

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
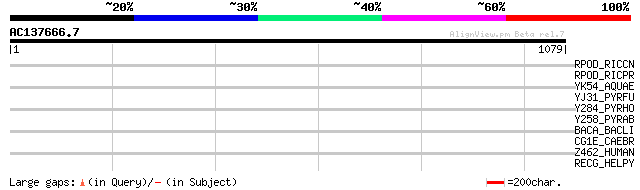
Query= AC137666.7 - phase: 0
(1079 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
RPOD_RICCN (Q92FZ8) RNA polymerase sigma factor rpoD (Sigma-70) 38 0.13
RPOD_RICPR (P33451) RNA polymerase sigma factor rpoD (Sigma-70) 37 0.29
YK54_AQUAE (O67838) Hypothetical protein AQ_2054 35 1.1
YJ31_PYRFU (Q8TZQ5) Hypothetical UPF0273 protein PF1931 34 1.9
Y284_PYRHO (O58022) Hypothetical UPF0273 protein PH0284 34 1.9
Y258_PYRAB (Q9V216) Hypothetical UPF0273 protein PYRAB02580 34 1.9
BACA_BACLI (O68006) Bacitracin synthetase 1 (BA1) [Includes: ATP... 34 1.9
CG1E_CAEBR (Q8MUK3) G1/S-specific cyclin E 34 2.4
Z462_HUMAN (Q96JM2) Zinc finger protein 462 32 7.1
RECG_HELPY (O26051) ATP-dependent DNA helicase recG (EC 3.6.1.-) 32 7.1
>RPOD_RICCN (Q92FZ8) RNA polymerase sigma factor rpoD (Sigma-70)
Length = 634
Score = 38.1 bits (87), Expect = 0.13
Identities = 28/109 (25%), Positives = 46/109 (41%), Gaps = 10/109 (9%)
Query: 475 QLYEKVKLLSSKFGKPFSGELVTGGWKKKSIFFELPYWKSLYVRH--FLDVMHIEKNVFD 532
+L EK + F + G ++ WK+K + + WK L + ++D M E +V +
Sbjct: 295 KLAEKYGVTRQNFLDEYIGSVINAAWKEKMLKNKKVAWKELMTKESDYIDQMIAELSVIE 354
Query: 533 SVVGTLLN--------IPGKSKDGINARLDMQSMGLRNELRPVKKDGKR 573
S G L+N I + + A+ DM LR + KK R
Sbjct: 355 SKTGLLVNDFKKLVNTIQKSERQTLQAKKDMIEANLRLVISIAKKYANR 403
>RPOD_RICPR (P33451) RNA polymerase sigma factor rpoD (Sigma-70)
Length = 635
Score = 37.0 bits (84), Expect = 0.29
Identities = 28/109 (25%), Positives = 46/109 (41%), Gaps = 10/109 (9%)
Query: 475 QLYEKVKLLSSKFGKPFSGELVTGGWKKKSIFFELPYWKSLYVRH--FLDVMHIEKNVFD 532
+L EK + F + G ++ WK+K + + WK L + ++D M E +V +
Sbjct: 296 KLAEKYGVTRQNFLDEYIGSVINAAWKEKMLKNKKVAWKELITKESDYIDQMINELSVIE 355
Query: 533 SVVGTLLN--------IPGKSKDGINARLDMQSMGLRNELRPVKKDGKR 573
S G L+N I + + A+ DM LR + KK R
Sbjct: 356 SNTGLLVNDFKKLVNTIQKSERQTLQAKKDMIEANLRLVISIAKKYANR 404
>YK54_AQUAE (O67838) Hypothetical protein AQ_2054
Length = 1116
Score = 35.0 bits (79), Expect = 1.1
Identities = 33/100 (33%), Positives = 46/100 (46%), Gaps = 10/100 (10%)
Query: 478 EKVKLLSSKFGKPFSGELVTGGWKKKSIFFELPYWKSLYVRHFLDVMHIEKNVFDSVVGT 537
E V++ S F +PF G + +K F L W + FL+ I KN D V
Sbjct: 760 EPVRINSLYFYEPF-GSAMNLELRKDFFAFALTGW---FREGFLNAYAISKNFRDFEVEF 815
Query: 538 LL-NIPGKSKDGINARLDMQSMGLRNELRPVKKDGKRTFL 576
+ N+P K KDGI AR++ G + V K+ K FL
Sbjct: 816 VYKNLPVKLKDGIRARIEADGKG-----KVVVKNFKEIFL 850
>YJ31_PYRFU (Q8TZQ5) Hypothetical UPF0273 protein PF1931
Length = 247
Score = 34.3 bits (77), Expect = 1.9
Identities = 22/68 (32%), Positives = 36/68 (52%), Gaps = 3/68 (4%)
Query: 89 STDFEKDTNYEFDQVEEILNVIEDDLRDCPKMFERVVIDAETPLYEGCTKFTRLSTILKL 148
S ++EK ++ + E + V+ +RD +RVV+D+ T LY R S IL+L
Sbjct: 99 SKEYEKYIVHDLTDIREFIEVLRQAIRDINA--KRVVVDSVTTLYINKPAMAR-SIILQL 155
Query: 149 YKLKAGNG 156
++ AG G
Sbjct: 156 KRVLAGTG 163
>Y284_PYRHO (O58022) Hypothetical UPF0273 protein PH0284
Length = 247
Score = 34.3 bits (77), Expect = 1.9
Identities = 22/68 (32%), Positives = 36/68 (52%), Gaps = 3/68 (4%)
Query: 89 STDFEKDTNYEFDQVEEILNVIEDDLRDCPKMFERVVIDAETPLYEGCTKFTRLSTILKL 148
S ++EK ++ + E + V+ +RD +RVV+D+ T LY R S IL+L
Sbjct: 99 SKEYEKYIVHDLTDIREFIEVLRQAIRDINA--KRVVVDSVTTLYINKPAMAR-SIILQL 155
Query: 149 YKLKAGNG 156
++ AG G
Sbjct: 156 KRVLAGTG 163
>Y258_PYRAB (Q9V216) Hypothetical UPF0273 protein PYRAB02580
Length = 247
Score = 34.3 bits (77), Expect = 1.9
Identities = 22/68 (32%), Positives = 36/68 (52%), Gaps = 3/68 (4%)
Query: 89 STDFEKDTNYEFDQVEEILNVIEDDLRDCPKMFERVVIDAETPLYEGCTKFTRLSTILKL 148
S ++EK ++ + E + V+ +RD +RVV+D+ T LY R S IL+L
Sbjct: 99 SKEYEKYIVHDLTDIREFIEVLRQAIRDINA--KRVVVDSVTTLYINKPALAR-SIILQL 155
Query: 149 YKLKAGNG 156
++ AG G
Sbjct: 156 KRVLAGTG 163
>BACA_BACLI (O68006) Bacitracin synthetase 1 (BA1) [Includes:
ATP-dependent isoleucine adenylase (IleA) (Isoleucine
activase); ATP-dependent cysteine adenylase (CysA)
(Cysteine activase); ATP-dependent leucine adenylase
(LeuA) (Leucine activase); ATP-depe
Length = 5255
Score = 34.3 bits (77), Expect = 1.9
Identities = 35/130 (26%), Positives = 63/130 (47%), Gaps = 11/130 (8%)
Query: 667 DPEKLPTLQNEIVVTLCELEMYFPPSFFDIMVHLTVHLIMETQYCGPAYMRWMYPIERYM 726
D EK L +E ++ +C ++M S+ I H H I+ +C + ++ + Y
Sbjct: 4311 DQEKGFDLSSEALMRVCLIKMS-DESYRLIWSH---HHILLDGWCLGIVLSELFSL--YG 4364
Query: 727 KILKGYVKN--QSRPEGCIV---ERYIVEEAVEFFTDFLSNVESIGLPMSRHSGRTTGEG 781
KI+KG + + +P G + E+ EEAV ++ D+L ES + + G T+ E
Sbjct: 4365 KIMKGESRRLKEPKPYGDYIKWLEKQDQEEAVAYWKDYLKGYESRSELPAFNRGATSEEY 4424
Query: 782 IGASKLVTIS 791
G K+++ S
Sbjct: 4425 CGKEKVISFS 4434
>CG1E_CAEBR (Q8MUK3) G1/S-specific cyclin E
Length = 518
Score = 33.9 bits (76), Expect = 2.4
Identities = 28/83 (33%), Positives = 37/83 (43%), Gaps = 3/83 (3%)
Query: 746 RYIVEEAVEFFTDFLSNVESIGLPMSRHSGRTTGEGIGASKLVTISDIELEQAHLYVLHN 805
R VE+A F + L NV P+ R R T E + K + E AH +H
Sbjct: 436 RSAVEKATGFIYEQLRNVIDYVTPICRAYDRFTQEHV--PKDIIPDYAPAEDAHNIQVHI 493
Query: 806 AD-EVEPYVKKHKEILQSLNPNR 827
EV+PYV+K +E PNR
Sbjct: 494 KHYEVDPYVEKERERRHRRGPNR 516
>Z462_HUMAN (Q96JM2) Zinc finger protein 462
Length = 1412
Score = 32.3 bits (72), Expect = 7.1
Identities = 22/84 (26%), Positives = 36/84 (42%), Gaps = 3/84 (3%)
Query: 36 IICPCLECCYTNVSANDLQDHLICNGIDKSYTCWTMHGEKK---TKSAKRRNERDPSTDF 92
++ C +C +T S LQ H+ + K Y C + E K + R+E S +F
Sbjct: 1204 VVFRCDKCTFTCSSDESLQQHIEKHNELKPYKCQLCYYETKHTEELDSHLRDEHKVSRNF 1263
Query: 93 EKDTNYEFDQVEEILNVIEDDLRD 116
E DQ+E++ +E D
Sbjct: 1264 ELVGRVNLDQLEQMKEKMESSSSD 1287
>RECG_HELPY (O26051) ATP-dependent DNA helicase recG (EC 3.6.1.-)
Length = 623
Score = 32.3 bits (72), Expect = 7.1
Identities = 19/59 (32%), Positives = 33/59 (55%), Gaps = 4/59 (6%)
Query: 578 PAAHTLSRKEKKIFCKFLHNVKVPEGYSSNIKSLVSMKDLKLKGLKSHDGHVLMGNFLP 636
P T K IF K ++ K+ E N++ L+S+++LK +G+K + H+L+ F P
Sbjct: 113 PKIITKFGKISLIFKKVKNHKKIQE----NLQKLISLENLKKEGVKENIAHLLLEIFFP 167
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.137 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 135,481,420
Number of Sequences: 164201
Number of extensions: 6072637
Number of successful extensions: 14092
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 14088
Number of HSP's gapped (non-prelim): 14
length of query: 1079
length of database: 59,974,054
effective HSP length: 121
effective length of query: 958
effective length of database: 40,105,733
effective search space: 38421292214
effective search space used: 38421292214
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 71 (32.0 bits)
Medicago: description of AC137666.7


