

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137825.4 - phase: 0
(161 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
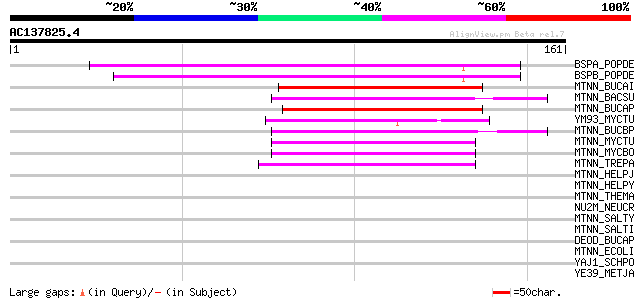
Score E
Sequences producing significant alignments: (bits) Value
BSPA_POPDE (Q07469) Bark storage protein A precursor 73 2e-13
BSPB_POPDE (Q09117) Bark storage protein B precursor 66 4e-11
MTNN_BUCAI (P57306) MTA/SAH nucleosidase (5'-methylthioadenosine... 49 5e-06
MTNN_BACSU (O32028) MTA/SAH nucleosidase (5'-methylthioadenosine... 49 6e-06
MTNN_BUCAP (O51931) MTA/SAH nucleosidase (5'-methylthioadenosine... 48 8e-06
YM93_MYCTU (Q50673) Hypothetical protein Rv2293c/MT2350 precursor 45 7e-05
MTNN_BUCBP (Q89AQ7) MTA/SAH nucleosidase (5'-methylthioadenosine... 44 1e-04
MTNN_MYCTU (P67656) MTA/SAH nucleosidase (5'-methylthioadenosine... 43 3e-04
MTNN_MYCBO (P67657) MTA/SAH nucleosidase (5'-methylthioadenosine... 43 3e-04
MTNN_TREPA (P96122) MTA/SAH nucleosidase (5'-methylthioadenosine... 43 4e-04
MTNN_HELPJ (Q9ZMY2) MTA/SAH nucleosidase (5'-methylthioadenosine... 40 0.002
MTNN_HELPY (O24915) MTA/SAH nucleosidase (5'-methylthioadenosine... 40 0.002
MTNN_THEMA (Q9X0L3) MTA/SAH nucleosidase (5'-methylthioadenosine... 38 0.011
NU2M_NEUCR (Q35140) NADH-ubiquinone oxidoreductase chain 2 (EC 1... 36 0.033
MTNN_SALTY (P60217) MTA/SAH nucleosidase (P46) (5'-methylthioade... 35 0.074
MTNN_SALTI (P60216) MTA/SAH nucleosidase (P46) (5'-methylthioade... 35 0.074
DEOD_BUCAP (Q8K937) Purine nucleoside phosphorylase (EC 2.4.2.1)... 34 0.16
MTNN_ECOLI (P24247) MTA/SAH nucleosidase (P46) (5'-methylthioade... 33 0.28
YAJ1_SCHPO (Q09901) Putative family 31 glucosidase C30D11.01c pr... 30 2.4
YE39_METJA (Q58834) Hypothetical protein MJ1439 30 3.1
>BSPA_POPDE (Q07469) Bark storage protein A precursor
Length = 312
Score = 73.2 bits (178), Expect = 2e-13
Identities = 40/126 (31%), Positives = 67/126 (52%), Gaps = 1/126 (0%)
Query: 24 INQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELE 83
I+ ++ E SN +G+V + L S + P ++ D AGR F G L
Sbjct: 11 IDPIAEIERSNCKIAHLRLGLVFTSDNNERALQDSGLYSPDSEDSSVDIAGRRFHSGTLN 70
Query: 84 KKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQI-GDVTIPKYWAHT 142
++ V +G +N Q+LL FNI GV+++G AG+L+ + + GDV++P+ A T
Sbjct: 71 GSSIVYVKTGSPSVNMATTLQILLVRFNIHGVIYFGNAGSLDKKTMVPGDVSVPQAVAFT 130
Query: 143 GLWHWQ 148
G+W+W+
Sbjct: 131 GVWNWK 136
>BSPB_POPDE (Q09117) Bark storage protein B precursor
Length = 312
Score = 65.9 bits (159), Expect = 4e-11
Identities = 37/119 (31%), Positives = 62/119 (52%), Gaps = 1/119 (0%)
Query: 31 EISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVV 90
E SN +G+V + L S + P ++ D AGR F G L ++ V
Sbjct: 18 ERSNCKIAHLRLGLVFTSDNNERALQDSGLYSPDSEDSSVDIAGRRFHSGTLNGSSIVYV 77
Query: 91 MSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQI-GDVTIPKYWAHTGLWHWQ 148
+G +N Q+LL F+I GV+++G AG+L+ + + GDV++P+ A TG+ +W+
Sbjct: 78 KTGSHSVNMATTLQILLARFSIHGVIYFGNAGSLDKKTMVPGDVSVPQAVAFTGVCNWK 136
>MTNN_BUCAI (P57306) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 232
Score = 48.9 bits (115), Expect = 5e-06
Identities = 22/59 (37%), Positives = 39/59 (65%)
Query: 79 IGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPK 137
IG+ + V ++ SG G ++A +AT +L+ L+ + +++ G AG+L S +IGD+ IPK
Sbjct: 35 IGKFKSHDVFLIKSGIGKVSASVATMILIDLYKPDTIINSGSAGSLQSFLKIGDIIIPK 93
>MTNN_BACSU (O32028) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 231
Score = 48.5 bits (114), Expect = 6e-06
Identities = 28/80 (35%), Positives = 43/80 (53%), Gaps = 5/80 (6%)
Query: 77 FRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIP 136
F GE E +VI++ SG G +NA ++T LLL + + V++ G AG + +GDV I
Sbjct: 33 FTTGEYEGTEVILLKSGIGKVNAAISTTLLLDRYKPDYVINTGSAGGFHHTLNVGDVVI- 91
Query: 137 KYWAHTGLWHWQVHTSCFNY 156
T + H V + F+Y
Sbjct: 92 ----STDVRHHDVDVTAFDY 107
>MTNN_BUCAP (O51931) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 235
Score = 48.1 bits (113), Expect = 8e-06
Identities = 20/58 (34%), Positives = 40/58 (68%)
Query: 80 GELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPK 137
G +K ++ +++SG G ++A ++T + + LF + +++ G AG+LN+ +IGD+ IPK
Sbjct: 36 GTFKKIEIFLILSGIGKVSASMSTTISINLFQPDFIINSGSAGSLNACLKIGDIIIPK 93
>YM93_MYCTU (Q50673) Hypothetical protein Rv2293c/MT2350 precursor
Length = 246
Score = 45.1 bits (105), Expect = 7e-05
Identities = 27/71 (38%), Positives = 36/71 (50%), Gaps = 7/71 (9%)
Query: 75 RHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFN------IEGVLHYGIAGNLNSRF 128
R + +G + KKVIV M+G G++NA T+ F I V+ G+AG R
Sbjct: 69 RRYYLGSISGKKVIVAMTGIGLVNATNTTETAFARFTCASSIAIAAVMFSGVAGGA-GRT 127
Query: 129 QIGDVTIPKYW 139
IGDV IP W
Sbjct: 128 SIGDVAIPARW 138
>MTNN_BUCBP (Q89AQ7) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 252
Score = 44.3 bits (103), Expect = 1e-04
Identities = 23/80 (28%), Positives = 42/80 (51%), Gaps = 5/80 (6%)
Query: 77 FRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIP 136
F IG + V++ SG G + +G+ LLL + ++ +++ G AG+LN + G + IP
Sbjct: 41 FYIGNIHNIHVVLAKSGVGKVFSGITCALLLQKYKVKFIINIGSAGSLNKNLKPGSIIIP 100
Query: 137 KYWAHTGLWHWQVHTSCFNY 156
T + + V+ + F Y
Sbjct: 101 -----TNVCYHDVNLTAFGY 115
>MTNN_MYCTU (P67656) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 255
Score = 43.1 bits (100), Expect = 3e-04
Identities = 20/59 (33%), Positives = 32/59 (53%)
Query: 77 FRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTI 135
F G+L+ +V++ +G G +N GL LL F ++ G+AG L+ IGD+ I
Sbjct: 35 FDSGQLDAHRVVLAAAGMGKVNTGLTATLLADRFGCRTIVFTGVAGGLDPELCIGDIVI 93
>MTNN_MYCBO (P67657) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 255
Score = 43.1 bits (100), Expect = 3e-04
Identities = 20/59 (33%), Positives = 32/59 (53%)
Query: 77 FRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTI 135
F G+L+ +V++ +G G +N GL LL F ++ G+AG L+ IGD+ I
Sbjct: 35 FDSGQLDAHRVVLAAAGMGKVNTGLTATLLADRFGCRTIVFTGVAGGLDPELCIGDIVI 93
>MTNN_TREPA (P96122) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 269
Score = 42.7 bits (99), Expect = 4e-04
Identities = 23/63 (36%), Positives = 36/63 (56%)
Query: 73 AGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGD 132
AG F + + +V+ V G G +NA L TQLL++ F +++ GIAG L+ R + D
Sbjct: 28 AGLTFYVVSVGALQVVYVCGGVGKVNAALCTQLLISEFGARVLINTGIAGALDERLCVFD 87
Query: 133 VTI 135
V +
Sbjct: 88 VLV 90
>MTNN_HELPJ (Q9ZMY2) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 230
Score = 40.4 bits (93), Expect = 0.002
Identities = 33/120 (27%), Positives = 52/120 (42%), Gaps = 9/120 (7%)
Query: 42 IGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVMSGEGMLNAGL 101
IGI+ E+ P+L F G F G K++IV S G +++ L
Sbjct: 4 IGILGAMREEITPILELFGV----DFEEIPLGGNVFHKGVYHNKEIIVAYSKIGKVHSTL 59
Query: 102 ATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHWQVHTSCFNYLDTFI 161
T ++ F ++ VL G+AG+L +I D+ + T L V S F++ FI
Sbjct: 60 TTTSMILAFGVQKVLFSGVAGSLVKDLKINDLLVA-----TQLVQHDVDLSAFDHPLGFI 114
>MTNN_HELPY (O24915) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 231
Score = 40.0 bits (92), Expect = 0.002
Identities = 26/94 (27%), Positives = 43/94 (45%), Gaps = 4/94 (4%)
Query: 42 IGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVMSGEGMLNAGL 101
IGI+ E+ P+L F G F G K++IV S G +++ L
Sbjct: 5 IGILGAMREEITPILELFGV----DFEEIPLGGNVFHKGVYHNKEIIVAYSKIGKVHSTL 60
Query: 102 ATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTI 135
T ++ F ++ VL G+AG+L +I D+ +
Sbjct: 61 TTTSMILAFGVQKVLFSGVAGSLVKDLKINDLLV 94
>MTNN_THEMA (Q9X0L3) MTA/SAH nucleosidase (5'-methylthioadenosine
nucleosidase) (EC 3.2.2.16) (S-adenosylhomocysteine
nucleosidase) (EC 3.2.2.9)
Length = 217
Score = 37.7 bits (86), Expect = 0.011
Identities = 22/61 (36%), Positives = 33/61 (54%), Gaps = 1/61 (1%)
Query: 75 RHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVT 134
R+++ G + + +V+V G + A L TQ L FNI+ V G AG L ++GDV
Sbjct: 30 RYYQRGVVGRNEVVVSYGFIGKVEAALVTQAFLDRFNIDAVFLTGNAGGLEG-VEVGDVV 88
Query: 135 I 135
I
Sbjct: 89 I 89
>NU2M_NEUCR (Q35140) NADH-ubiquinone oxidoreductase chain 2 (EC
1.6.5.3)
Length = 583
Score = 36.2 bits (82), Expect = 0.033
Identities = 28/114 (24%), Positives = 52/114 (45%), Gaps = 10/114 (8%)
Query: 5 LGVLVMIVLLGNSIYAYGAINQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVPH 64
L V ++++ +G S+Y Y N ++ + +INN + + + LNPLL S +
Sbjct: 381 LNVFIILITIGFSLYGY-VWNNKEYKNLLDINNSPVQLISQLKGYFYLNPLLSLSLAI-- 437
Query: 65 NKFPYFDFAGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHY 118
F FAG +G K+ V+ + N + L+ L ++ G ++Y
Sbjct: 438 ---TIFSFAGIPPLVGFFAKQMVL----SAALDNGYIFLSLIAILTSVVGAVYY 484
>MTNN_SALTY (P60217) MTA/SAH nucleosidase (P46)
(5'-methylthioadenosine nucleosidase) (EC 3.2.2.16)
(S-adenosylhomocysteine nucleosidase) (EC 3.2.2.9)
Length = 232
Score = 35.0 bits (79), Expect = 0.074
Identities = 18/56 (32%), Positives = 31/56 (55%)
Query: 80 GELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTI 135
G+L +V ++ SG G + A L LLL + +++ G AG L S ++GD+ +
Sbjct: 36 GQLNGTEVALLKSGIGKVAAALGATLLLEHCKPDVIINTGSAGGLASTLKVGDIVV 91
>MTNN_SALTI (P60216) MTA/SAH nucleosidase (P46)
(5'-methylthioadenosine nucleosidase) (EC 3.2.2.16)
(S-adenosylhomocysteine nucleosidase) (EC 3.2.2.9)
Length = 232
Score = 35.0 bits (79), Expect = 0.074
Identities = 18/56 (32%), Positives = 31/56 (55%)
Query: 80 GELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTI 135
G+L +V ++ SG G + A L LLL + +++ G AG L S ++GD+ +
Sbjct: 36 GQLNGTEVALLKSGIGKVAAALGATLLLEHCKPDVIINTGSAGGLASTLKVGDIVV 91
>DEOD_BUCAP (Q8K937) Purine nucleoside phosphorylase (EC 2.4.2.1)
(Inosine phosphorylase) (PNP)
Length = 236
Score = 33.9 bits (76), Expect = 0.16
Identities = 14/56 (25%), Positives = 31/56 (55%)
Query: 80 GELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTI 135
G + KK+ V+ G G+ +A L + L+ +N++ ++ G G + + ++ D+ I
Sbjct: 51 GYYKNKKISVMSHGMGIPSASLYVRELILEYNVKKIIRVGTCGTIQNNIKLRDIVI 106
>MTNN_ECOLI (P24247) MTA/SAH nucleosidase (P46)
(5'-methylthioadenosine nucleosidase) (EC 3.2.2.16)
(S-adenosylhomocysteine nucleosidase) (EC 3.2.2.9)
Length = 232
Score = 33.1 bits (74), Expect = 0.28
Identities = 17/56 (30%), Positives = 30/56 (53%)
Query: 80 GELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTI 135
G+L +V ++ SG G + A L LLL + +++ G AG L ++GD+ +
Sbjct: 36 GQLNGTEVALLKSGIGKVAAALGATLLLEHCKPDVIINTGSAGGLAPTLKVGDIVV 91
>YAJ1_SCHPO (Q09901) Putative family 31 glucosidase C30D11.01c
precursor (EC 3.2.1.-)
Length = 993
Score = 30.0 bits (66), Expect = 2.4
Identities = 36/151 (23%), Positives = 59/151 (38%), Gaps = 15/151 (9%)
Query: 2 EFLLGVLVMIVLLGNSIYAYGAINQLSWREISNINNEGPSIGIVVPNAYELN-------- 53
E+ GVL ++ L G++ YAYG +S E I I N +
Sbjct: 92 EYSYGVLAILELAGDACYAYGTDYPYLLLNVSYDTEERVHISISDLNQTQFQLSNRRDVW 151
Query: 54 --PLLHSSSFVPHNKFPYFDFAGRHFR--IGELEKKKVIVVMSGEGMLNAGLATQLLLTL 109
PL + SS N F F F I + +V+ G ++ +L +
Sbjct: 152 DAPLFYRSSNFSGNLQYNFSFNTDPFEFWITRIADDQVLFDTRGNPLIFEDQYIELTTNM 211
Query: 110 FNIEGVLHYGIAGNLNSRFQIGDVTIPKYWA 140
+E YG++G+ S F++G+ +WA
Sbjct: 212 --VEDYNVYGLSGSQQS-FRLGNNLTKTFWA 239
>YE39_METJA (Q58834) Hypothetical protein MJ1439
Length = 207
Score = 29.6 bits (65), Expect = 3.1
Identities = 29/123 (23%), Positives = 44/123 (35%), Gaps = 17/123 (13%)
Query: 42 IGIVVPNAYELNP-----LLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVVMSGEGM 96
+G+ P ++ N LL+ + + + +HF EL+ K VI+V E
Sbjct: 82 LGVDTPEIHKRNNPYEYYLLNGTPITDTKYLKEWGYKAKHFAEKELKNKTVIIVFDNEAP 141
Query: 97 LNAGLATQLLLTLFN--------IEGVLHYGIAGNLNSRFQIGD----VTIPKYWAHTGL 144
L N E +L YG A S F++ D V GL
Sbjct: 142 KKDKYGRYLAYIFINNSNNLINFNEELLKYGYARVYISNFELKDEFLNVEREAKENRVGL 201
Query: 145 WHW 147
W+W
Sbjct: 202 WNW 204
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.142 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,240,544
Number of Sequences: 164201
Number of extensions: 847862
Number of successful extensions: 2262
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 21
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 2238
Number of HSP's gapped (non-prelim): 29
length of query: 161
length of database: 59,974,054
effective HSP length: 101
effective length of query: 60
effective length of database: 43,389,753
effective search space: 2603385180
effective search space used: 2603385180
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC137825.4


