIRMCLLQTKV----WRFVGFA---SAAVGLLCYALSTSFNHLFGNWNLLKIFL 53
++P ++ + + +L T + F GF +A V L F L F
Sbjct: 75 LLPFVVILLLIILNTNTNDSYYLFAGFYYNDAAVVFFKNIILIGFLIFTFAIKQYLSYFK 134
Query: 54 YTLFSFIMILYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAA 113
Y F FI++L+ +++ ++ L+L + S FF +I +
Sbjct: 135 YYDFEFILVLFISLF-----------SSLLILNSNDLISLFF------------IIELQS 171
Query: 114 FAIMSMSLSRQ-SQCGIEVDLLYFFLGCLIMQLMKIKLQLVIVGAG-FSYS-LIIIRSSL 170
+ S+Q S E L YF LGC ++ + L+ G SY+ L + S +
Sbjct: 172 LTFYILVASKQTSSFSTESGLKYFILGCFSSGIILFGISLIYGFTGLLSYTDLTLFLSEV 231
Query: 171 SSTNSVPENEYFGVGLRGENSVVIEMDSLLRPSN--------DIYSGMMQQLTTCINALQ 222
TN + ++ +F +++ + L + + D+Y G +T ++ L
Sbjct: 232 YVTNFILDSSFFSFSGFLIGFLLLTVGFLFKLGSAPFHMWMPDVYEGSPLLITAYLSTLP 291
Query: 223 QDSL 226
+ SL
Sbjct: 292 KISL 295
>DSC3_HUMAN (Q14574) Desmocollin 3 precursor (Desmocollin 4) (HT-CP)
Length = 896
Score = 33.5 bits (75), Expect = 1.9
Identities = 34/131 (25%), Positives = 54/131 (40%), Gaps = 17/131 (12%)
Query: 2 IPVIIRIRMCLLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFL--YTLFSF 59
IP+ ++ R TK+ R V C A S S + G W +L I L LFS
Sbjct: 648 IPITVKDRAGQAATKLLR-VNLCECTHPTQCRATSRSTGVILGKWAILAILLGIALLFSV 706
Query: 60 IMILYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAAFAIMSM 119
++ L ++ ++ RF A L I++ + D+V C+A M+
Sbjct: 707 LLTLVCGVFGATKGKRFPEDLAQQNLIISNTEAPGDDRV------------CSANGFMTQ 754
Query: 120 SLSRQSQ--CG 128
+ + SQ CG
Sbjct: 755 TTNNSSQGFCG 765
>RBP1_PLAVB (Q00798) Reticulocyte binding protein 1 precursor
Length = 2873
Score = 32.7 bits (73), Expect = 3.2
Identities = 36/171 (21%), Positives = 75/171 (43%), Gaps = 13/171 (7%)
Query: 433 VPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNE--VEASSFFLKQ 490
V +++ YG ++Q+ + S + L +E + + +N +E ++ + +
Sbjct: 2395 VKTLEEIDRDYGDNYQIVEEHKKQFSILIDRTNALMDDIEIFKKENNYNLMEVNTETIHR 2454
Query: 491 MEQIMRMLQRKLIVKS-------ENCK--DRALRHIFMLNNRSHIEAMNKFSRLETIFGN 541
+ + + KL+ EN K D L++IF L S IE + + N
Sbjct: 2455 VNDYIEKITNKLVQAKTEYEQILENIKQNDDMLQNIF-LKKVSIIEYFENVKKKKESILN 2513
Query: 542 DWFQNNKA-KIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFN 591
D ++ + KI ++LD KR+ + + + E ++K LL++K + N
Sbjct: 2514 DLYEQERLLKIGEHLDEIKRNVTETLSSYEIDQKMEMMSKNLLEKKSKMMN 2564
>KINH_HUMAN (P33176) Kinesin heavy chain (Ubiquitous kinesin heavy
chain) (UKHC)
Length = 963
Score = 32.7 bits (73), Expect = 3.2
Identities = 39/186 (20%), Positives = 76/186 (39%), Gaps = 23/186 (12%)
Query: 447 HQMTIQVMSYVSSACRKRRKLEQILEEYPEVH--------NEVEASSFF---LKQMEQIM 495
H+M + ++ V +A ++ +EQ ++ + E H +EVEA + L+ Q M
Sbjct: 681 HEMEKEHLNKVQTANEVKQAVEQQIQSHRETHQKQISSLRDEVEAKAKLITDLQDQNQKM 740
Query: 496 RMLQRKLIVKSENC------KDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKA 549
+ Q +L V+ E K R L + ++ +R + ET+ +N
Sbjct: 741 MLEQERLRVEHEKLKATDQEKSRKLHELTVMQDRREQARQDLKGLEETVAKELQTLHNLR 800
Query: 550 KIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFNNRFEAICRVQSAWFIYGS 609
K L+ + V ++D++++ K+KI N E + +V +
Sbjct: 801 K------LFVQDLATRVKKSAEIDSDDTGGSAAQKQKISFLENNLEQLTKVHKQLVRDNA 854
Query: 610 QLRGEI 615
LR E+
Sbjct: 855 DLRCEL 860
>MATK_HEDHE (Q8WH60) Maturase K (Intron maturase)
Length = 505
Score = 32.3 bits (72), Expect = 4.2
Identities = 25/115 (21%), Positives = 51/115 (43%), Gaps = 10/115 (8%)
Query: 425 KMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEAS 484
K+ +FI+ + +++ Y H ++ +Q + Y L L EY + +A
Sbjct: 142 KISHFIYVL----EILIPYPVHLEILVQTLRYWVKDASSLHLLRFFLHEYCNWNTPNKAG 197
Query: 485 SFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIF 539
S F K+ +++ L + + E+ IF+ N SH+ + + + LE I+
Sbjct: 198 SSFSKRNQRLFFFLYNSHLCEYESI------FIFLRNQSSHLRSTSSGTLLERIY 246
>DUSC_HUMAN (Q9UNI6) Dual specificity protein phosphatase 12 (EC
3.1.3.48) (EC 3.1.3.16) (Dual-specificity tyrosine
phosphatase YVH1)
Length = 340
Score = 32.0 bits (71), Expect = 5.4
Identities = 20/75 (26%), Positives = 38/75 (50%), Gaps = 5/75 (6%)
Query: 452 QVMSY---VSSACRKRRKLEQILEEYPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSEN 508
Q M Y SSA K+ +L+++ E+YPE+ N + F + + L+ +++ K
Sbjct: 166 QAMGYEVDTSSAIYKQYRLQKVTEKYPELQNLPQ--ELFAVDPTTVSQGLKDEVLYKCRK 223
Query: 509 CKDRALRHIFMLNNR 523
C+ R +L++R
Sbjct: 224 CRRSLFRSSSILDHR 238
>SYC_HELPY (P41259) Cysteinyl-tRNA synthetase (EC 6.1.1.16)
(Cysteine--tRNA ligase) (CysRS)
Length = 465
Score = 31.6 bits (70), Expect = 7.1
Identities = 31/105 (29%), Positives = 51/105 (48%), Gaps = 9/105 (8%)
Query: 488 LKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMN-KFSRLETIFGNDWFQN 546
L +E ++ KL +N K++AL+ + N + E + F F ++
Sbjct: 358 LSVLESMLSSTNEKL---DQNPKNKALKGEILANLKFIEELLGIGFKDPSAYFQLGVSES 414
Query: 547 NKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFN 591
K +I+ ++ KR+ E DFLK D SI +ELLK+KI L +
Sbjct: 415 EKQEIENKIEERKRAK--ERKDFLKAD---SIREELLKQKIALMD 454
>MRAW_RICPR (Q9ZCY2) S-adenosyl-methyltransferase mraW (EC 2.1.1.-)
Length = 306
Score = 31.6 bits (70), Expect = 7.1
Identities = 18/70 (25%), Positives = 39/70 (55%), Gaps = 2/70 (2%)
Query: 477 VHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLE 536
++NE+E FL ++ I++ R ++V + +DR +++ F N+ + A +K+++ E
Sbjct: 206 INNELEELEQFLANVKNILKKDGRLVVVSFHSLEDRIVKNFFKENSEKQV-ARSKYAKDE 264
Query: 537 -TIFGNDWFQ 545
I N W +
Sbjct: 265 IKIDPNKWLK 274
>Y8K6_ENCCU (Q8SUI0) Hypothetical protein ECU08_2060
Length = 263
Score = 31.2 bits (69), Expect = 9.3
Identities = 38/165 (23%), Positives = 64/165 (38%), Gaps = 32/165 (19%)
Query: 41 HLFGNWNLLKIFLYTLFSFIMILYANIWKNSRS-----------LRFKAHTAFLVLTITS 89
HL LL FLY+ ++ +LY N WKN L T F +++I S
Sbjct: 56 HLLRLITLLLPFLYSTIQYLFLLYTN-WKNGHKQEDILYNILYYLLNLLLTTFSIISILS 114
Query: 90 VYS------------FFFDKVVNGKPDAYSLISCAAFAIMSMSLSRQSQCGIEVDLLYFF 137
+ + FFF ++ P Y L+S + + + I +D+L
Sbjct: 115 IIALIINRRENDDDLFFFSVILPSMPLTY-LLSTSCRLVPGQIGFIDTGINIFIDIL--I 171
Query: 138 LGCLIMQLM-----KIKLQLVIVGAGFSYSLIIIRSSLSSTNSVP 177
L CL+ + K L + + + + ++ + LSS S P
Sbjct: 172 LSCLVSLTLICIEPKDYLCFIAISSALTLVRLLKKKYLSSKQSPP 216
>ALLP_BACSU (P94575) Probable allantoin permease (Allantoin
transport protein)
Length = 490
Score = 31.2 bits (69), Expect = 9.3
Identities = 17/59 (28%), Positives = 29/59 (48%), Gaps = 2/59 (3%)
Query: 2 IPVIIRIRMCLLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSFI 60
IP ++R ++ + F G S A+ +L + + + G WN+L I L L SF+
Sbjct: 107 IPALLRAFTAIMWLGIQTFAG--STALNILLLNMWPGWGEIGGEWNILGIHLSGLLSFV 163
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.139 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 71,759,477
Number of Sequences: 164201
Number of extensions: 2876452
Number of successful extensions: 10308
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 10297
Number of HSP's gapped (non-prelim): 20
length of query: 672
length of database: 59,974,054
effective HSP length: 117
effective length of query: 555
effective length of database: 40,762,537
effective search space: 22623208035
effective search space used: 22623208035
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC138012.9


