

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137826.3 - phase: 0 /pseudo
(1546 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
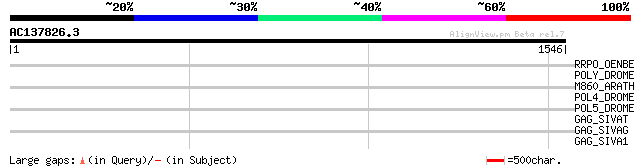
RRPO_OENBE (P31843) RNA-directed DNA polymerase homolog (Reverse... 45 0.001
POLY_DROME (P10401) Retrovirus-related Pol polyprotein from tran... 42 0.013
M860_ARATH (P92523) Hypothetical mitochondrial protein AtMg00860... 37 0.42
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 37 0.55
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 35 2.1
GAG_SIVAT (P05892) Gag polyprotein [Contains: Core protein p17; ... 33 8.0
GAG_SIVAG (P27978) Gag polyprotein [Contains: Core protein p17; ... 33 8.0
GAG_SIVA1 (P27972) Gag polyprotein [Contains: Core protein p17; ... 33 8.0
>RRPO_OENBE (P31843) RNA-directed DNA polymerase homolog (Reverse
transcriptase homolog)
Length = 142
Score = 45.4 bits (106), Expect = 0.001
Identities = 19/37 (51%), Positives = 29/37 (78%)
Query: 742 SMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASF 778
S+ MC DYR L +VT +NKY +P +DDLF++L +A++
Sbjct: 5 SLRMCIDYRALTKVTIKNKYPIPRVDDLFDRLAQATW 41
>POLY_DROME (P10401) Retrovirus-related Pol polyprotein from
transposon gypsy [Contains: Reverse transcriptase (EC
2.7.7.49); Endonuclease]
Length = 1035
Score = 42.0 bits (97), Expect = 0.013
Identities = 32/106 (30%), Positives = 54/106 (50%), Gaps = 15/106 (14%)
Query: 837 LGFILIVCVDDILLFLESK*DKVDHL-----CIV*DVHGR---QKLYA*FLSVSFC*LQL 888
+G I V VDD+++F E++ D V H+ C++ D + R +K SV +
Sbjct: 332 IGKICYVYVDDVIIFSENESDHVRHIDTVLKCLI-DANMRVSQEKTRFFKESVEY----- 385
Query: 889 PIGNVVMNERIMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGF 934
+G +V + D + ++ + + P V + R F+GLASYYR F
Sbjct: 386 -LGFIVSKDGTKSDPEKVKAIQEYPEPDCVYKVRSFLGLASYYRVF 430
>M860_ARATH (P92523) Hypothetical mitochondrial protein AtMg00860
(ORF158)
Length = 158
Score = 37.0 bits (84), Expect = 0.42
Identities = 15/43 (34%), Positives = 23/43 (52%)
Query: 892 NVVMNERIMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGF 934
+++ E + D +E ++ W P E RGF+GL YYR F
Sbjct: 37 HIISGEGVSADPAKLEAMVGWPEPKNTTELRGFLGLTGYYRRF 79
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 36.6 bits (83), Expect = 0.55
Identities = 15/31 (48%), Positives = 22/31 (70%)
Query: 748 DYRQLNRVTTRNKYTLPLIDDLFNQLWRASF 778
DYRQ+N+ +K+ LP IDD+ +QL RA +
Sbjct: 375 DYRQINKKLLADKFPLPRIDDILDQLGRAKY 405
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 34.7 bits (78), Expect = 2.1
Identities = 26/99 (26%), Positives = 45/99 (45%), Gaps = 1/99 (1%)
Query: 837 LGFILIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVM 895
+G + V +DDI++F E +L +V + L F Q+ +G +V
Sbjct: 273 IGKVCYVYIDDIIVFSEDYDTHWKNLRLVLASLSKANLQVNLEKSHFLDTQVEFLGYIVT 332
Query: 896 NERIMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGF 934
+ I D + + + P+ V E + F+G+ SYYR F
Sbjct: 333 ADGIKADPKKVRAISEMPPPTSVKELKRFLGMTSYYRKF 371
>GAG_SIVAT (P05892) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 519
Score = 32.7 bits (73), Expect = 8.0
Identities = 11/17 (64%), Positives = 13/17 (75%)
Query: 405 YNCGDFRHISRYCPKPR 421
YNCG F H+ R CP+PR
Sbjct: 400 YNCGKFGHMQRQCPEPR 416
>GAG_SIVAG (P27978) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 521
Score = 32.7 bits (73), Expect = 8.0
Identities = 11/17 (64%), Positives = 13/17 (75%)
Query: 405 YNCGDFRHISRYCPKPR 421
YNCG F H+ R CP+PR
Sbjct: 405 YNCGKFGHMQRQCPEPR 421
>GAG_SIVA1 (P27972) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 520
Score = 32.7 bits (73), Expect = 8.0
Identities = 11/17 (64%), Positives = 13/17 (75%)
Query: 405 YNCGDFRHISRYCPKPR 421
YNCG F H+ R CP+PR
Sbjct: 401 YNCGKFGHMQRQCPEPR 417
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.359 0.161 0.583
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 153,193,060
Number of Sequences: 164201
Number of extensions: 5685046
Number of successful extensions: 20195
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 20184
Number of HSP's gapped (non-prelim): 12
length of query: 1546
length of database: 59,974,054
effective HSP length: 123
effective length of query: 1423
effective length of database: 39,777,331
effective search space: 56603142013
effective search space used: 56603142013
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 73 (32.7 bits)
Medicago: description of AC137826.3


