

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137822.15 + phase: 0
(279 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
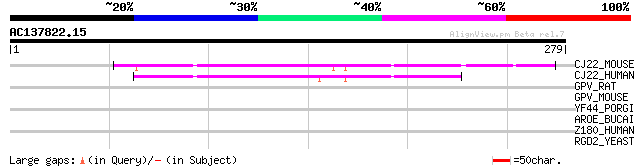
CJ22_MOUSE (Q6PDY2) Protein C10orf22 homolog 91 2e-18
CJ22_HUMAN (Q96SZ5) Protein C10orf22 87 4e-17
GPV_RAT (O08770) Platelet glycoprotein V precursor (GPV) (CD42D) 37 0.041
GPV_MOUSE (O08742) Platelet glycoprotein V precursor (GPV) (CD42D) 33 0.59
YF44_PORGI (Q7MUH3) Hypothetical UPF0246 protein PG1544 32 2.2
AROE_BUCAI (Q44607) Shikimate dehydrogenase (EC 1.1.1.25) 31 3.8
Z180_HUMAN (Q9UJW8) Zinc finger protein 180 (HHZ168) 30 8.5
RGD2_YEAST (P43556) Rho-GTPase-activating protein RGD2 30 8.5
>CJ22_MOUSE (Q6PDY2) Protein C10orf22 homolog
Length = 251
Score = 91.3 bits (225), Expect = 2e-18
Identities = 72/241 (29%), Positives = 108/241 (43%), Gaps = 25/241 (10%)
Query: 53 TFKGPGTVPS------PRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTT 106
TF+G T P ++ L +L ++ ED+ ++ +P + + VTY
Sbjct: 15 TFRGSSTGSEGPAPGFPENLSLLKSLLTQVRAEDLNIAPRKALPQP--LPRNLPPVTYMH 72
Query: 107 VYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNL----- 161
+Y+ + FSL +F L IPLH+HPGM K+L G + I D +D A H
Sbjct: 73 IYETEGFSLGVFLLKSGTCIPLHDHPGMHGMLKVLYGTVRISCMDKLDTGAGHRRPPPEQ 132
Query: 162 -----LQPSSK--LRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAIT-PCAVLDVIGP 213
LQP + +R L++ +T VL P N+H+ A+ P A LD++ P
Sbjct: 133 QFEPPLQPLEREAVRPGVLRSRAEYTEASGPCVLTPHR-DNLHQIDAVDGPAAFLDILAP 191
Query: 214 PYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEEIDMPENCQMDGIEYLGPP 273
PY EDGRDC YY+ +E G D LL E ++ +G Y GP
Sbjct: 192 PYDPEDGRDCHYYR--VVEPIRPKEASGSACDLPREVWLL-ETPQADDFWCEGEPYPGPK 248
Query: 274 I 274
+
Sbjct: 249 V 249
>CJ22_HUMAN (Q96SZ5) Protein C10orf22
Length = 228
Score = 87.0 bits (214), Expect = 4e-17
Identities = 56/180 (31%), Positives = 87/180 (48%), Gaps = 18/180 (10%)
Query: 63 PRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFLPE 122
P ++ KL +L ++ ED+ ++ +P + VTY +Y+ D FSL +F L
Sbjct: 6 PENLSKLKSLLTQLRAEDLNIAPRKATLQP--LPPNLPPVTYMHIYETDGFSLGVFLLKS 63
Query: 123 RGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVD------------HEASHNLLQPSSK--L 168
IPLH+HPGM K+L G + I D +D + LQP + +
Sbjct: 64 GTSIPLHDHPGMHGMLKVLYGTVRISCMDKLDAGGGQRPRALPPEQQFEPPLQPREREAV 123
Query: 169 RLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAIT-PCAVLDVIGPPYSKEDGRDCSYYK 227
R L++ +T +L P N+H+ A+ P A LD++ PPY +DGRDC YY+
Sbjct: 124 RPGVLRSRAEYTEASGPCILTPHR-DNLHQIDAVEGPAAFLDILAPPYDPDDGRDCHYYR 182
>GPV_RAT (O08770) Platelet glycoprotein V precursor (GPV) (CD42D)
Length = 567
Score = 37.4 bits (85), Expect = 0.041
Identities = 25/77 (32%), Positives = 36/77 (46%), Gaps = 4/77 (5%)
Query: 112 NFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNLLQPSSKLRLA 171
N + + F +RGV+ H+ GMTV +L+L HI + D + N L LRL
Sbjct: 51 NLTHILLFRMDRGVLQSHSFSGMTVLQRLMLSDSHISAID----PGTFNDLVKLKTLRLT 106
Query: 172 KLKANKTFTAPCDTSVL 188
+ K + A D VL
Sbjct: 107 RNKISHLPRAILDKMVL 123
>GPV_MOUSE (O08742) Platelet glycoprotein V precursor (GPV) (CD42D)
Length = 567
Score = 33.5 bits (75), Expect = 0.59
Identities = 22/77 (28%), Positives = 36/77 (46%), Gaps = 4/77 (5%)
Query: 112 NFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNLLQPSSKLRLA 171
N + + F ++G++ H+ GMTV + +L HI + D + N L LRL
Sbjct: 51 NLTHILLFRMDQGILRNHSFSGMTVLQRQMLSDSHISAID----PGTFNDLVKLKTLRLT 106
Query: 172 KLKANKTFTAPCDTSVL 188
+ K ++ A D VL
Sbjct: 107 RNKISRLPRAILDKMVL 123
>YF44_PORGI (Q7MUH3) Hypothetical UPF0246 protein PG1544
Length = 256
Score = 31.6 bits (70), Expect = 2.2
Identities = 13/30 (43%), Positives = 19/30 (63%)
Query: 250 YGLLEEIDMPENCQMDGIEYLGPPIDDTMF 279
YGLL DM +M+G +LG P++D +F
Sbjct: 114 YGLLRPSDMIRPYRMEGFAHLGAPVEDEVF 143
>AROE_BUCAI (Q44607) Shikimate dehydrogenase (EC 1.1.1.25)
Length = 273
Score = 30.8 bits (68), Expect = 3.8
Identities = 34/135 (25%), Positives = 55/135 (40%), Gaps = 29/135 (21%)
Query: 44 QELFDSCKQTFKGPGTVPSPRDVHKL--CHILDNMKPEDVGLSRDLQFFKPGNIIKENQR 101
QE + C + K S + K+ C+IL + + +GL DL N IK+N
Sbjct: 71 QEAYFFCNKLTKRAEVAQSVNTLKKIDNCNILGD-NTDGIGLLSDLIRL---NFIKKNYS 126
Query: 102 VTYTTV-------------YKCDNFSLCIF----FLPERGVIPLHNHPGMTVFSKLLLGQ 144
+ Y C S+CIF E+ V+ H + + VF+
Sbjct: 127 ILIIGAGGAARGVLFPLLSYGC---SICIFNRTVLNAEKLVLQFHKYGNINVFNT---NS 180
Query: 145 MHIKSYDWVDHEASH 159
+H+KS+D + + SH
Sbjct: 181 LHVKSFDLIINATSH 195
>Z180_HUMAN (Q9UJW8) Zinc finger protein 180 (HHZ168)
Length = 692
Score = 29.6 bits (65), Expect = 8.5
Identities = 18/71 (25%), Positives = 34/71 (47%), Gaps = 2/71 (2%)
Query: 90 FKPGNIIKENQRV-TYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIK 148
F+ + + ++QR T ++C+ F L R ++ H G F+ + G+ I
Sbjct: 614 FRQSSCLTQHQRTHTGEKPFECNQCGKT-FSLSARLIVHQRTHTGEKPFTCIQCGKAFIN 672
Query: 149 SYDWVDHEASH 159
SY + H+A+H
Sbjct: 673 SYKLIRHQATH 683
>RGD2_YEAST (P43556) Rho-GTPase-activating protein RGD2
Length = 714
Score = 29.6 bits (65), Expect = 8.5
Identities = 22/84 (26%), Positives = 39/84 (46%), Gaps = 5/84 (5%)
Query: 9 VGHVNKVCYVKRVIAKKRNKPYNHRRVKKHVPKALQELFDSCKQTFKGPGTVPSPRDVHK 68
+ H ++ Y ++ + N N R+ KK+V K +E F+ C+Q + T + ++
Sbjct: 113 IEHRERIQYSEKTLLTNVN---NFRKSKKYVGKLEKEYFNKCRQLEEFKRTHFNEDELAN 169
Query: 69 LCHIL--DNMKPEDVGLSRDLQFF 90
L N EDV +D +FF
Sbjct: 170 AMKSLKIQNKYEEDVAREKDHRFF 193
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.140 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,372,395
Number of Sequences: 164201
Number of extensions: 1772535
Number of successful extensions: 3072
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 3065
Number of HSP's gapped (non-prelim): 8
length of query: 279
length of database: 59,974,054
effective HSP length: 109
effective length of query: 170
effective length of database: 42,076,145
effective search space: 7152944650
effective search space used: 7152944650
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC137822.15


