

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137670.5 + phase: 0
(409 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
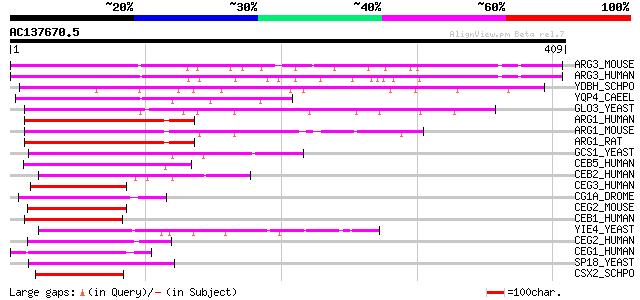
Score E
Sequences producing significant alignments: (bits) Value
ARG3_MOUSE (Q9D8S3) ADP-ribosylation factor GTPase-activating pr... 181 4e-45
ARG3_HUMAN (Q9NP61) ADP-ribosylation factor GTPase-activating pr... 179 1e-44
YDBH_SCHPO (Q10367) Hypothetical protein C22E12.17c in chromosome I 154 4e-37
YQP4_CAEEL (Q09531) Hypothetical protein F07F6.4 in chromosome III 139 1e-32
GLO3_YEAST (P38682) ADP-ribosylation factor GTPase-activating pr... 137 4e-32
ARG1_HUMAN (Q8N6T3) ADP-ribosylation factor GTPase activating pr... 109 1e-23
ARG1_MOUSE (Q9EPJ9) ADP-ribosylation factor GTPase activating pr... 107 6e-23
ARG1_RAT (Q62848) ADP-ribosylation factor GTPase activating prot... 107 7e-23
GCS1_YEAST (P35197) ADP-ribosylation factor GTPase-activating pr... 99 1e-20
CEB5_HUMAN (Q96P50) Centaurin beta 5 (Cnt-b5) 90 9e-18
CEB2_HUMAN (Q15057) Centaurin beta 2 (Cnt-b2) 84 7e-16
CEG3_HUMAN (Q96P47) Centaurin gamma 3 82 3e-15
CG1A_DROME (Q9NGC3) Centaurin gamma 1A protein 80 7e-15
CEG2_MOUSE (Q8BXK8) Centaurin gamma 2 (ARF-GAP with GTP-binding ... 80 1e-14
CEB1_HUMAN (Q15027) Centaurin beta 1 (Cnt-b1) 80 1e-14
YIE4_YEAST (P40529) Hypothetical 32.6 kDa protein in MET30-PIG2 ... 79 2e-14
CEG2_HUMAN (Q9UPQ3) Centaurin gamma 2 (ARF-GAP with GTP-binding ... 79 2e-14
CEG1_HUMAN (Q99490) Centaurin gamma 1 79 3e-14
SP18_YEAST (P32572) Sporulation protein SPS18 (Sporulation-speci... 70 7e-12
CSX2_SCHPO (Q9UUE2) Protein csx2 67 6e-11
>ARG3_MOUSE (Q9D8S3) ADP-ribosylation factor GTPase-activating
protein 3 (ARF GAP 3)
Length = 525
Score = 181 bits (458), Expect = 4e-45
Identities = 164/531 (30%), Positives = 233/531 (42%), Gaps = 133/531 (25%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
M S D A+F++L++ NK CFDC AKNP+WAS++YG+FLCIDCS HRSLGVH+S
Sbjct: 1 MGDPSKQDILAIFKRLRSVPTNKVCFDCGAKNPSWASISYGVFLCIDCSGSHRSLGVHLS 60
Query: 61 FVRSTNLDS-WTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSK 119
F+RST LDS W+ QL+ M GGNS A FF QHG AKY SRAA+LY++ +
Sbjct: 61 FIRSTELDSNWSWFQLRCMQVGGNSNASSFFHQHGC-ATKDTNAKYNSRAAQLYREKIKT 119
Query: 120 EVAKSMSEEA------ALSAPPAA---------SSQSAQGTNGLPDVKTNEVPIEKT--- 161
++ + +APP + +S ++ +G T
Sbjct: 120 LATQATRRHGTDLWLDSCAAPPVSPPPKEEDFFASHASLEVSGAMQASAQPESASSTPWG 179
Query: 162 VEKTVEKPE------------KTESSSSPRAYTAV----SNNLKKPIGAKKTGKSGGLGA 205
+E T EK E T ++P +++ N KK +GAKK G LGA
Sbjct: 180 LETTPEKHEGGPGQGPSVEGLNTPGKAAPAEVSSIIKKKPNQAKKGLGAKK----GSLGA 235
Query: 206 RKLT----------------RKPSESLYEQKPEELPAPVSSSTITKNNL----------- 238
+KLT RK E L P+E + VSS + +L
Sbjct: 236 QKLTNTSFTEIEKQAQAVDKRKEQEDLARGAPKE-ESIVSSLRLAYKDLEISRKKDERLN 294
Query: 239 -----------------------------------PSGPPLTS-RFEYTEDVQSSELNSG 262
P L R +Y ED + S +S
Sbjct: 295 LSGQKKVEAERLGMGFGSCRSGISHSVTSDMQTIEQESPTLAKPRRKYQEDPEDSYFSSS 354
Query: 263 G----SNVTGHVSVPKS--SSSFFS-DFGMDSGFQKKSG---------------PSSSK- 299
+ + + P SSSF S D G DS ++K S PS+ +
Sbjct: 355 SKWSEQSSSRYFDDPMELRSSSFSSWDDGADSYWKKDSSRDPEPAMRSTGSSDRPSARRK 414
Query: 300 ---VQIQESDEARKKFSNAKSISSSQFFGDQNKANADAQATLSKFSGSSAISSADLFGDS 356
I +DEA+KKF N K+ISS +FG Q + + + +A L + S SS+ISSADLF +
Sbjct: 415 PEYEPIGSTDEAQKKFGNVKAISSDMYFGIQAQTDFETRARLERLSTSSSISSADLFDEQ 474
Query: 357 SDNVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDR 407
A + ++ + A D++ K KL+ A+ +MT +QDR
Sbjct: 475 RKQT--AGNYNLSNVLPNA-PDMAQFKQGVRSVAGKLSVFANGVMTSIQDR 522
>ARG3_HUMAN (Q9NP61) ADP-ribosylation factor GTPase-activating
protein 3 (ARF GAP 3)
Length = 516
Score = 179 bits (454), Expect = 1e-44
Identities = 156/517 (30%), Positives = 233/517 (44%), Gaps = 114/517 (22%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
M S D +F++L++ NK CFDC AKNP+WAS+TYG+FLCIDCS HRSLGVH+S
Sbjct: 1 MGDPSKQDILTIFKRLRSVPTNKVCFDCGAKNPSWASITYGVFLCIDCSGSHRSLGVHLS 60
Query: 61 FVRSTNLDS-WTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSK 119
F+RST LDS W+ QL+ M GGN+ A FF QHG + AKY SRAA+LY++ +
Sbjct: 61 FIRSTELDSNWSWFQLRCMQVGGNASASSFFHQHGCS-TNDTNAKYNSRAAQLYREKIKS 119
Query: 120 EVAKSMSEEAA-----------LSAPPAAS-----------SQSAQGTNGLPDVKTNEVP 157
+++ + LS PP S +A + P
Sbjct: 120 LASQATRKHGTDLWLDSCVVPPLSPPPKEEDFFASHVSPEVSDTAWASAIAEPSSLTSRP 179
Query: 158 IEKTVEKT---VEKPEKTESSSSPRAYTAVSNNL--KKPIGAKK--TGKSGGLGARKLT- 209
+E T+E E+ E + P T +++ KKP AKK K G LGA+KL
Sbjct: 180 VETTLENNEGGQEQGPSVEGLNVPTKATLEVSSIIKKKPNQAKKGLGAKKGSLGAQKLAN 239
Query: 210 ------RKPSESLYEQKPEELPAP--------VSSSTITKNNLPSGPPLTSRFEYT--ED 253
K +++ + K +E A VSS + +L + + ++
Sbjct: 240 TCFNEIEKQAQAADKMKEQEDLAKVVSKEESIVSSLRLAYKDLEIQMKKDEKMNISGKKN 299
Query: 254 VQSSELNSGGSN---VTGH---------------VSVPK-------------SSSSFFS- 281
V S L G N V H ++ P+ SSSS+F
Sbjct: 300 VDSDRLGMGFGNCRSVISHSVTSDMQTIEQESPIMAKPRKKYNDDSDDSYFTSSSSYFDE 359
Query: 282 ------------DFGMDSGFQKKSGPSSSKV-------------------QIQESDEARK 310
D DS ++K++ + V ++ +DEA+K
Sbjct: 360 PVELRSSSFSSWDDSSDSYWKKETSKDTETVLKTTGYSDRPTARRKPDYEPVENTDEAQK 419
Query: 311 KFSNAKSISSSQFFGDQNKANADAQATLSKFSGSSAISSADLFGDSSDNVDLAASDLINR 370
KF N K+ISS +FG Q++A+ + +A L + S SS+ISSADLF + A + ++
Sbjct: 420 KFGNVKAISSDMYFGRQSQADYETRARLERLSASSSISSADLFEEPRKQP--AGNYSLSS 477
Query: 371 ISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDR 407
+ A D++ K KL+ A+ ++T +QDR
Sbjct: 478 VLPNA-PDMAQFKQGVRSVAGKLSVFANGVVTSIQDR 513
>YDBH_SCHPO (Q10367) Hypothetical protein C22E12.17c in chromosome I
Length = 486
Score = 154 bits (389), Expect = 4e-37
Identities = 134/468 (28%), Positives = 211/468 (44%), Gaps = 81/468 (17%)
Query: 8 DKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFV---RS 64
+ + L+++ +NK CFDC AKNPTW+S T+GI+LC+DCSA HR++GVHISFV RS
Sbjct: 7 ESQKLLTSLRSQRDNKVCFDCGAKNPTWSSTTFGIYLCLDCSAAHRNMGVHISFVRFLRS 66
Query: 65 TNLDSWTPEQLKMMSFGGNSRAQVFFRQHG---WNGDGKVEAKYTSRAAELYKQLLSKEV 121
T LDSWT QL++M GGN A+ +F++HG KY+S+ A+ Y + L
Sbjct: 67 TVLDSWTYAQLRVMRVGGNENARNYFKRHGGVSLLNSKDCRLKYSSKTAKQYLEKLKSLA 126
Query: 122 AK--------------SMSEEAALSAP---------PAASSQSAQGTNGLPDVKTN---- 154
+ S + E + +A A S + ++ D KT+
Sbjct: 127 VEDEANYPDILDMDFLSNTHEGSSAADTTNEDDDFFSAWDKASVKKSDDNLDDKTDLAST 186
Query: 155 -----------EVPIEKTVEKT-VEKPEKTESSSSPRAYT-AVSNNLKKPIGAKKTGKSG 201
+ P+ T EKT V P + +S+S+ ++ T ++S+ +PI A +
Sbjct: 187 SSSVVVESGEKDEPVVVTEEKTMVSPPSRPDSTSTTKSKTSSISSARARPIRASSRPTAS 246
Query: 202 GLGA---RKLTRKPSES------------LYEQKPEELPAPVSSSTITKNNLPSGPPLTS 246
LGA +KL K + + E P + PA V+S T + L
Sbjct: 247 KLGASRPQKLGIKKANADIDFDEFEKAVLSSESAPTKKPAAVASKESTVDTLVDNGVEEV 306
Query: 247 RFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSF--------FSDFGMDSGFQKKSGPSS- 297
+ + VQ + + S S+ F F S + K+ +
Sbjct: 307 KESTSTTVQGKPVKPVLKSAASAKSTKSDDSNLNANFARLGFGQFAAASNARAKAAAKAR 366
Query: 298 --SKVQIQESDEARKKFSNAKSISSSQFFGDQN---KANADAQATLSKFSGSSAISSADL 352
K ++ AR F++ KSISS Q+FG + +A A+AQ LS F ++AISS
Sbjct: 367 ELKKNEVNAPTYARDHFASQKSISSDQYFGRGSFDPEAAAEAQERLSSFRDATAISSKSY 426
Query: 353 FGDSSDNVDLAASD------LINRISFQAQQDISSLKNIAGETGKKLT 394
FG+ D + S + I+ A +DI ++K + +KL+
Sbjct: 427 FGEEEDENEEGESSHRPDSAYLRDIAETATEDIEAIKVAIHQGAEKLS 474
>YQP4_CAEEL (Q09531) Hypothetical protein F07F6.4 in chromosome III
Length = 529
Score = 139 bits (351), Expect = 1e-32
Identities = 86/222 (38%), Positives = 117/222 (51%), Gaps = 19/222 (8%)
Query: 5 SFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRS 64
S D RK++ NK CFDC A+NPTW +VTYG+FLCIDCSAVHR+LGVH++FVRS
Sbjct: 8 SKVDLQTAMRKMRALPPNKLCFDCGARNPTWCTVTYGVFLCIDCSAVHRNLGVHLTFVRS 67
Query: 65 TNLD-SWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLS---KE 120
TNLD +WT QL+ M GGN A FF+ HG N + + KY SRAA++Y+ LS +E
Sbjct: 68 TNLDTNWTWLQLRAMQLGGNGNANQFFKAHGCN-TTEAQQKYKSRAAQMYRDKLSTLCQE 126
Query: 121 VAKSMSEEAALSAPPAASSQSAQGTN-----------GLPDVKTNEVPIEKTVEKTVEKP 169
+ + + A + A+ + + ++ + E + P
Sbjct: 127 AQRKFGTQLIIDTVTHAEEKPAEEEDFFAQDFGHTSASATSLSSDAYIADHKSEDSTHGP 186
Query: 170 EKTESSSSPRAYTA--VSNNLKKPIGAKKTG-KSGGLGARKL 208
SS T+ VS LKKPI G K LGA+K+
Sbjct: 187 SVDHLDSSVAVPTSAPVSVILKKPIKKATLGAKKNALGAQKV 228
Score = 43.1 bits (100), Expect = 0.001
Identities = 32/100 (32%), Positives = 53/100 (53%), Gaps = 16/100 (16%)
Query: 307 EARKKFSNAKSISSSQFFGDQNKANADAQATLSKFSGSSAISSADLFGDSSDNVDLAASD 366
+ +KKF NAK+ISS +FG + + + ++ L+K G ++ S DL+G+ S
Sbjct: 429 DLQKKFGNAKAISSDMYFGTP-EMDCETRSALTKCEGQTSFGSEDLWGNGSQ-------- 479
Query: 367 LINRISFQAQQDISSLKNI----AGETGKKLTSLASSLMT 402
R S Q D+S LK+ A + +K ++L+SS T
Sbjct: 480 --QRQSSQV-PDMSDLKDSFRAGASKVAEKWSTLSSSFST 516
>GLO3_YEAST (P38682) ADP-ribosylation factor GTPase-activating
protein GLO3
Length = 493
Score = 137 bits (346), Expect = 4e-32
Identities = 118/416 (28%), Positives = 181/416 (43%), Gaps = 71/416 (17%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
VF+KL + EN+ CFDC KNPTW SV +G+ LCI CSAVHR++GVHI+FV+S+ LD WT
Sbjct: 18 VFQKLGSNMENRVCFDCGNKNPTWTSVPFGVMLCIQCSAVHRNMGVHITFVKSSTLDKWT 77
Query: 72 PEQLKMMSFGGNSRAQVFFRQHGW-------NGDGKVEAKYTSRAAELYKQLLSKEVAKS 124
L+ GGN +A+ FF ++ N D K KYTS A+ YK L K+V K
Sbjct: 78 INNLRRFKLGGNHKARDFFLKNNGKQLLNTANVDAK--TKYTSPVAKKYKIHLDKKVQKD 135
Query: 125 MS---EEAALSAPPAA---------SSQSAQGTNGLPDVKTN-EVPIEKTVEK------- 164
M E L+ ++ +S+S N + D +N + P + K
Sbjct: 136 MELYPSELVLNGQDSSDSPLDTDSDASRSTSKENSVDDFFSNWQKPSSNSSSKLNVNTGS 195
Query: 165 --------------TVEKPEKTESSSSPRAYTAVSNNLKKPIGAKKTGKSGGLGARKLTR 210
TV K + ++S + S + KK + K L A+K+ +
Sbjct: 196 LAPKNNTTGSTPKTTVTKTRSSILTASRKKPVLNSQDKKKHSILSSSRKPTRLTAKKVDK 255
Query: 211 KPSESLYEQ--------KPEELPAPVSSSTITKNNLPSGPPLTSRFEYTEDV-----QSS 257
+E L++Q K +E SS+ I +N+ S S+ + Q
Sbjct: 256 SQAEDLFDQFKKEAQQEKEDEFTNSSSSTKIRQNDYDSQFMNNSKGNNNNSIDDINTQPD 315
Query: 258 ELNSGGSNVTGHVS----------VPKSSSSFFSDFGMDSGFQKKSGPSSSKVQ--IQES 305
E N ++ + PK + F D+ K S K+ + +
Sbjct: 316 EFNDFLNDTSNSFDTTRKEQQDTLTPKFAKLGFGMTMNDANDLAKQQKESQKIAQGPRYT 375
Query: 306 DEARKKFSNAKSISSSQFFGD---QNKANADAQATLSKFSGSSAISSADLFGDSSD 358
+++ K+ISS Q FG AN +A L F +++ISS+ FG+ +
Sbjct: 376 GRIAERYGTQKAISSDQLFGRGSFDEAANREAHDKLKTFDNATSISSSSYFGEDKE 431
>ARG1_HUMAN (Q8N6T3) ADP-ribosylation factor GTPase activating
protein 1 (ADP-ribosylation factor 1 GTPase activating
protein) (ARF1 GAP) (ARF1-directed GTPase-activating
protein) (GAP protein)
Length = 406
Score = 109 bits (272), Expect = 1e-23
Identities = 53/126 (42%), Positives = 79/126 (62%), Gaps = 4/126 (3%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
V ++++ + EN CF+C A NP W SVTYGI++C++CS HR LGVH+SFVRS +D W
Sbjct: 9 VLKEVRVQDENNVCFECGAFNPQWVSVTYGIWICLECSGRHRGLGVHLSFVRSVTMDKWK 68
Query: 72 PEQLKMMSFGGNSRAQVFFR-QHGWNGDGKVEAKYTSRAAELYKQLLSKEVAKSMSEEAA 130
+L+ M GGN++ + F Q ++ ++ KY SRAA L++ K VA + E +
Sbjct: 69 DIELEKMKAGGNAKFREFLESQEDYDPCWSLQEKYNSRAAALFR---DKVVALAEGREWS 125
Query: 131 LSAPPA 136
L + PA
Sbjct: 126 LESSPA 131
>ARG1_MOUSE (Q9EPJ9) ADP-ribosylation factor GTPase activating
protein 1 (ADP-ribosylation factor 1 GTPase activating
protein) (ARF1 GAP) (ARF1-directed GTPase-activating
protein) (GAP protein)
Length = 414
Score = 107 bits (267), Expect = 6e-23
Identities = 91/333 (27%), Positives = 145/333 (43%), Gaps = 53/333 (15%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
V ++++ + EN CF+C A NP W SVTYGI++C++CS HR LGVH+SFVRS +D W
Sbjct: 9 VLKEVRAQDENNVCFECGAFNPQWVSVTYGIWICLECSGRHRGLGVHLSFVRSVTMDKWK 68
Query: 72 PEQLKMMSFGGNSRAQVFFR-QHGWNGDGKVEAKYTSRAAELYKQLLSKEVAKSMSEEAA 130
+L+ M GGN++ + F Q + ++ KY+SRAA L++ K + +E +
Sbjct: 69 DIELEKMKAGGNAKFREFLETQDDYEPSWSLQDKYSSRAAALFR---DKVATLAEGKEWS 125
Query: 131 LSAPPAAS---------------------SQSAQGTNGLPDVKTNEVPIEKTVEK----- 164
L + PA + S +A G D +++ + ++
Sbjct: 126 LESSPAQNWTPPQPKTLQFTAHRASGQPQSAAASGDKAFEDWLNDDLGSYQGAQENRYVG 185
Query: 165 --TVEKPEKTESSSSPRAYTAVSNNLKK-PIGAKKTGKSGGLGARKLTRKPSESLYEQKP 221
P+K E A +++ + GA K + GA K + S QK
Sbjct: 186 FGNTVPPQKREDDFLNNAMSSLYSGWSSFTTGASKFASAAKEGATKFGSQAS-----QKA 240
Query: 222 EELPAPVSSSTITKNNLPSGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVP-KSSSSFF 280
EL ++ +N L +DV SS ++ S V G S + ++FF
Sbjct: 241 SEL-----GHSLNENVLKPAQEKVKEGRIFDDV-SSGVSQLASKVQGVGSKGWRDVTTFF 294
Query: 281 SDFGMDS--------GFQKKSGPSSSKVQIQES 305
S DS +Q SG +S I +S
Sbjct: 295 SGKAEDSSDRPLEGHSYQNSSGDNSQNSNIDQS 327
>ARG1_RAT (Q62848) ADP-ribosylation factor GTPase activating protein
1 (ADP-ribosylation factor 1 GTPase activating protein)
(ARF1 GAP) (ARF1-directed GTPase-activating protein)
(GAP protein)
Length = 415
Score = 107 bits (266), Expect = 7e-23
Identities = 51/126 (40%), Positives = 78/126 (61%), Gaps = 4/126 (3%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
V ++++ + EN CF+C A NP W SVTYGI++C++CS HR LGVH+SFVRS +D W
Sbjct: 9 VLKEVRAQDENNVCFECGAFNPQWVSVTYGIWICLECSGRHRGLGVHLSFVRSVTMDKWK 68
Query: 72 PEQLKMMSFGGNSRAQVFFR-QHGWNGDGKVEAKYTSRAAELYKQLLSKEVAKSMSEEAA 130
+L+ M GGN++ + F Q + ++ KY+SRAA L++ K + +E +
Sbjct: 69 DIELEKMKAGGNAKFREFLEAQDDYEPSWSLQDKYSSRAAALFR---DKVATLAEGKEWS 125
Query: 131 LSAPPA 136
L + PA
Sbjct: 126 LESSPA 131
>GCS1_YEAST (P35197) ADP-ribosylation factor GTPase-activating
protein GCS1
Length = 352
Score = 99.4 bits (246), Expect = 1e-20
Identities = 67/232 (28%), Positives = 99/232 (41%), Gaps = 31/232 (13%)
Query: 15 KLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPEQ 74
+L+ NK C DC A NP WA+ +G F+C++C+ +HR LGVHISFVRS +D + PE+
Sbjct: 16 QLQKIGANKKCMDCGAPNPQWATPKFGAFICLECAGIHRGLGVHISFVRSITMDQFKPEE 75
Query: 75 LKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLS---------------- 118
L M GGN +F+ H + + KY + AE YK+ L+
Sbjct: 76 LLRMEKGGNEPLTEWFKSHNIDLSLPQKVKYDNPVAEDYKEKLTCLCEDRVFEEREHLDF 135
Query: 119 -KEVAKSMSEEAALSAPPAASSQS------------AQGTNGLPDVK-TNEVPIEKTVEK 164
+ S+ AA + P A S+ A +NG K NE + +K
Sbjct: 136 DASKLSATSQTAASATPGVAQSREGTPLENRRSATPANSSNGANFQKEKNEAYFAELGKK 195
Query: 165 TVEKPEKTESSSSPRAYTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESL 216
+P+ S + Y + KP + G S L P +L
Sbjct: 196 NQSRPDHLPPSQGGK-YQGFGSTPAKPPQERSAGSSNTLSLENFQADPLGTL 246
>CEB5_HUMAN (Q96P50) Centaurin beta 5 (Cnt-b5)
Length = 759
Score = 90.1 bits (222), Expect = 9e-18
Identities = 49/127 (38%), Positives = 71/127 (55%), Gaps = 3/127 (2%)
Query: 11 AVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSW 70
+V +++++ + N C DC +P WAS+ G+ LCI+CS +HRSLGVH S VRS LDSW
Sbjct: 362 SVLQRVQSVAGNSQCGDCGQPDPRWASINLGVLLCIECSGIHRSLGVHCSKVRSLTLDSW 421
Query: 71 TPEQLKMMSFGGNSRA-QVFFRQHGWNGDGKVEAKYTSRAAELY--KQLLSKEVAKSMSE 127
PE LK+M GNS Q++ Q G K A + + E + + + K+ +
Sbjct: 422 EPELLKLMCELGNSAVNQIYEAQCEGAGSRKPTASSSRQDKEAWIKDKYVEKKFLRKAPM 481
Query: 128 EAALSAP 134
AL AP
Sbjct: 482 APALEAP 488
>CEB2_HUMAN (Q15057) Centaurin beta 2 (Cnt-b2)
Length = 778
Score = 84.0 bits (206), Expect = 7e-16
Identities = 59/175 (33%), Positives = 82/175 (46%), Gaps = 20/175 (11%)
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPEQLKMMSFG 81
N SC DC +P WAS+ GI LCI+CS +HRSLGVH S VRS LD+W PE LK+M
Sbjct: 411 NASCCDCGLADPRWASINLGITLCIECSGIHRSLGVHFSKVRSLTLDTWEPELLKLMCEL 470
Query: 82 GNSRAQVFFR-----------QHGWNGDGK--VEAKYTSRA-AELYKQLLS-----KEVA 122
GN + Q G + + + AKY R + Y LS K+
Sbjct: 471 GNDVINRVYEANVEKMGIKKPQPGQRQEKEAYIRAKYVERKFVDKYSISLSPPEQQKKFV 530
Query: 123 KSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSS 177
SEE LS + + V++N+ I+++ + E T S++S
Sbjct: 531 SKSSEEKRLSISKFGPGDQVR-ASAQSSVRSNDSGIQQSSDDGRESLPSTVSANS 584
>CEG3_HUMAN (Q96P47) Centaurin gamma 3
Length = 876
Score = 81.6 bits (200), Expect = 3e-15
Identities = 35/71 (49%), Positives = 47/71 (65%)
Query: 16 LKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPEQL 75
++T N C DC+A NP WAS+ G +CI+CS +HR LG H+S VRS +LD W PE L
Sbjct: 633 VRTVRGNSFCIDCDAPNPDWASLNLGALMCIECSGIHRHLGAHLSRVRSLDLDDWPPELL 692
Query: 76 KMMSFGGNSRA 86
+M+ GN+ A
Sbjct: 693 AVMTAMGNALA 703
>CG1A_DROME (Q9NGC3) Centaurin gamma 1A protein
Length = 995
Score = 80.5 bits (197), Expect = 7e-15
Identities = 44/109 (40%), Positives = 57/109 (51%), Gaps = 5/109 (4%)
Query: 7 TDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTN 66
TD A+ + N C DC A NP WAS+ G+ +CI+CS VHR+LG HIS VRS
Sbjct: 699 TDLAAMLAIRQRVPGNGFCVDCGAPNPEWASLNLGVLMCIECSGVHRNLGSHISKVRSLG 758
Query: 67 LDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQ 115
LD W L +M GNS A W + + K TS+A+ K+
Sbjct: 759 LDDWPSPHLSVMLAIGNSLANSV-----WESNTRQRVKPTSQASREDKE 802
>CEG2_MOUSE (Q8BXK8) Centaurin gamma 2 (ARF-GAP with GTP-binding
protein-like, ankyrin repeat and pleckstrin homology
domains 1) (AGAP1)
Length = 857
Score = 79.7 bits (195), Expect = 1e-14
Identities = 33/73 (45%), Positives = 47/73 (64%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
+ ++ N C DC+ +NP WAS+ G +CI+CS +HR+LG H+S VRS +LD W E
Sbjct: 613 QSIRNMRGNSHCVDCDTQNPNWASLNLGALMCIECSGIHRNLGTHLSRVRSLDLDDWPME 672
Query: 74 QLKMMSFGGNSRA 86
+K+MS GN A
Sbjct: 673 LIKVMSSIGNELA 685
>CEB1_HUMAN (Q15027) Centaurin beta 1 (Cnt-b1)
Length = 740
Score = 79.7 bits (195), Expect = 1e-14
Identities = 36/72 (50%), Positives = 45/72 (62%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWT 71
V ++++ N C DC P WAS+ G+ LCI CS +HRSLGVH S VRS LDSW
Sbjct: 407 VVAQVQSVDGNAQCCDCREPAPEWASINLGVTLCIQCSGIHRSLGVHFSKVRSLTLDSWE 466
Query: 72 PEQLKMMSFGGN 83
PE +K+M GN
Sbjct: 467 PELVKLMCELGN 478
>YIE4_YEAST (P40529) Hypothetical 32.6 kDa protein in MET30-PIG2
intergenic region
Length = 298
Score = 79.3 bits (194), Expect = 2e-14
Identities = 80/275 (29%), Positives = 124/275 (45%), Gaps = 31/275 (11%)
Query: 22 NKSCFDCNAK-NPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPEQL-KMMS 79
N C DC A+ +P WAS + G+F+CI C+ +HRSLG HIS V+S +LD+W E L K++
Sbjct: 20 NSHCADCKAQLHPRWASWSLGVFICIKCAGIHRSLGTHISKVKSVDLDTWKEEHLVKLIQ 79
Query: 80 FGGNSRAQVFFRQHGWNGDGKVEAKYTSRAA---------ELYKQL--LSKEVAKSMSEE 128
F N RA ++ D + K T ++ E K + LS + S E
Sbjct: 80 FKNNLRANSYY--EATLADELKQRKITDTSSLQNFIKNKYEYKKWIGDLSSIEGLNDSTE 137
Query: 129 AALSAP------PAASSQSAQGTNGLPDVKTNEVP--IEKTVEKTVEKPEKTESSSSPRA 180
L P PA++++ Q +N L +T + + T + S S +
Sbjct: 138 PVLHKPSANHSLPASNARLDQSSNSLQKTQTQPPSHLLSTSRSNTSLLNLQVSSLSKTTS 197
Query: 181 YTAVSNNLKKPIGAKKT---GKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNN 237
T+V+++ IGA T + G G R +K SLY KP +S +
Sbjct: 198 NTSVTSSATS-IGAANTKTGNRVGEFGQRNDLKKSILSLY-SKPSAQTQSQNSFFTSTTP 255
Query: 238 LPSGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSV 272
P P S F T + ++ NS SN + ++S+
Sbjct: 256 QPCNTP--SPFVNT-GITATNNNSMNSNSSSNISL 287
>CEG2_HUMAN (Q9UPQ3) Centaurin gamma 2 (ARF-GAP with GTP-binding
protein-like, ankyrin repeat and pleckstrin homology
domains 1) (AGAP1)
Length = 857
Score = 79.3 bits (194), Expect = 2e-14
Identities = 37/106 (34%), Positives = 58/106 (53%), Gaps = 3/106 (2%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
+ ++ N C DC +NP WAS+ G +CI+CS +HR+LG H+S VRS +LD W E
Sbjct: 613 QSIRNMRGNSHCVDCETQNPNWASLNLGALMCIECSGIHRNLGTHLSRVRSLDLDDWPIE 672
Query: 74 QLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSK 119
+K+MS GN A + + + G+ + S E + + +K
Sbjct: 673 LIKVMSSIGNELANSVWEE---SSQGRTKPSVDSTREEKERWIRAK 715
>CEG1_HUMAN (Q99490) Centaurin gamma 1
Length = 836
Score = 78.6 bits (192), Expect = 3e-14
Identities = 39/104 (37%), Positives = 58/104 (55%), Gaps = 6/104 (5%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
+ +DS ++ A+ + ++ N C DC A NPTWAS+ G +CI+CS +HR+LG H+S
Sbjct: 567 LRTDSQSEAVAI-QAIRNAKGNSICVDCGAPNPTWASLNLGALICIECSGIHRNLGTHLS 625
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAK 104
VRS +LD W E +++ GN A W D + AK
Sbjct: 626 RVRSLDLDDWPRELTLVLTAIGNDTA-----NRVWESDTRGRAK 664
>SP18_YEAST (P32572) Sporulation protein SPS18 (Sporulation-specific
protein SPX18)
Length = 300
Score = 70.5 bits (171), Expect = 7e-12
Identities = 33/108 (30%), Positives = 65/108 (59%), Gaps = 1/108 (0%)
Query: 15 KLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPEQ 74
+ K + N +CF+C + NP + S ++GIF+C++C+ + R +G +I V+S +D++ +
Sbjct: 18 RAKKAAGNNNCFECKSVNPQFVSCSFGIFICVNCANLLRGMGTNIFCVKSITMDNFEEKD 77
Query: 75 LKMMSFGGNSRAQVFFRQHGWNGDG-KVEAKYTSRAAELYKQLLSKEV 121
++ + GN+R F ++G +G + KY + A+ YK+ L+ EV
Sbjct: 78 VRRVEKSGNNRFGSFLSKNGILQNGIPLREKYDNLFAKSYKRRLANEV 125
>CSX2_SCHPO (Q9UUE2) Protein csx2
Length = 870
Score = 67.4 bits (163), Expect = 6e-11
Identities = 29/66 (43%), Positives = 42/66 (62%), Gaps = 1/66 (1%)
Query: 20 SENKSCFDCNAK-NPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPEQLKMM 78
S ++SC DCN W ++ + + LCIDCS +HRSLG HI+ +RS LD + PE + ++
Sbjct: 681 SSDQSCADCNTTARVEWCAINFPVVLCIDCSGIHRSLGTHITKIRSLTLDKFNPETVDLL 740
Query: 79 SFGGNS 84
GNS
Sbjct: 741 YATGNS 746
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.307 0.123 0.332
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 43,934,618
Number of Sequences: 164201
Number of extensions: 1777593
Number of successful extensions: 6803
Number of sequences better than 10.0: 259
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 193
Number of HSP's that attempted gapping in prelim test: 5831
Number of HSP's gapped (non-prelim): 564
length of query: 409
length of database: 59,974,054
effective HSP length: 113
effective length of query: 296
effective length of database: 41,419,341
effective search space: 12260124936
effective search space used: 12260124936
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC137670.5


