

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137823.16 + phase: 0
(621 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
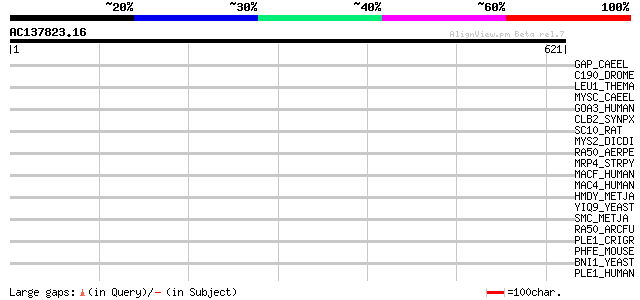
Score E
Sequences producing significant alignments: (bits) Value
GAP_CAEEL (P34288) GTPase-activating protein GAP (CeGAP) 44 0.001
C190_DROME (Q9VJE5) Restin homolog (Cytoplasmic linker protein 1... 40 0.018
LEU1_THEMA (Q9WZ23) 2-isopropylmalate synthase (EC 2.3.3.13) (Al... 39 0.053
MYSC_CAEEL (P12845) Myosin heavy chain C (MHC C) 36 0.26
GOA3_HUMAN (Q08378) Golgi autoantigen, golgin subfamily A member... 36 0.26
CLB2_SYNPX (Q7U3T3) Chaperone clpB 2 35 0.45
SC10_RAT (P97878) Exocyst complex component Sec10 (71 kDa compon... 35 0.59
MYS2_DICDI (P08799) Myosin II heavy chain, non muscle 35 0.59
RA50_AERPE (Q9YFZ1) DNA double-strand break repair rad50 ATPase 35 0.76
MRP4_STRPY (P30141) Fibrinogen- and Ig-binding protein precursor... 35 0.76
MACF_HUMAN (Q9UPN3) Microtubule-actin crosslinking factor 1, iso... 35 0.76
MAC4_HUMAN (Q96PK2) Microtubule-actin crosslinking factor 1, iso... 35 0.76
HMDY_METJA (Q58734) H(2)-forming methylenetetrahydromethanopteri... 35 0.76
YIQ9_YEAST (P40442) Hypothetical 99.7 kDa protein in SDL1 5'regi... 34 1.3
SMC_METJA (Q59037) Chromosome partition protein smc homolog 34 1.3
RA50_ARCFU (O29230) DNA double-strand break repair rad50 ATPase 34 1.3
PLE1_CRIGR (Q9JI55) Plectin 1 (PLTN) (PCN) (300-kDa intermediate... 34 1.3
PHFE_MOUSE (Q9D4H9) PHD finger protein 14 34 1.3
BNI1_YEAST (P41832) BNI1 protein (Synthetic lethal 39) 34 1.3
PLE1_HUMAN (Q15149) Plectin 1 (PLTN) (PCN) (Hemidesmosomal prote... 33 1.7
>GAP_CAEEL (P34288) GTPase-activating protein GAP (CeGAP)
Length = 1317
Score = 43.9 bits (102), Expect = 0.001
Identities = 50/167 (29%), Positives = 71/167 (41%), Gaps = 16/167 (9%)
Query: 296 KASLNTLMSEIQRLNK--------LCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTAL 347
KA+++ L + Q L+ L E +E ++KK + + R EL Y+
Sbjct: 1081 KAAIDALANHSQHLHLASSPAFEVLSEETREKIRRMQKKQSWHDTKELRSGELLKTYSPT 1140
Query: 348 LKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLY 407
K TDA S S ST LST P A + NS+ D + S R+P S
Sbjct: 1141 -KDLTDALSCTSDY-STTSSAPLSTNPPLAVACADQPNSSSDYASSDPSPCARNPSTSPA 1198
Query: 408 MLPSS---PQALLEAIGSSGSSGQEAVANAEISAAILTARAGARDPS 451
PS+ A L A SSG S Q + +I L + G+RDP+
Sbjct: 1199 SRPSNLAISPAQLHATSSSGQSHQPMSRSQKIR---LRTKLGSRDPA 1242
>C190_DROME (Q9VJE5) Restin homolog (Cytoplasmic linker protein 190)
(Microtubule binding protein 190) (d-CLIP-190)
Length = 1690
Score = 40.0 bits (92), Expect = 0.018
Identities = 63/297 (21%), Positives = 116/297 (38%), Gaps = 29/297 (9%)
Query: 206 KLGADVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRS 265
KL ++ SQ + + + Q L + AA L E+ K H +T
Sbjct: 858 KLHDEISQLKSQAEETQSELKSTQSNLEAKSKQLEAANG-SLEEEAKKSGHLLEQITKLK 916
Query: 266 IGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKK 325
+G T +L DV +K +++ A+L +++NK AE + L+
Sbjct: 917 SEVGETQ---AALSSCHTDVESKTKQLEAANAAL-------EKVNKEYAESRAEASDLQD 966
Query: 326 KWKKIE-----EFDGRRSELESIYTALLKAN----------TDAASFWSQQPSTAREYAL 370
K K+I E RS +++T L K + T A WSQ+ +E L
Sbjct: 967 KVKEITDTLHAELQAERSSSSALHTKLSKFSDEIATGHKELTSKADAWSQE-MLQKEKEL 1025
Query: 371 STIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEA 430
+ ++ K E++ +F S N + + LE + ++ ++
Sbjct: 1026 QELRQQLQDSQDSQTKLKAEGERKEKSFEESIKNLQEEVTKAKTENLELSTGTQTTIKDL 1085
Query: 431 VANAEISAAILT--ARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGLASILESLE 485
EI+ A L + + D I + + A+Q A + ++A L+++LE L+
Sbjct: 1086 QERLEITNAELQHKEKMASEDAQKIADLKTLVEAIQVANANISATNAELSTVLEVLQ 1142
>LEU1_THEMA (Q9WZ23) 2-isopropylmalate synthase (EC 2.3.3.13)
(Alpha-isopropylmalate synthase) (Alpha-IPM synthetase)
Length = 513
Score = 38.5 bits (88), Expect = 0.053
Identities = 41/213 (19%), Positives = 84/213 (39%), Gaps = 25/213 (11%)
Query: 268 IGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKW 327
IG +S+ + R K+ E G+K T ++ +L +KE D
Sbjct: 320 IGRSSETLVLGRHSGKHALRKKLETYGIKLDEETFQKVFEKFTELADRKKEVYD------ 373
Query: 328 KKIEEFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSA 387
+L SI + +L+ + T +T++P + V++ N
Sbjct: 374 ----------DDLFSIVSEVLREPINGYKLVHFHVHTG-----NTLLPTAAVVLQVGNEK 418
Query: 388 KDLIEK---EVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVANAEISAAILTAR 444
++ E V A +++ D +L + P + +++A+G+ ++ E I + + R
Sbjct: 419 REAAEAGNGPVDAIFKAIDKALGIQPKLEEYIIQAVGTGKNAQGEVKLTLRIDNELYSGR 478
Query: 445 AGARDPSAIPSICRVSAALQY-AAGGLEGSDAG 476
+ D +I ++A +Y A GL + G
Sbjct: 479 GVSTDIVEASAIAYINAINKYLIAKGLLRKNGG 511
>MYSC_CAEEL (P12845) Myosin heavy chain C (MHC C)
Length = 1947
Score = 36.2 bits (82), Expect = 0.26
Identities = 37/147 (25%), Positives = 68/147 (46%), Gaps = 15/147 (10%)
Query: 215 GSQNQLLERQKAHVQQFLATEDALNNAA------EARDLCEKLMKRLHGGTDVTSRSIGI 268
GS ++ ER A +Q +A E L +A+ EAR + K+L V + +
Sbjct: 912 GSTREVEERMTAMNEQKVALEGKLADASKKLEVEEARAVEINKQKKL-----VEAECADL 966
Query: 269 GATSQNVG-SLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKW 327
Q+V SLR+++ + AKE ++ L+ + I +LNK ERK E+ KK
Sbjct: 967 KKNCQDVDLSLRKVEAEKNAKEHQIRALQDEMRQQDENISKLNK---ERKNQEEQNKKLT 1023
Query: 328 KKIEEFDGRRSELESIYTALLKANTDA 354
+ ++ + + + L+++ D+
Sbjct: 1024 EDLQAAEEQNLAANKLKAKLMQSLEDS 1050
>GOA3_HUMAN (Q08378) Golgi autoantigen, golgin subfamily A member 3
(Golgin-160) (Golgi complex-associated protein of 170
kDa) (GCP170)
Length = 1498
Score = 36.2 bits (82), Expect = 0.26
Identities = 43/177 (24%), Positives = 76/177 (42%), Gaps = 17/177 (9%)
Query: 221 LERQKAHVQQFLAT-EDALNNAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLR 279
L R+ A V+Q +A E L +A + RD E ++ L + + T N +
Sbjct: 904 LHREVAQVRQHMADLEGHLQSAQKERDEMETHLQSLQFDKEQM-----VAVTEANEALKK 958
Query: 280 QLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSE 339
Q++ E + KA + + Q++ +L ++ A+ +K K K E G S
Sbjct: 959 QIE------ELQQEARKA----ITEQKQKMRRLGSDLTSAQKEMKTKHKAYENAVGILSR 1008
Query: 340 -LESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEV 395
L+ A A+ + +Q S+ AL I A A ++ + +K L+EKE+
Sbjct: 1009 RLQEALAAKEAADAELGQLRAQGGSSDSSLALHERIQALEAELQAVSHSKTLLEKEL 1065
>CLB2_SYNPX (Q7U3T3) Chaperone clpB 2
Length = 900
Score = 35.4 bits (80), Expect = 0.45
Identities = 88/378 (23%), Positives = 155/378 (40%), Gaps = 63/378 (16%)
Query: 125 GQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYKYENNI 184
G++PD ++ + + L L+ A + + ++ +E++ R+D+ ++ + E +
Sbjct: 264 GEVPDSLQGLRLIALDLGALIAGA--KFRGQFEERLRSVLEEVS-RSDSGVVLFIDELHT 320
Query: 185 VM--DVSSSDGSSPLQYPLYGNGKL---GADVPPGGSQNQLLERQKA---HVQQFLATED 236
V+ D SS+D S L+ P G L GA P + +E+ A QQ L E
Sbjct: 321 VVGSDRSSTDAGSLLK-PALARGDLRCIGATTPE--EYRRTVEKDPALNRRFQQVLIREP 377
Query: 237 ALNNAAEA-RDLCEKLMKRLHGGTDVTSRSIGIG----------------ATSQNVGSLR 279
L + E R L E+ LH G +T +I A +
Sbjct: 378 DLELSLEILRGLRERY--ELHHGVTITDEAIQTANRLADRYISDRCLPDKAIDLIDEAAA 435
Query: 280 QLQLDVWAKEREVSGLKASLNTL------MSEIQRLNKLCAERKEAE-----DSLKKKWK 328
QL+++V +K + V +A L + E ++ +R+ E D L+++W+
Sbjct: 436 QLKIEVTSKPQVVEEAEADLRRVELALLAAEEAPEEERIQLQRQRLEVSSRLDDLRRRWQ 495
Query: 329 K----IEEFDGRRSELESIYTALLKANTDAASFWS-----------QQPSTAREYALSTI 373
+ +EE + E + A+ +A + A + QQ A E + +
Sbjct: 496 EERTQLEELGQLLQQDEDLRHAIAEAEREGALEEAARLQYDQLHTVQQRREALEASQAEA 555
Query: 374 IPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQ-EAVA 432
A +A++ A D+ + V+ + P L LE+ S GQ EAVA
Sbjct: 556 QSAGTALLREQVEAGDIADL-VARWTGIPVQRLLAGERRKLLALESHLSERVIGQVEAVA 614
Query: 433 NAEISAAILTARAGARDP 450
++AAI ARAG +DP
Sbjct: 615 --AVAAAIRRARAGMKDP 630
>SC10_RAT (P97878) Exocyst complex component Sec10 (71 kDa component
of rsec6/8 secretory complex) (p71)
Length = 708
Score = 35.0 bits (79), Expect = 0.59
Identities = 34/138 (24%), Positives = 55/138 (39%), Gaps = 15/138 (10%)
Query: 5 YDRQCDEASRIFAEYHKRLCYYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYSTVKGS 64
+DR + S+I ++YH C I + AQR G S E AV KG
Sbjct: 175 FDRFSEVKSKIASKYHDLECQLIQEFTSAQRRG---------EVSRMREVAAVLLHFKGY 225
Query: 65 KSSDDVIVIETTREKNIRKACESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLGFD-F 123
DV + + +R A + ++ + + SNP+A AKL + F
Sbjct: 226 SHCIDVYIKQCQEGAYLRNDIFEDAAILCQRVNK-----QVGDIFSNPEAVLAKLIQNVF 280
Query: 124 DGQIPDEVRTVIVNCLKS 141
+ ++ V+ + C KS
Sbjct: 281 EVKLQSFVKDQLEECRKS 298
>MYS2_DICDI (P08799) Myosin II heavy chain, non muscle
Length = 2116
Score = 35.0 bits (79), Expect = 0.59
Identities = 45/194 (23%), Positives = 78/194 (40%), Gaps = 41/194 (21%)
Query: 217 QNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVG 276
Q + LE + VQ+ LA E A A EA + K+L G + T++
Sbjct: 1032 QKKKLEEELKQVQEALAAETAAKLAQEAAN------KKLQGEYTELNEKFNSEVTAR--- 1082
Query: 277 SLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKI------ 330
S ++ S TL S++ +N E K+ D+L+KK K +
Sbjct: 1083 ----------------SNVEKSKKTLESQLVAVNNELDEEKKNRDALEKKKKALDAMLEE 1126
Query: 331 --EEFDGRRSELESIYTALLKANTDAASFWSQ----QPSTAR-EYALSTI---IPACSAV 380
++ + E +S+Y +K +D + +Q Q + A+ E ST+ +
Sbjct: 1127 MKDQLESTGGEKKSLYDLKVKQESDMEALRNQISELQSTIAKLEKIKSTLEGEVARLQGE 1186
Query: 381 VETSNSAKDLIEKE 394
+E AK +EK+
Sbjct: 1187 LEAEQLAKSNVEKQ 1200
>RA50_AERPE (Q9YFZ1) DNA double-strand break repair rad50 ATPase
Length = 919
Score = 34.7 bits (78), Expect = 0.76
Identities = 31/126 (24%), Positives = 53/126 (41%), Gaps = 6/126 (4%)
Query: 216 SQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNV 275
S+++ LER + LAT + DL EK + L G S A + +
Sbjct: 611 SESKKLERMLVSKAEDLATRLGITAYRSLDDLLEKAREALEGVDKELS------AIERRL 664
Query: 276 GSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDG 335
R+L+ + + E + L L +E ++L K + + E E LK+ + E D
Sbjct: 665 EEARRLKEEAAKLKWEAEQVMKRLEELEAEEKKLRKEVSRKSEIEARLKEVQNTLAELDD 724
Query: 336 RRSELE 341
R S ++
Sbjct: 725 RISRID 730
>MRP4_STRPY (P30141) Fibrinogen- and Ig-binding protein precursor
(MRP protein)
Length = 388
Score = 34.7 bits (78), Expect = 0.76
Identities = 57/271 (21%), Positives = 111/271 (40%), Gaps = 30/271 (11%)
Query: 214 GGSQNQLLERQKA-------------HVQQFLATED----ALNNAAEARD--LCEKLMKR 254
G N++ E +KA H+ Q ++ + AL + AE ++ E L +
Sbjct: 56 GKEANKVFEERKALEKQARDLGDTINHMSQTISEQSRKIAALKSEAELKNQQALEALNNK 115
Query: 255 LHGGTDVTSRSIGIGATSQN-VGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLC 313
+D+T+ + + + V +++ ++ AK++E++ K+ L +EI+ L +
Sbjct: 116 NKQISDLTNENAQLKEAIEGYVQTIQNASREIAAKQQELAAAKSQLEAKNAEIEALKQQD 175
Query: 314 AERKEAEDSLKKKWKKIEEFDG-RRSELESIYTALLKANTDAASFWSQQP-------STA 365
A + E L+ + +E G + EL + L A + A SQ S
Sbjct: 176 ASKTEEIAKLQSEAATLENLLGSAKRELTELQAKLDTATAEKAKLESQVTTLENLLGSAK 235
Query: 366 REYA-LSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSG 424
RE L + A +A E S +EK++ A + + L ++ Q + +
Sbjct: 236 RELTDLQAKLDAANAEKEKLQSQAATLEKQLEATKKELADLQAKLAATNQEKEKLEAEAK 295
Query: 425 SSGQEAVANAEISAAILTARA-GARDPSAIP 454
+ ++ AE A + +A GA+ P P
Sbjct: 296 ALKEQLAKQAEELAKLKADKASGAQKPDTKP 326
>MACF_HUMAN (Q9UPN3) Microtubule-actin crosslinking factor 1, isoforms
1/2/3 (Actin cross-linking family protein 7) (Macrophin
1) (Trabeculin-alpha) (620 kDa actin-binding protein)
(ABP620)
Length = 5430
Score = 34.7 bits (78), Expect = 0.76
Identities = 40/178 (22%), Positives = 68/178 (37%), Gaps = 43/178 (24%)
Query: 210 DVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHG------------ 257
D P G +QLL + + Q+FL + +N+ C + +L G
Sbjct: 3017 DAPDGSDASQLLHQAEVAQQEFLEVKQRVNSG------CVMMENKLEGIGQFHCRVREMF 3070
Query: 258 --GTDVTSRSIGIGATSQNVGSLRQ--------------LQLDVWAKEREVSGLKASLNT 301
D+ G+GA ++ SL+ L+LD+ A E E + T
Sbjct: 3071 SQLADLDDELDGMGAIGRDTDSLQSQIEDVRLFLNKIHVLKLDIEASEAECRHMLEEEGT 3130
Query: 302 -----LMSEIQRLNKLCAE----RKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKA 350
L E++ LNK C + K ++ L+ ++E+F + L TA +A
Sbjct: 3131 LDLLGLKRELEALNKQCGKLTERGKARQEQLELTLGRVEDFYRKLKGLNDATTAAEEA 3188
>MAC4_HUMAN (Q96PK2) Microtubule-actin crosslinking factor 1, isoform
4
Length = 5938
Score = 34.7 bits (78), Expect = 0.76
Identities = 40/178 (22%), Positives = 68/178 (37%), Gaps = 43/178 (24%)
Query: 210 DVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHG------------ 257
D P G +QLL + + Q+FL + +N+ C + +L G
Sbjct: 3519 DAPDGSDASQLLHQAEVAQQEFLEVKQRVNSG------CVMMENKLEGIGQFHCRVREMF 3572
Query: 258 --GTDVTSRSIGIGATSQNVGSLRQ--------------LQLDVWAKEREVSGLKASLNT 301
D+ G+GA ++ SL+ L+LD+ A E E + T
Sbjct: 3573 SQLADLDDELDGMGAIGRDTDSLQSQIEDVRLFLNKIHVLKLDIEASEAECRHMLEEEGT 3632
Query: 302 -----LMSEIQRLNKLCAE----RKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKA 350
L E++ LNK C + K ++ L+ ++E+F + L TA +A
Sbjct: 3633 LDLLGLKRELEALNKQCGKLTERGKARQEQLELTLGRVEDFYRKLKGLNDATTAAEEA 3690
>HMDY_METJA (Q58734) H(2)-forming methylenetetrahydromethanopterin
dehydrogenase related protein MJ1338
Length = 353
Score = 34.7 bits (78), Expect = 0.76
Identities = 43/177 (24%), Positives = 70/177 (39%), Gaps = 25/177 (14%)
Query: 409 LPSSPQALLEAIGSSGSSGQEAVANAEISAAILTARAGARDPSAIP-------------- 454
+P +PQ IG + G+E +I A+ A++ ++ +P
Sbjct: 159 VPGTPQHGHYVIGGKTTDGKELATEEQIKKAVELAKSAGKEAYVVPADVSSVVADMGSLV 218
Query: 455 SICRVSAALQYAAGGLEGSDAGLASILESLEFCLKRRGS--EASVLEDLLKAINLVHIRR 512
+ +S L Y G + +A I + + L+ S E S +E ++KA+N + R
Sbjct: 219 TAVALSGVLDYYTVGRKIINAPKKMIEQQVIMTLQTMASLVETSGIEGMVKALNPELLIR 278
Query: 513 DLVQSGHALLNHAYFVQQDYERTTNFSLNLAAEQERAVMEKWLPELKTGVLNAQQSL 569
S LL+ + E N L AE E+A E+K L A QSL
Sbjct: 279 S--ASSMKLLDRQKDLDAALEILQNLDETLKAEVEKA-------EIKPTTLVAAQSL 326
>YIQ9_YEAST (P40442) Hypothetical 99.7 kDa protein in SDL1 5'region
precursor
Length = 995
Score = 33.9 bits (76), Expect = 1.3
Identities = 29/148 (19%), Positives = 69/148 (46%), Gaps = 10/148 (6%)
Query: 345 TALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDN 404
+AL + +++ S S ST+ + S + S++ ETS+SA D++ + +
Sbjct: 20 SALGQYYSNSTSISSNSSSTSVVSSSSGSVSISSSIAETSSSATDILSSITQSASSTSGV 79
Query: 405 SLYMLPSSPQALLEAIGSSGSS--------GQEAVANAEISAAILTARAGARDPSAIPSI 456
S + PSS + ++ S SS Q + + +++S+++ + + D S+ S+
Sbjct: 80 SSSVGPSSSSVVSSSVSQSSSSVSDVSSSVSQSSSSASDVSSSVSQSASSTSDVSS--SV 137
Query: 457 CRVSAALQYAAGGLEGSDAGLASILESL 484
+ S++ + + S + + + S+
Sbjct: 138 SQSSSSASDVSSSVSQSSSSASDVSSSV 165
>SMC_METJA (Q59037) Chromosome partition protein smc homolog
Length = 1169
Score = 33.9 bits (76), Expect = 1.3
Identities = 23/70 (32%), Positives = 37/70 (52%), Gaps = 1/70 (1%)
Query: 302 LMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWS-Q 360
++ EI + + ++K+AE+ LKK + IE D R SE+E+ L K DA +
Sbjct: 164 IIDEISGIAEFDEKKKKAEEELKKARELIEMIDIRISEVENNLKKLKKEKEDAEKYIKLN 223
Query: 361 QPSTAREYAL 370
+ A +YAL
Sbjct: 224 EELKAAKYAL 233
>RA50_ARCFU (O29230) DNA double-strand break repair rad50 ATPase
Length = 886
Score = 33.9 bits (76), Expect = 1.3
Identities = 36/188 (19%), Positives = 78/188 (41%), Gaps = 5/188 (2%)
Query: 215 GSQNQLLERQKAHVQQFLATEDALNNAAEARDL-CEKLMKRLHGGTDVTSR-SIGIGATS 272
G+ ++LER+K +++FL+ E+ + E + E++ + + + + S +
Sbjct: 162 GAVIRMLEREKERLKEFLSQEEQIKRQKEEKKAEIERISEEIKSIESLREKLSEEVRNLE 221
Query: 273 QNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEE 332
+ L + + + + ++ S + + L +++ L K E E + L+KK K+++E
Sbjct: 222 SRLKELEEHKSRLESLRKQESSVLQEVRGLEEKLRELEKQLKEVVERIEDLEKKAKEVKE 281
Query: 333 FDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIE 392
+ + L + N ++ RE A I A E NS + I
Sbjct: 282 LKPKAERYSILEKLLSEINQALRDVEKREGDLTREAA---GIQAQLKKAEEDNSKLEEIT 338
Query: 393 KEVSAFYR 400
K + R
Sbjct: 339 KRIEELER 346
>PLE1_CRIGR (Q9JI55) Plectin 1 (PLTN) (PCN) (300-kDa intermediate
filament-associated protein) (IFAP300) (Fragment)
Length = 4473
Score = 33.9 bits (76), Expect = 1.3
Identities = 31/127 (24%), Positives = 54/127 (42%), Gaps = 10/127 (7%)
Query: 277 SLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAER-------KEAEDSLKKKWKK 329
SL++L+ A++ L+ L +RL + ER +E + L ++W+
Sbjct: 1057 SLKKLRAQAEAQQPVFDTLRDELRGAQEVGERLQQRHGERDVEVERWRERVNQLLERWQA 1116
Query: 330 I-EEFDGRRSELESIYTAL--LKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNS 386
+ + D R+ ELE + L + + D S W Q +E + IP A E
Sbjct: 1117 VLAQIDVRQRELEQLGRQLRYYRESADPLSSWLQDAKRRQEQIQAVPIPNSQAAREQLRQ 1176
Query: 387 AKDLIEK 393
K L+E+
Sbjct: 1177 EKALLEE 1183
>PHFE_MOUSE (Q9D4H9) PHD finger protein 14
Length = 881
Score = 33.9 bits (76), Expect = 1.3
Identities = 31/120 (25%), Positives = 59/120 (48%), Gaps = 7/120 (5%)
Query: 258 GTDVTSRSIGIGATSQNVGSLRQLQL--DVWAKEREVSGLKASLNTLMSEIQRLNKLCAE 315
G++ S+ G G+ S + ++ + +L D+ KE + LK S +M ++L K+ E
Sbjct: 45 GSEDPSKDSGEGSCSDSEENILEEELNEDIQVKEEQ---LKNSTEEIMPSDKQLIKM--E 99
Query: 316 RKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIP 375
+KE E++ ++ KK E+ + E E +++ AAS P+T+ S +P
Sbjct: 100 KKEEEENGERPRKKKEKEKEKEKEREKDKEKATVSDSAAASAAGTTPATSPPAVTSPSVP 159
>BNI1_YEAST (P41832) BNI1 protein (Synthetic lethal 39)
Length = 1953
Score = 33.9 bits (76), Expect = 1.3
Identities = 48/235 (20%), Positives = 102/235 (42%), Gaps = 30/235 (12%)
Query: 202 YGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDV 261
Y NGKL DVP + ++H Q + N++ + L + + H + +
Sbjct: 99 YMNGKLSGDVPVSS------QHARSHSMQSKYSYSKRNSSQASNKLTRQHTGQSHSASSL 152
Query: 262 TSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEA-- 319
S+ + S+ ++ L++ + EV L +M + L +++EA
Sbjct: 153 LSQG-SLTNLSKFTTPDGKIYLEMPSDPYEVEVL---FEDIMYKRNIFQSLSEDKQEALM 208
Query: 320 EDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSA 379
S++KKW +++ ++EL+ ++ANT ++S S+ + + + T + S
Sbjct: 209 GYSIEKKWLIVKQ--DLQNELKK-----MRANTTSSSTASRTSMASDHHPILTANSSLS- 260
Query: 380 VVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVANA 434
S K ++ S SP +++Y + L ++G+S S G++ V+ +
Sbjct: 261 ------SPKSVLMTSAS----SPTSTVYSNSLNHSTTLSSVGTSTSKGKKLVSGS 305
>PLE1_HUMAN (Q15149) Plectin 1 (PLTN) (PCN) (Hemidesmosomal protein 1)
(HD1)
Length = 4684
Score = 33.5 bits (75), Expect = 1.7
Identities = 60/287 (20%), Positives = 107/287 (36%), Gaps = 21/287 (7%)
Query: 223 RQKAHVQQ-FLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQL 281
RQ+ V++ LA + + AA + E + R+ + T RS + + RQ
Sbjct: 2055 RQRRQVEEEILALKASFEKAAAGKAELELELGRIRSNAEDTLRS----KEQAELEAARQR 2110
Query: 282 QLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELE 341
QL + R + +L +E + +RK A + +++ K+EE R E
Sbjct: 2111 QLAAEEERRRREAEERVQKSLAAE----EEAARQRKAALEEVERLKAKVEEARSLRERAE 2166
Query: 342 SIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRS 401
L+ +AA Q A +A+ ++ S D + E A R+
Sbjct: 2167 QESARQLQLAQEAAQKRLQAEEKAHAFAVQQKEQELQQTLQQEQSVLDRLRGEAEAARRA 2226
Query: 402 PDNSLYM-------LPSSPQALLEAIGSSGSSGQEAVANAEISAAILTARAGARDPSAIP 454
+ + S + + EA S+ ++A A A+ AA R A +A
Sbjct: 2227 AEEAEEARVQAEREAAQSRRQVEEAERLKQSAEEQAQARAQAQAAAEKLRKEAEQEAA-- 2284
Query: 455 SICRVSAALQYAAGGLEGSDAGLASILESLEFCLKRRGSEASVLEDL 501
R + A Q A + +DA + + E L+++ L L
Sbjct: 2285 ---RRAQAEQAALRQKQAADAEMEKHKKFAEQTLRQKAQVEQELTTL 2328
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,408,677
Number of Sequences: 164201
Number of extensions: 2711451
Number of successful extensions: 7792
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 56
Number of HSP's that attempted gapping in prelim test: 7727
Number of HSP's gapped (non-prelim): 128
length of query: 621
length of database: 59,974,054
effective HSP length: 116
effective length of query: 505
effective length of database: 40,926,738
effective search space: 20668002690
effective search space used: 20668002690
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 69 (31.2 bits)
Medicago: description of AC137823.16


