

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137667.13 - phase: 0 /pseudo
(267 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
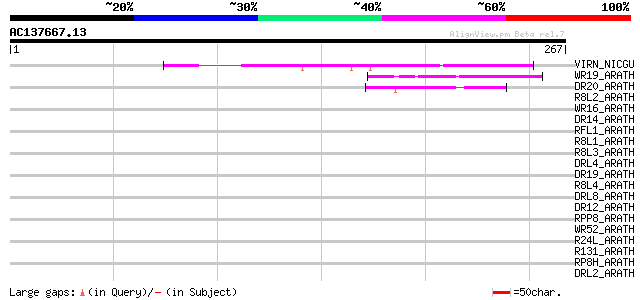
Score E
Sequences producing significant alignments: (bits) Value
VIRN_NICGU (Q40392) TMV resistance protein N 49 1e-05
WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY ... 45 1e-04
DR20_ARATH (Q9SH22) Putative disease resistance protein At1g63360 44 3e-04
R8L2_ARATH (Q9MAG6) Probable disease resistance RPP8-like protein 2 41 0.003
WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY ... 39 0.017
DR14_ARATH (Q9SI85) Probable disease resistance protein At1g6263... 38 0.029
RFL1_ARATH (Q8L3R3) Disease resistance protein RFL1 (RPS5-like p... 37 0.050
R8L1_ARATH (O04093) Putative disease resistance protein RPP8-lik... 37 0.065
R8L3_ARATH (Q9FJB5) Probable disease resistance RPP8-like protein 3 36 0.085
DRL4_ARATH (Q9SX38) Putative disease resistance protein At1g50180 36 0.085
DR19_ARATH (Q9C8T9) Putative disease resistance protein At1g63350 36 0.085
R8L4_ARATH (Q9FJK8) Probable disease resistance RPP8-like protein 4 36 0.11
DRL8_ARATH (Q8W3K3) Putative disease resistance protein At1g58400 36 0.11
DR12_ARATH (Q9LQ54) Probable disease resistance protein At1g5962... 35 0.19
RPP8_ARATH (Q8W4J9) Disease resistance protein RPP8 (Resistance ... 35 0.25
WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY ... 34 0.42
R24L_ARATH (Q9C646) Probable disease resistance protein RXW24L 34 0.42
R131_ARATH (Q9LRR4) Putative disease resistance RPP13-like prote... 34 0.42
RP8H_ARATH (P59584) Disease resistance protein RPH8A (RPP8 homol... 33 0.55
DRL2_ARATH (P60839) Probable disease resistance protein At1g12290 33 0.55
>VIRN_NICGU (Q40392) TMV resistance protein N
Length = 1144
Score = 49.3 bits (116), Expect = 1e-05
Identities = 47/201 (23%), Positives = 91/201 (44%), Gaps = 44/201 (21%)
Query: 75 DSLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKP 134
+S IVVFS+NYA+S +CL ++ I+ K+ ++
Sbjct: 65 ESQFAIVVFSENYATSR--------------------WCLNELVKIMECKTRFKQTVIPI 104
Query: 135 FSNMN----KDSNKTLRWCKDGGKLKHKSPISG---WDVRHNDAQ--------------- 172
F +++ ++ ++ + + K+K + G W + N+A
Sbjct: 105 FYDVDPSHVRNQKESFAKAFEEHETKYKDDVEGIQRWRIALNEAANLKGSCDNRDKTDAD 164
Query: 173 -IEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKK 231
I +IV++I + L S S + G+D+ E++E LL ++ VR++G+ GMGG+GK
Sbjct: 165 CIRQIVDQISSKLCKISLSYLQNIVGIDTHLEKIES-LLEIGINGVRIMGIWGMGGVGKT 223
Query: 232 AIATALYNKIFHQFPVLFLID 252
IA A+++ + + + D
Sbjct: 224 TIARAIFDTLLGRMDSSYQFD 244
>WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY
DNA-binding protein 19)
Length = 1895
Score = 45.4 bits (106), Expect = 1e-04
Identities = 30/84 (35%), Positives = 47/84 (55%), Gaps = 4/84 (4%)
Query: 173 IEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKA 232
I++IV + + +L S N + GMD EE+ L ++S+D VR +G+ G GIGK
Sbjct: 797 IDEIVRDALKVLC--SADKVNMI-GMDMQVEEILSLLCIESLD-VRSIGIWGTVGIGKTT 852
Query: 233 IATALYNKIFHQFPVLFLIDDLRK 256
IA ++ KI Q+ ++ DL K
Sbjct: 853 IAEEIFRKISVQYETCVVLKDLHK 876
>DR20_ARATH (Q9SH22) Putative disease resistance protein At1g63360
Length = 884
Score = 44.3 bits (103), Expect = 3e-04
Identities = 22/75 (29%), Positives = 41/75 (54%), Gaps = 10/75 (13%)
Query: 172 QIEKIVEEIMNIL-------GYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCG 224
++EK+ E+ ++ ++ L + G D++ ++ KHL+ D V ++G+ G
Sbjct: 123 EVEKLKGEVFGVITEQASTSAFEERPLQPTIVGQDTMLDKAGKHLMEDGVG---IMGMYG 179
Query: 225 MGGIGKKAIATALYN 239
MGG+GK + T LYN
Sbjct: 180 MGGVGKTTLLTQLYN 194
>R8L2_ARATH (Q9MAG6) Probable disease resistance RPP8-like protein 2
Length = 1271
Score = 41.2 bits (95), Expect = 0.003
Identities = 24/49 (48%), Positives = 30/49 (60%), Gaps = 6/49 (12%)
Query: 195 LAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFH 243
L G+D EEL HL+ + D V+VV V GMGGIGK T L ++FH
Sbjct: 163 LVGLDQSVEELVDHLVEN--DSVQVVSVSGMGGIGK----TTLARQVFH 205
>WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY
DNA-binding protein 16)
Length = 1372
Score = 38.5 bits (88), Expect = 0.017
Identities = 22/66 (33%), Positives = 37/66 (55%), Gaps = 1/66 (1%)
Query: 197 GMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDDLRK 256
G+ S E+EK + +D +R VG+ GM GIGK +A A+++++ +F I+D K
Sbjct: 144 GIYSKLLEIEKMINKQPLD-IRCVGIWGMPGIGKTTLAKAVFDQMSGEFDAHCFIEDYTK 202
Query: 257 IYRHDG 262
+ G
Sbjct: 203 AIQEKG 208
>DR14_ARATH (Q9SI85) Probable disease resistance protein At1g62630
(pNd4)
Length = 893
Score = 37.7 bits (86), Expect = 0.029
Identities = 19/75 (25%), Positives = 38/75 (50%), Gaps = 10/75 (13%)
Query: 172 QIEKIVEEIMNIL-------GYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCG 224
++EK+ E+ ++ ++ L + G + ++ KHL+ D ++G+ G
Sbjct: 123 EVEKLKGEVFGVITEQASTSAFEERPLQPTIVGQKKMLDKAWKHLMEDGTG---IMGMYG 179
Query: 225 MGGIGKKAIATALYN 239
MGG+GK + T L+N
Sbjct: 180 MGGVGKTTLLTQLFN 194
>RFL1_ARATH (Q8L3R3) Disease resistance protein RFL1 (RPS5-like
protein 1) (pNd13/pNd14)
Length = 885
Score = 37.0 bits (84), Expect = 0.050
Identities = 21/65 (32%), Positives = 35/65 (53%), Gaps = 3/65 (4%)
Query: 176 IVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIAT 235
IV E I + + + + G DS+ +++ L+ D V +VG+ GMGG+GK + T
Sbjct: 138 IVTEAAPIAEVEELPIQSTIVGQDSMLDKVWNCLMEDKV---WIVGLYGMGGVGKTTLLT 194
Query: 236 ALYNK 240
+ NK
Sbjct: 195 QINNK 199
>R8L1_ARATH (O04093) Putative disease resistance protein RPP8-like
protein 1
Length = 821
Score = 36.6 bits (83), Expect = 0.065
Identities = 20/49 (40%), Positives = 30/49 (60%), Gaps = 6/49 (12%)
Query: 195 LAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFH 243
L G++ E L HL+ + D+++VV + GMGGIGK T L ++FH
Sbjct: 140 LVGVEQSVEALAGHLVEN--DNIQVVSISGMGGIGK----TTLARQVFH 182
>R8L3_ARATH (Q9FJB5) Probable disease resistance RPP8-like protein 3
Length = 901
Score = 36.2 bits (82), Expect = 0.085
Identities = 20/49 (40%), Positives = 30/49 (60%), Gaps = 6/49 (12%)
Query: 195 LAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFH 243
L G++ EEL ++ +D+++VV + GMGGIGK T L +IFH
Sbjct: 163 LVGVEQSVEELVGPMV--EIDNIQVVSISGMGGIGK----TTLARQIFH 205
>DRL4_ARATH (Q9SX38) Putative disease resistance protein At1g50180
Length = 839
Score = 36.2 bits (82), Expect = 0.085
Identities = 19/67 (28%), Positives = 37/67 (54%), Gaps = 6/67 (8%)
Query: 195 LAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYN-----KIFHQFPVLF 249
L G++ E+L L+ + +RV +CGMGG+GK +A +++ + F +F ++
Sbjct: 164 LVGLEQSLEKLVNDLVSGG-EKLRVTSICGMGGLGKTTLAKQIFHHHKVRRHFDRFAWVY 222
Query: 250 LIDDLRK 256
+ D R+
Sbjct: 223 VSQDCRR 229
>DR19_ARATH (Q9C8T9) Putative disease resistance protein At1g63350
Length = 898
Score = 36.2 bits (82), Expect = 0.085
Identities = 18/76 (23%), Positives = 40/76 (51%), Gaps = 10/76 (13%)
Query: 172 QIEKIVEEIMNILGYKSTS-------LPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCG 224
++EK+ + ++ ++++ L + G +++ + HL+ D V ++G+ G
Sbjct: 123 EVEKLERRVFEVISDQASTSEVEEQQLQPTIVGQETMLDNAWNHLMEDGVG---IMGLYG 179
Query: 225 MGGIGKKAIATALYNK 240
MGG+GK + T + NK
Sbjct: 180 MGGVGKTTLLTQINNK 195
>R8L4_ARATH (Q9FJK8) Probable disease resistance RPP8-like protein 4
Length = 908
Score = 35.8 bits (81), Expect = 0.11
Identities = 20/49 (40%), Positives = 29/49 (58%), Gaps = 6/49 (12%)
Query: 195 LAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFH 243
L G++ EEL HL+ + + +VV + GMGGIGK T L ++FH
Sbjct: 165 LVGVEQSVEELVGHLVENDI--YQVVSIAGMGGIGK----TTLARQVFH 207
>DRL8_ARATH (Q8W3K3) Putative disease resistance protein At1g58400
Length = 910
Score = 35.8 bits (81), Expect = 0.11
Identities = 21/63 (33%), Positives = 37/63 (58%), Gaps = 9/63 (14%)
Query: 185 GYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYN--KIF 242
GY+S G++ ++L +L+ + DD+++V V GMGG+GK +A ++N +
Sbjct: 159 GYESD-----FVGLEVNVKKLVGYLVEE--DDIQIVSVTGMGGLGKTTLARQVFNHEDVK 211
Query: 243 HQF 245
HQF
Sbjct: 212 HQF 214
>DR12_ARATH (Q9LQ54) Probable disease resistance protein At1g59620
(CW9)
Length = 695
Score = 35.0 bits (79), Expect = 0.19
Identities = 18/52 (34%), Positives = 31/52 (59%), Gaps = 1/52 (1%)
Query: 188 STSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYN 239
S + + L G++ ++L HL+ + D +VV + GMGGIGK +A ++N
Sbjct: 132 SNNNESVLVGLEENVKKLVGHLV-EVEDSSQVVSITGMGGIGKTTLARQVFN 182
>RPP8_ARATH (Q8W4J9) Disease resistance protein RPP8 (Resistance to
Peronospora parasitica protein 8)
Length = 908
Score = 34.7 bits (78), Expect = 0.25
Identities = 20/49 (40%), Positives = 29/49 (58%), Gaps = 6/49 (12%)
Query: 195 LAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFH 243
L G++ +EL HL+ + V +VV + GMGGIGK T L ++FH
Sbjct: 165 LVGVEQSVKELVGHLVENDVH--QVVSIAGMGGIGK----TTLARQVFH 207
>WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY
DNA-binding protein 52) (Disease resistance protein
RRS1) (Resistance to Ralstonia solanacearum 1 protein)
Length = 1378
Score = 33.9 bits (76), Expect = 0.42
Identities = 16/46 (34%), Positives = 25/46 (53%)
Query: 217 VRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDDLRKIYRHDG 262
+R VG+ GM GIGK +A A+++++ F I+D K G
Sbjct: 172 IRCVGIWGMPGIGKTTLAKAVFDQMSSAFDASCFIEDYDKSIHEKG 217
>R24L_ARATH (Q9C646) Probable disease resistance protein RXW24L
Length = 899
Score = 33.9 bits (76), Expect = 0.42
Identities = 16/47 (34%), Positives = 29/47 (61%), Gaps = 2/47 (4%)
Query: 193 NYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYN 239
N GM++ ++L +L+ DD ++V + GMGG+GK +A ++N
Sbjct: 160 NDFVGMEANVKKLVGYLV--EKDDYQIVSLTGMGGLGKTTLARQVFN 204
>R131_ARATH (Q9LRR4) Putative disease resistance RPP13-like protein
1
Length = 1054
Score = 33.9 bits (76), Expect = 0.42
Identities = 31/125 (24%), Positives = 60/125 (47%), Gaps = 23/125 (18%)
Query: 137 NMNKDSNKTLRWCKDGGKLKHKSPISGWDVRHNDAQIEKIVEEI------MNILGYK--- 187
N+ +S+ + R + G++ + G + H + ++EK+ + NILG K
Sbjct: 95 NIGAESSSSNRLRQLRGRMSLGDFLDG-NSEHLETRLEKVTIRLERLASQRNILGLKELT 153
Query: 188 ---------STSL--PNYLAGMDSLTEELEKHLLLDSVDD--VRVVGVCGMGGIGKKAIA 234
+TSL + + G D +E+ + L+ ++ D + VV + G+GG+GK ++
Sbjct: 154 AMIPKQRLPTTSLVDESEVFGRDDDKDEIMRFLIPENGKDNGITVVAIVGIGGVGKTTLS 213
Query: 235 TALYN 239
LYN
Sbjct: 214 QLLYN 218
>RP8H_ARATH (P59584) Disease resistance protein RPH8A (RPP8 homolog
A)
Length = 910
Score = 33.5 bits (75), Expect = 0.55
Identities = 20/49 (40%), Positives = 28/49 (56%), Gaps = 6/49 (12%)
Query: 195 LAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFH 243
L G++ EL HL+ + V +VV + GMGGIGK T L ++FH
Sbjct: 165 LVGVEQSVTELVCHLVENDVH--QVVSIAGMGGIGK----TTLARQVFH 207
>DRL2_ARATH (P60839) Probable disease resistance protein At1g12290
Length = 884
Score = 33.5 bits (75), Expect = 0.55
Identities = 14/46 (30%), Positives = 28/46 (60%), Gaps = 3/46 (6%)
Query: 195 LAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNK 240
+ G +++ E+ HL+ D +++G+ GMGG+GK + T + N+
Sbjct: 156 IVGQETILEKAWDHLMDDGT---KIMGLYGMGGVGKTTLLTQINNR 198
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.339 0.149 0.474
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,689,779
Number of Sequences: 164201
Number of extensions: 1117771
Number of successful extensions: 5061
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 5015
Number of HSP's gapped (non-prelim): 62
length of query: 267
length of database: 59,974,054
effective HSP length: 108
effective length of query: 159
effective length of database: 42,240,346
effective search space: 6716215014
effective search space used: 6716215014
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC137667.13


