

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137546.15 + phase: 0 /pseudo
(355 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
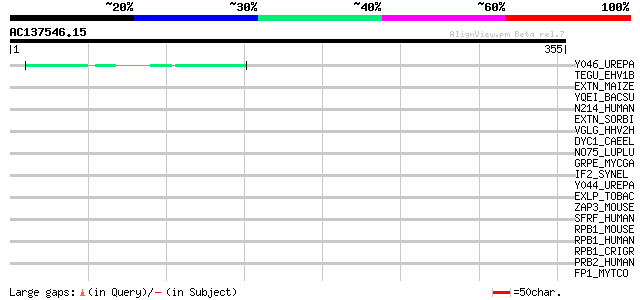
Score E
Sequences producing significant alignments: (bits) Value
Y046_UREPA (Q9PR99) Hypothetical protein UU046 44 5e-04
TEGU_EHV1B (P28955) Large tegument protein 43 0.001
EXTN_MAIZE (P14918) Extensin precursor (Proline-rich glycoprotein) 41 0.005
YQEI_BACSU (P54454) Hypothetical UPF0044 protein yqeI 40 0.009
N214_HUMAN (P35658) Nuclear pore complex protein Nup214 (Nucleop... 39 0.015
EXTN_SORBI (P24152) Extensin precursor (Proline-rich glycoprotein) 39 0.015
VGLG_HHV2H (P13290) Glycoprotein G 39 0.026
DYC1_CAEEL (Q8STF6) Dystrophin-like protein 1 (Dyb-1 binding and... 39 0.026
NO75_LUPLU (Q06841) Early nodulin 75 protein (N-75) (NGM-75) (Fr... 38 0.034
GRPE_MYCGA (Q7NBE4) GrpE protein (HSP-70 cofactor) 38 0.034
IF2_SYNEL (Q8DK04) Translation initiation factor IF-2 37 0.058
Y044_UREPA (Q9PRA1) Hypothetical protein UU044 37 0.076
EXLP_TOBAC (Q03211) Pistil-specific extensin-like protein precur... 37 0.076
ZAP3_MOUSE (Q9R0I7) Nuclear protein ZAP3 37 0.099
SFRF_HUMAN (O95104) Splicing factor, arginine/serine-rich 15 (CT... 36 0.13
RPB1_MOUSE (P08775) DNA-directed RNA polymerase II largest subun... 36 0.13
RPB1_HUMAN (P24928) DNA-directed RNA polymerase II largest subun... 36 0.13
RPB1_CRIGR (P11414) DNA-directed RNA polymerase II largest subun... 36 0.13
PRB2_HUMAN (P02812) Basic salivary proline-rich protein 2 (Saliv... 36 0.13
FP1_MYTCO (Q25434) Adhesive plaque matrix protein precursor (Foo... 36 0.17
>Y046_UREPA (Q9PR99) Hypothetical protein UU046
Length = 791
Score = 44.3 bits (103), Expect = 5e-04
Identities = 37/141 (26%), Positives = 54/141 (38%), Gaps = 26/141 (18%)
Query: 11 ISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDG 70
I+ N D RP+ K N PK PQ P + + P P+ P+ ++ +
Sbjct: 52 INKDNLDYQKARPSIKDNNLKEIPKPKPQPKPEPQPTPFP----DPIPTPPKKEELKK-- 105
Query: 71 VSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEF 130
P+ KP + ++P P P+ P+P P PP + K KE
Sbjct: 106 -------------------PEIKPEEPKKPEIKP-EPIPKPKPQPIPQPTPPVETKPKEE 145
Query: 131 DSFVLPPPHKKGVKPVQSPGP 151
PPP K+ KP P P
Sbjct: 146 LLPPSPPPPKEEPKPEPDPQP 166
>TEGU_EHV1B (P28955) Large tegument protein
Length = 3421
Score = 43.1 bits (100), Expect = 0.001
Identities = 45/164 (27%), Positives = 60/164 (36%), Gaps = 20/164 (12%)
Query: 28 NKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEFKYSYT 87
N P+ PK P K ++ + P P+ + S ++ G P FKY
Sbjct: 2480 NAMPAPPKTTPVVESKQKSDSVDRAPTLP----PKAAPLPPSDASAIMSGKPV-FKY--- 2531
Query: 88 ETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPH---KKGVK 144
TP +K PP VP P P P PLP S K LPP + K
Sbjct: 2532 -TPGNKSAV---PPSVPAPPTLPPAP-----PLPQSTSKAASGPPPTLPPAPPLPQSTSK 2582
Query: 145 PVQSPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVRSTLKSS 188
P P LP P + + G P T L +ST K++
Sbjct: 2583 AASGPPPTLPPAPPLPQSTSKAASGPPPTLPPAPPLPQSTSKAA 2626
>EXTN_MAIZE (P14918) Extensin precursor (Proline-rich glycoprotein)
Length = 267
Score = 40.8 bits (94), Expect = 0.005
Identities = 38/140 (27%), Positives = 47/140 (33%), Gaps = 10/140 (7%)
Query: 23 PTGKPNKNPSKPKVDPQSH---PALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAP 79
PT P+ P PK P ++ P + P K P TP + +P
Sbjct: 93 PTYTPSPKPPTPKPTPPTYTPSPKPPATKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSP 152
Query: 80 FEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRP-PLPPSKKKLKEFDSFVLPPP 138
TPK P P P P P P T P P PP+ K + PP
Sbjct: 153 ------KPPTPKPTPPTYTPSPKPPTHPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPP 206
Query: 139 HKKGVKPVQSPGPFLPGTSP 158
K P +P P P T P
Sbjct: 207 TPKPTPPTYTPSPKPPATKP 226
Score = 36.6 bits (83), Expect = 0.099
Identities = 37/139 (26%), Positives = 48/139 (33%), Gaps = 24/139 (17%)
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEF 82
PT P+ P PK P ++ P K TP S ++ P
Sbjct: 130 PTYTPSPKPPTPKPTPPTYT-------PSPKPPTPKPTPPTYTPSPKPPTHP---TPKPT 179
Query: 83 KYSYTETPKSKPVQMREPPFVPFG--PVTMPRPWTGRP-PLPPSKKKLKEFDSFVLPPPH 139
+YT +PK + P + P P P P T P P PP+ K PP
Sbjct: 180 PPTYTPSPKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPATK-----------PPT 228
Query: 140 KKGVKPVQSPGPFLPGTSP 158
K P +P P P T P
Sbjct: 229 PKPTPPTYTPTPKPPATKP 247
Score = 35.8 bits (81), Expect = 0.17
Identities = 40/158 (25%), Positives = 51/158 (31%), Gaps = 19/158 (12%)
Query: 5 LATTFPISASNADQSSRRPTGKPNKNPSKPKVDPQSH---PALKFSNIPKQKLKPVNKTP 61
+ F I ++ S PT P+ P PK P ++ P S P K P TP
Sbjct: 1 MCPAFSIFFNSRRYSLTPPTYTPSPKPPTPKPTPPTYTPSPKPPASKPPTPKPTPPTYTP 60
Query: 62 ENVKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRP-PL 120
P T TP KP + P P P P P P P
Sbjct: 61 S-------------PKPPTPKPTPPTYTPSPKPPATKPPTPKPTPPTYTPSPKPPTPKPT 107
Query: 121 PPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSP 158
PP+ + + PP K P +P P P P
Sbjct: 108 PPTYTPSPKPPA--TKPPTPKPTPPTYTPSPKPPTPKP 143
>YQEI_BACSU (P54454) Hypothetical UPF0044 protein yqeI
Length = 96
Score = 40.0 bits (92), Expect = 0.009
Identities = 21/54 (38%), Positives = 32/54 (58%), Gaps = 4/54 (7%)
Query: 290 LTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRV----NCQ 339
LT ++ +R K LTPI ++GK GV N++ + EA E EL++V NC+
Sbjct: 2 LTGKQKRFLRSKAHHLTPIFQVGKGGVNDNMIKQIAEALEARELIKVSVLQNCE 55
>N214_HUMAN (P35658) Nuclear pore complex protein Nup214
(Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein)
Length = 2090
Score = 39.3 bits (90), Expect = 0.015
Identities = 38/132 (28%), Positives = 53/132 (39%), Gaps = 22/132 (16%)
Query: 33 KPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEFKYSYTETPKS 92
KP ++ P++ NI + P + + V +SE +F + T TP S
Sbjct: 554 KPTLESTPVPSVSAPNIAMKSSFPPSTSAVKVNLSE------------KFTAAATSTPVS 601
Query: 93 KPVQMREPPFVPFGPVTMPR---PWTGRPPL--PPSKKKLKEFDSFVLPPPHKKGVKPVQ 147
PP PF + P P + PL PPS LK S VLP P + +
Sbjct: 602 S--SQSAPPMSPFSSASKPAASGPLSHPTPLSAPPSSVPLK---SSVLPSPSGRSAQGSS 656
Query: 148 SPGPFLPGTSPR 159
SP P + SPR
Sbjct: 657 SPVPSMVQKSPR 668
>EXTN_SORBI (P24152) Extensin precursor (Proline-rich glycoprotein)
Length = 283
Score = 39.3 bits (90), Expect = 0.015
Identities = 34/112 (30%), Positives = 41/112 (36%), Gaps = 21/112 (18%)
Query: 50 PKQKLKPVNKTPENVKISEDGVSYVIEGAPFEFKYS---YTETPKSKPVQMREPPFVPFG 106
P KP K P+ K G + E P E K + YT +PK P PP P
Sbjct: 35 PTPTPKPPAKGPKPEKPPTKGHGHKPEKPPKEHKPTPPTYTPSPKPTP-----PPATP-- 87
Query: 107 PVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSP 158
P PP+ + S V PPP K P +P P P T P
Sbjct: 88 -----------KPTPPTYTPSPKPKSPVYPPPPKASTPPTYTPSPKPPATKP 128
Score = 38.1 bits (87), Expect = 0.034
Identities = 38/133 (28%), Positives = 48/133 (35%), Gaps = 12/133 (9%)
Query: 26 KPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEFKYS 85
KP K P + K P ++ P KP TP S S V P +
Sbjct: 59 KPEKPPKEHKPTPPTYTPSPKPTPPPATPKP---TPPTYTPSPKPKSPVYPPPP-KASTP 114
Query: 86 YTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKP 145
T TP KP + P + P+P +PP PP + V PP K P
Sbjct: 115 PTYTPSPKPPATKPPTY------PTPKPPATKPPTPPVYTPSPKPP--VTKPPTPKPTPP 166
Query: 146 VQSPGPFLPGTSP 158
V +P P P T P
Sbjct: 167 VYTPNPKPPVTKP 179
Score = 37.4 bits (85), Expect = 0.058
Identities = 42/145 (28%), Positives = 51/145 (34%), Gaps = 16/145 (11%)
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEF 82
PT P+ P+ P P+ P ++ PK K PV P +Y P
Sbjct: 72 PTYTPSPKPTPPPATPKPTPPT-YTPSPKPK-SPVYPPPPKASTPP---TYTPSPKPPAT 126
Query: 83 KYSYTETPKSKPVQMREPPFV---PFGPVTMPRPWTGRPPL------PPSKKKLKEFDSF 133
K TPK + PP P PVT P PP+ PP K S
Sbjct: 127 KPPTYPTPKPPATKPPTPPVYTPSPKPPVTKPPTPKPTPPVYTPNPKPPVTKPPTHTPS- 185
Query: 134 VLPPPHKKGVKPVQSPGPFLPGTSP 158
PP K PV +P P P SP
Sbjct: 186 -PKPPTSKPTPPVYTPSPKPPKPSP 209
Score = 30.4 bits (67), Expect = 7.1
Identities = 41/154 (26%), Positives = 54/154 (34%), Gaps = 21/154 (13%)
Query: 8 TFPISASNADQSSRRPTGKPNKNP--SKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVK 65
T+P A + P P+ P +KP P+ P + N KP TP
Sbjct: 130 TYPTPKPPATKPPTPPVYTPSPKPPVTKPPT-PKPTPPVYTPNPKPPVTKPPTHTPSPKP 188
Query: 66 ISEDGVSYVIEGAPFEFKYSY-TETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSK 124
+ V +P K S T TP KP + P P T P+P T P P +
Sbjct: 189 PTSKPTPPVYTPSPKPPKPSPPTYTPTPKPPATKPPTSTP----THPKP-TPHTPYPQAH 243
Query: 125 KKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSP 158
PP +K KP P P P +P
Sbjct: 244 -----------PPTYKPAPKP-SPPAPTPPTYTP 265
>VGLG_HHV2H (P13290) Glycoprotein G
Length = 699
Score = 38.5 bits (88), Expect = 0.026
Identities = 33/126 (26%), Positives = 45/126 (35%), Gaps = 16/126 (12%)
Query: 11 ISASNADQSSRRPTGKPNKNP-------SKPKVDPQSHPALKFSNIPKQKL----KPVNK 59
+ A D SS PT P K P P VDP + P + P ++ V
Sbjct: 343 LMALTEDTSSDSPTSAPEKTPLPVSATAMAPSVDPSAEPTAPATTTPPDEMATQAATVAV 402
Query: 60 TPENVKISEDGVSYVIEGAPFEFKYSYT----ETPKSKPVQMREPPFVPFGPVTMPRPWT 115
TPE ++ + +E +P + T T S + PP P P T P T
Sbjct: 403 TPEETAVASPPATASVESSPLPAAAAATPGAGHTNTSSASAAKTPPTTP-APTTPPPTST 461
Query: 116 GRPPLP 121
P P
Sbjct: 462 HATPRP 467
>DYC1_CAEEL (Q8STF6) Dystrophin-like protein 1 (Dyb-1 binding and
CAPON-related protein)
Length = 887
Score = 38.5 bits (88), Expect = 0.026
Identities = 47/188 (25%), Positives = 74/188 (39%), Gaps = 32/188 (17%)
Query: 24 TGKPNKNPSKPKVDPQSHP--ALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFE 81
T P P P++ PQ P AL + P Q L + + N S+ + +
Sbjct: 269 TSHPILQPKAPELVPQLQPQTALPYQQKP-QSLLNIQQQQFNTLPSQMPSTQTLP----- 322
Query: 82 FKYSYTETPKSK--PVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFV-LPPP 138
S +E P P M PP +P+ T+P P T P +P ++ + V + P
Sbjct: 323 ---SLSENPGQSHIPRMMTMPPNMPYPTATLPHPRTWAPQIPSYPNSMQSLEQNVPMYYP 379
Query: 139 HKKGVKPVQSPGPFLPGTSPRYVMSREEVL---------------GEPLTKEEIN-ELVR 182
G+ P S PF G S ++S L G +T ++ N +L+R
Sbjct: 380 QMPGMLPSSSSLPF--GLSSPVMVSPYATLQLNMQSQQLDQPDHSGSQITMDQYNQQLMR 437
Query: 183 STLKSSRQ 190
S L ++Q
Sbjct: 438 SQLDQAQQ 445
>NO75_LUPLU (Q06841) Early nodulin 75 protein (N-75) (NGM-75)
(Fragment)
Length = 434
Score = 38.1 bits (87), Expect = 0.034
Identities = 34/126 (26%), Positives = 45/126 (34%), Gaps = 17/126 (13%)
Query: 21 RRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPF 80
R P P + P V P H P +K PV+ P E + E P
Sbjct: 263 RSPPVHPPSHEKPPFVYPPPHEKSPMHEPPYEKAPPVHLPPHEKPPIEYAPPH--EKPPV 320
Query: 81 EFKYSYTETPKSKPVQMREPPFVPFGPVTMP---RPWTGRPPL--PPSKKKLKEFDSFVL 135
E+ + + P P +PP P+ P +P PP PPS V
Sbjct: 321 EYPPPHVKPPVEYPSPAEKPPTHEKPPIEFPPHEKPPVYEPPYEKPPS----------VC 370
Query: 136 PPPHKK 141
PPPH+K
Sbjct: 371 PPPHEK 376
Score = 34.7 bits (78), Expect = 0.38
Identities = 34/130 (26%), Positives = 48/130 (36%), Gaps = 29/130 (22%)
Query: 33 KPKVD-PQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEFKYSYTETPK 91
KP ++ P H P +K PV+ P+ E P E+ + P
Sbjct: 178 KPPIEYPPPHEKPPVYEPPYEKPPPVHPPPD-------------EKPPIEYPPPLEKPPV 224
Query: 92 SKPVQMREPPFVP----FGPVTMPRPWTGRPPL--------PPSKKKLKEFDSFVLPPPH 139
+P + PP P P+ P P +PP+ PP E FV PPPH
Sbjct: 225 HEPPYEKPPPAQPPPHDKPPIEYPPPHE-KPPVYEPPYERSPPVHPPSHEKPPFVYPPPH 283
Query: 140 KKGVKPVQSP 149
+K P+ P
Sbjct: 284 EK--SPMHEP 291
>GRPE_MYCGA (Q7NBE4) GrpE protein (HSP-70 cofactor)
Length = 298
Score = 38.1 bits (87), Expect = 0.034
Identities = 35/125 (28%), Positives = 49/125 (39%), Gaps = 13/125 (10%)
Query: 35 KVDPQSHPALKFSNIPKQKLKPVNKTPENVKIS-EDGVSYVIEGAPFEFKYSYTETPKSK 93
K +PA + K K ++ + ++ DGV YV E S E P++
Sbjct: 177 KASDPDYPADHVCKVMKSGYKLYDRVVRHAMVAVSDGVGYV------EPASSEQEQPQTP 230
Query: 94 PVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQS--PGP 151
+ PP T P +P PP+ V PPH+K V+P QS PGP
Sbjct: 231 AKSTQTPP----AQETASAPAKTKPTAPPTNNLNANKPPVVSNPPHQKPVQPPQSNTPGP 286
Query: 152 FLPGT 156
P T
Sbjct: 287 KQPIT 291
>IF2_SYNEL (Q8DK04) Translation initiation factor IF-2
Length = 957
Score = 37.4 bits (85), Expect = 0.058
Identities = 27/93 (29%), Positives = 42/93 (45%), Gaps = 4/93 (4%)
Query: 87 TETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPV 146
+ETP++ Q+++PP P P RP PP P + K ++ + PP K P
Sbjct: 93 SETPEAPKPQLQKPPARPQPPQAPTRP---TPPAPVAPKPVEPVAAKPPAPPAKPEPTPP 149
Query: 147 QSPGPFLPGTSPRYVMSREEVLGEPLTKEEINE 179
+ P P L R E+V P ++E+ E
Sbjct: 150 R-PVPTLVPPPTRPTKKEEKVAATPPPRKELKE 181
>Y044_UREPA (Q9PRA1) Hypothetical protein UU044
Length = 782
Score = 37.0 bits (84), Expect = 0.076
Identities = 34/114 (29%), Positives = 48/114 (41%), Gaps = 15/114 (13%)
Query: 66 ISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKK 125
I++D + Y + P + E PK KP +P P P+ P PP K+
Sbjct: 52 INKDNLDYQ-KARPSIKDSNLKEIPKPKPQPKPKP---------QPTPFPDPIPTPPKKE 101
Query: 126 KLKEFDSFVLP-PPHKKGVKPVQSPGPFLPGTSPRYVMSREEVL--GEPLTKEE 176
+LK+ D + P P K +KP P P P +EE+L P KEE
Sbjct: 102 ELKKPD--IKPEEPKKPEIKPEPKPEPIPQPAPPIETKPKEELLPPNPPPPKEE 153
Score = 36.6 bits (83), Expect = 0.099
Identities = 35/141 (24%), Positives = 52/141 (36%), Gaps = 30/141 (21%)
Query: 11 ISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDG 70
I+ N D RP+ K + PK PQ P + + P P+ P+ ++ +
Sbjct: 52 INKDNLDYQKARPSIKDSNLKEIPKPKPQPKPKPQPTPFP----DPIPTPPKKEELKK-- 105
Query: 71 VSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEF 130
P KP + ++P P P+P P PP + K KE
Sbjct: 106 -------------------PDIKPEEPKKPEIKP-----EPKPEPIPQPAPPIETKPKEE 141
Query: 131 DSFVLPPPHKKGVKPVQSPGP 151
PPP K+ KP +P P
Sbjct: 142 LLPPNPPPPKEEPKPEPNPQP 162
>EXLP_TOBAC (Q03211) Pistil-specific extensin-like protein precursor
(PELP)
Length = 426
Score = 37.0 bits (84), Expect = 0.076
Identities = 23/67 (34%), Positives = 29/67 (42%), Gaps = 8/67 (11%)
Query: 88 ETPKSKPVQMREPPFVPF---GPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVK 144
+ P P + PP P P +P T +PP PP KK S +LPPP
Sbjct: 187 KAPSPSPAKQPPPPPPPVKAPSPSPATQPPTKQPPPPPRAKK-----SPLLPPPPPVAYP 241
Query: 145 PVQSPGP 151
PV +P P
Sbjct: 242 PVMTPSP 248
Score = 30.8 bits (68), Expect = 5.4
Identities = 22/75 (29%), Positives = 27/75 (35%), Gaps = 2/75 (2%)
Query: 85 SYTETPKSKPVQMREPPFVPFGPVTMPRPWTGR-PPLPPSKKKLKEFDSFVLPPPHKKGV 143
S + P+ P + PP P PV P P + PP PP K PP +
Sbjct: 164 SAKQPPQPPPAKQPSPPPPP-PPVKAPSPSPAKQPPPPPPPVKAPSPSPATQPPTKQPPP 222
Query: 144 KPVQSPGPFLPGTSP 158
P P LP P
Sbjct: 223 PPRAKKSPLLPPPPP 237
>ZAP3_MOUSE (Q9R0I7) Nuclear protein ZAP3
Length = 1386
Score = 36.6 bits (83), Expect = 0.099
Identities = 21/62 (33%), Positives = 27/62 (42%), Gaps = 6/62 (9%)
Query: 100 PPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPR 159
PPFVP+ + P P P LPPS V+PP + P P P +P + P
Sbjct: 482 PPFVPYSQMPPPLPTMPPPVLPPS------LPPPVMPPALPTSIPPPGMPPPVMPPSLPT 535
Query: 160 YV 161
V
Sbjct: 536 SV 537
Score = 31.2 bits (69), Expect = 4.2
Identities = 27/84 (32%), Positives = 31/84 (36%), Gaps = 13/84 (15%)
Query: 87 TETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPS----------KKKLKEFDSFVLP 136
T P P + P P P ++P P P +PPS L VLP
Sbjct: 496 TMPPPVLPPSLPPPVMPPALPTSIPPPGMPPPVMPPSLPTSVPPPGMPPSLSSGPPPVLP 555
Query: 137 PP--HKKGVKPVQSPGPFLPGTSP 158
PP G PV P P LPG P
Sbjct: 556 PPALSSAGSPPVLPP-PALPGGPP 578
>SFRF_HUMAN (O95104) Splicing factor, arginine/serine-rich 15
(CTD-binding SR-like protein RA4) (Fragment)
Length = 1157
Score = 36.2 bits (82), Expect = 0.13
Identities = 35/114 (30%), Positives = 54/114 (46%), Gaps = 17/114 (14%)
Query: 55 KPVNKTPENVKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPW 114
K + K PEN +++++G GA E ++ +P KP+ + PP P+T+P P
Sbjct: 641 KGIPKKPEN-EVAQNG------GA--ETSHTEPVSPIPKPLPVPVPPIPVPAPITVPPPQ 691
Query: 115 T-GRPPLPPSKKKLKEFDSF-----VLPPPHKKGVKPVQSPGPFL-PGTSPRYV 161
P PP L+ +F + PP GV P P PFL PG +P ++
Sbjct: 692 VPPHQPGPPVVGALQP-PAFTPPLGIPPPGFGPGVPPPPPPPPFLRPGFNPMHL 744
>RPB1_MOUSE (P08775) DNA-directed RNA polymerase II largest subunit
(EC 2.7.7.6) (RPB1)
Length = 1970
Score = 36.2 bits (82), Expect = 0.13
Identities = 43/160 (26%), Positives = 52/160 (31%), Gaps = 14/160 (8%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISED 69
P S QS P+ +PS P P S S P TP + K S
Sbjct: 1806 PSSPRYTPQSPTYTPSSPSYSPSSPSYSPTSPKYTPTS--PSYSPSSPEYTPASPKYSPT 1863
Query: 70 GVSYVIEGAPFEFKYSYTE------TPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPS 123
Y +P KYS T TPK P P P T P+ P P+
Sbjct: 1864 SPKY----SPTSPKYSPTSPTYSPTTPKYSPTSPTYSPTSPVYTPTSPKYSPTSPTYSPT 1919
Query: 124 KKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPRYVMS 163
K P K SPG + P TSP Y ++
Sbjct: 1920 SPKYSPTSPTYSPTSPKGSTYSPTSPG-YSP-TSPTYSLT 1957
Score = 33.5 bits (75), Expect = 0.84
Identities = 41/155 (26%), Positives = 54/155 (34%), Gaps = 13/155 (8%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISED 69
P S S + S P+ +PS P+ PQS S P + +P + K +
Sbjct: 1785 PTSPSYSPTSPSYSPTSPSYSPSSPRYTPQSPTYTPSS--PSYSPSSPSYSPTSPKYTPT 1842
Query: 70 GVSY---VIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKK 126
SY E P KYS T +PK P + P P T P+ P P+
Sbjct: 1843 SPSYSPSSPEYTPASPKYSPT-SPKYSPTSPKYSPTSPTYSPTTPKYSPTSPTYSPTS-- 1899
Query: 127 LKEFDSFVLPPPHKKGVKPVQSP-GPFLPGTSPRY 160
+ P P SP P TSP Y
Sbjct: 1900 ----PVYTPTSPKYSPTSPTYSPTSPKYSPTSPTY 1930
>RPB1_HUMAN (P24928) DNA-directed RNA polymerase II largest subunit
(EC 2.7.7.6) (RPB1)
Length = 1970
Score = 36.2 bits (82), Expect = 0.13
Identities = 43/160 (26%), Positives = 52/160 (31%), Gaps = 14/160 (8%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISED 69
P S QS P+ +PS P P S S P TP + K S
Sbjct: 1806 PSSPRYTPQSPTYTPSSPSYSPSSPSYSPTSPKYTPTS--PSYSPSSPEYTPTSPKYSPT 1863
Query: 70 GVSYVIEGAPFEFKYSYTE------TPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPS 123
Y +P KYS T TPK P P P T P+ P P+
Sbjct: 1864 SPKY----SPTSPKYSPTSPTYSPTTPKYSPTSPTYSPTSPVYTPTSPKYSPTSPTYSPT 1919
Query: 124 KKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPRYVMS 163
K P K SPG + P TSP Y ++
Sbjct: 1920 SPKYSPTSPTYSPTSPKGSTYSPTSPG-YSP-TSPTYSLT 1957
Score = 33.5 bits (75), Expect = 0.84
Identities = 41/155 (26%), Positives = 54/155 (34%), Gaps = 13/155 (8%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISED 69
P S S + S P+ +PS P+ PQS S P + +P + K +
Sbjct: 1785 PTSPSYSPTSPSYSPTSPSYSPSSPRYTPQSPTYTPSS--PSYSPSSPSYSPTSPKYTPT 1842
Query: 70 GVSY---VIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKK 126
SY E P KYS T +PK P + P P T P+ P P+
Sbjct: 1843 SPSYSPSSPEYTPTSPKYSPT-SPKYSPTSPKYSPTSPTYSPTTPKYSPTSPTYSPTS-- 1899
Query: 127 LKEFDSFVLPPPHKKGVKPVQSP-GPFLPGTSPRY 160
+ P P SP P TSP Y
Sbjct: 1900 ----PVYTPTSPKYSPTSPTYSPTSPKYSPTSPTY 1930
>RPB1_CRIGR (P11414) DNA-directed RNA polymerase II largest subunit
(EC 2.7.7.6) (RPB1) (Fragment)
Length = 467
Score = 36.2 bits (82), Expect = 0.13
Identities = 43/160 (26%), Positives = 52/160 (31%), Gaps = 14/160 (8%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISED 69
P S QS P+ +PS P P S S P TP + K S
Sbjct: 303 PSSPRYTPQSPTYTPSSPSYSPSSPSYSPTSPKYTPTS--PSYSPSSPEYTPTSPKYSPT 360
Query: 70 GVSYVIEGAPFEFKYSYTE------TPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPS 123
Y +P KYS T TPK P P P T P+ P P+
Sbjct: 361 SPKY----SPTSPKYSPTSPTYSPTTPKYSPTSPTYSPTSPVYTPTSPKYSPTSPTYSPT 416
Query: 124 KKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPRYVMS 163
K P K SPG + P TSP Y ++
Sbjct: 417 SPKYSPTSPTYSPTSPKGSTYSPTSPG-YSP-TSPTYSLT 454
Score = 33.5 bits (75), Expect = 0.84
Identities = 41/155 (26%), Positives = 54/155 (34%), Gaps = 13/155 (8%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISED 69
P S S + S P+ +PS P+ PQS S P + +P + K +
Sbjct: 282 PTSPSYSPTSPSYSPTSPSYSPSSPRYTPQSPTYTPSS--PSYSPSSPSYSPTSPKYTPT 339
Query: 70 GVSY---VIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKK 126
SY E P KYS T +PK P + P P T P+ P P+
Sbjct: 340 SPSYSPSSPEYTPTSPKYSPT-SPKYSPTSPKYSPTSPTYSPTTPKYSPTSPTYSPTS-- 396
Query: 127 LKEFDSFVLPPPHKKGVKPVQSP-GPFLPGTSPRY 160
+ P P SP P TSP Y
Sbjct: 397 ----PVYTPTSPKYSPTSPTYSPTSPKYSPTSPTY 427
>PRB2_HUMAN (P02812) Basic salivary proline-rich protein 2 (Salivary
proline-rich protein) (Con1 glycoprotein) [Contains:
Basic peptide P-F] (Fragment)
Length = 382
Score = 36.2 bits (82), Expect = 0.13
Identities = 42/150 (28%), Positives = 48/150 (32%), Gaps = 26/150 (17%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISED 69
P N QS+R P GKP P + PQ P KP P+ S
Sbjct: 191 PPQGDNKSQSARSPPGKPQGPPPQGGNQPQGP--------PPPPGKPQGPPPQGGNKS-- 240
Query: 70 GVSYVIEGAPFEFK-YSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLK 128
+G P K SK R PP P GP PP PP K +
Sbjct: 241 ------QGPPPPGKPQGPPPQGGSKSRSSRSPPGKPQGPPPQGGNQPQGPPPPPGKPQ-- 292
Query: 129 EFDSFVLPPPHKKGVKPVQSPGPFLPGTSP 158
PP + G KP P P P P
Sbjct: 293 -------GPPPQGGNKPQGPPPPGKPQGPP 315
Score = 31.2 bits (69), Expect = 4.2
Identities = 38/149 (25%), Positives = 43/149 (28%), Gaps = 25/149 (16%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISED 69
P N Q P GKP P + PQ P KP P+ +
Sbjct: 6 PPQGGNKPQGPPSPPGKPQGPPPQGGNQPQGPPP--------PPGKPQGPPPQGGNKPQ- 56
Query: 70 GVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKE 129
G P K K R PP P GP PP PP K +
Sbjct: 57 -------GPPPPGKPQGPPPQGDKSRSPRSPPGKPQGPPPQGGNQPQGPPPPPGKPQ--- 106
Query: 130 FDSFVLPPPHKKGVKPVQSPGPFLPGTSP 158
PP + G KP P P P P
Sbjct: 107 ------GPPPQGGNKPQGPPPPGKPQGPP 129
>FP1_MYTCO (Q25434) Adhesive plaque matrix protein precursor (Foot
protein 1) (MCFP1)
Length = 872
Score = 35.8 bits (81), Expect = 0.17
Identities = 36/132 (27%), Positives = 44/132 (33%), Gaps = 22/132 (16%)
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEF 82
PT KP KP P P + + P K KP TP K S + P +
Sbjct: 290 PTYKP-----KPSYPPTYKPKITYP--PTYKPKPSYPTPYKQKPSYPPIYKSKSSYPTSY 342
Query: 83 KYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKG 142
K T P KP +T P + +P PPS K K + P K
Sbjct: 343 KSKKTYPPTYKP------------KITYPPTYKPKPSYPPSYKPKKTYSPTYKP---KIT 387
Query: 143 VKPVQSPGPFLP 154
P P P P
Sbjct: 388 YPPTYKPKPSYP 399
Score = 30.8 bits (68), Expect = 5.4
Identities = 35/140 (25%), Positives = 47/140 (33%), Gaps = 27/140 (19%)
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEF 82
PT KP K P P + + P K KP TP K S + P +
Sbjct: 570 PTYKP-----KASYPPTYKPKITYP--PTYKPKPSYPTPYKQKPSYPPIYKSKSSYPTAY 622
Query: 83 KYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPP---PH 139
K T P KP +T P + +P PPS + PP P
Sbjct: 623 KSKKTYPPTYKP------------KITYPPTYKPKPSYPPSYR-----PKITYPPTYKPK 665
Query: 140 KKGVKPVQSPGPFLPGTSPR 159
K + +S G + P P+
Sbjct: 666 KSYPQAYKSKGSYPPSYQPK 685
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,991,409
Number of Sequences: 164201
Number of extensions: 2309037
Number of successful extensions: 7788
Number of sequences better than 10.0: 119
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 114
Number of HSP's that attempted gapping in prelim test: 7452
Number of HSP's gapped (non-prelim): 329
length of query: 355
length of database: 59,974,054
effective HSP length: 111
effective length of query: 244
effective length of database: 41,747,743
effective search space: 10186449292
effective search space used: 10186449292
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 66 (30.0 bits)
Medicago: description of AC137546.15


