

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137546.13 - phase: 0
(363 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
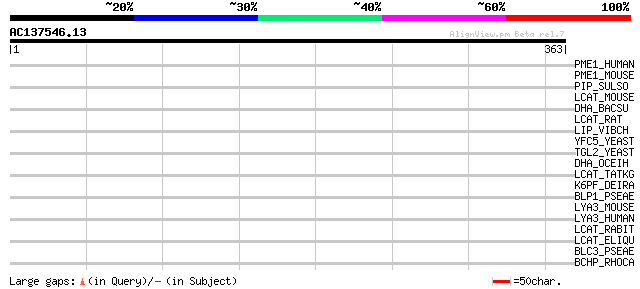
Score E
Sequences producing significant alignments: (bits) Value
PME1_HUMAN (Q9Y570) Protein phosphatase methylesterase 1 (EC 3.1... 39 0.027
PME1_MOUSE (Q8BVQ5) Protein phosphatase methylesterase 1 (EC 3.1... 38 0.035
PIP_SULSO (Q97UA2) Proline iminopeptidase (EC 3.4.11.5) (PIP) (P... 35 0.30
LCAT_MOUSE (P16301) Phosphatidylcholine-sterol acyltransferase p... 34 0.51
DHA_BACSU (Q08352) Alanine dehydrogenase (EC 1.4.1.1) (Stage V s... 34 0.66
LCAT_RAT (P18424) Phosphatidylcholine-sterol acyltransferase pre... 32 1.9
LIP_VIBCH (P15493) Lipase precursor (EC 3.1.1.3) (Triacylglycero... 32 2.5
YFC5_YEAST (P43571) Hypothetical 117.8 kDa protein in STE2-FRS2 ... 31 4.3
TGL2_YEAST (P54857) Lipase 2 (EC 3.1.1.3) (Triacylglycerol lipase) 31 5.6
DHA_OCEIH (Q8CX61) Alanine dehydrogenase (EC 1.4.1.1) 31 5.6
LCAT_TATKG (O35840) Phosphatidylcholine-sterol acyltransferase (... 30 7.3
K6PF_DEIRA (Q9RWN1) 6-phosphofructokinase (EC 2.7.1.11) (Phospho... 30 7.3
BLP1_PSEAE (Q03170) Beta-lactamase PSE-1 precursor (EC 3.5.2.6) ... 30 7.3
LYA3_MOUSE (Q8VEB4) 1-O-acylceramide synthase precursor (EC 2.3.... 30 9.6
LYA3_HUMAN (Q8NCC3) 1-O-acylceramide synthase precursor (EC 2.3.... 30 9.6
LCAT_RABIT (P53761) Phosphatidylcholine-sterol acyltransferase p... 30 9.6
LCAT_ELIQU (O35573) Phosphatidylcholine-sterol acyltransferase (... 30 9.6
BLC3_PSEAE (P37322) Beta-lactamase CARB-3 precursor (EC 3.5.2.6)... 30 9.6
BCHP_RHOCA (P26172) Geranylgeranyl hydrogenase 30 9.6
>PME1_HUMAN (Q9Y570) Protein phosphatase methylesterase 1 (EC
3.1.1.-) (PME-1) (PRO0750)
Length = 386
Score = 38.5 bits (88), Expect = 0.027
Identities = 32/104 (30%), Positives = 50/104 (47%), Gaps = 4/104 (3%)
Query: 97 EASVEHNAMEIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGKVAGLALVQ 156
+ S E A ++ +E +Y P+ML+GHS GG A A+ S+L + GL ++
Sbjct: 125 DLSAETMAKDVGNVVEAMYGDLPPPIMLIGHSMGG---AIAVHTASSNLVPSLLGLCMID 181
Query: 157 SPYGGTPIASDILREGQIGDKETRRILELIICKIIK-GDIRALE 199
G A + ++ G +T + LE I +K G IR LE
Sbjct: 182 VVEGTAMDALNSMQNFLRGRPKTFKSLENAIEWSVKSGQIRNLE 225
>PME1_MOUSE (Q8BVQ5) Protein phosphatase methylesterase 1 (EC
3.1.1.-) (PME-1)
Length = 386
Score = 38.1 bits (87), Expect = 0.035
Identities = 33/108 (30%), Positives = 51/108 (46%), Gaps = 4/108 (3%)
Query: 93 KVHSEASVEHNAMEIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGKVAGL 152
K + S E A ++ +E +Y PVML+GHS GG A A+ ++L + GL
Sbjct: 121 KNSEDLSAETMAKDVGNVVEAMYGDLPPPVMLIGHSMGG---AIAVHTAAANLVPSLLGL 177
Query: 153 ALVQSPYGGTPIASDILREGQIGDKETRRILELIICKIIK-GDIRALE 199
++ G A + ++ G +T + LE I +K G IR LE
Sbjct: 178 CMIDVVEGTAMDALNSMQNFLRGRPKTFKSLENAIEWSVKSGQIRNLE 225
>PIP_SULSO (Q97UA2) Proline iminopeptidase (EC 3.4.11.5) (PIP)
(Prolyl aminopeptidase) (PAP) (Tricorn protease
interacting factor F1)
Length = 310
Score = 35.0 bits (79), Expect = 0.30
Identities = 20/64 (31%), Positives = 36/64 (56%), Gaps = 2/64 (3%)
Query: 96 SEASVEHNAMEIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGKVAGLALV 155
S+ +++H E+++ ++++ G+ K ++LLGHS GG A A Y L+G + L
Sbjct: 86 SDYTIDHGLEELEELRKQVF-GNDK-IVLLGHSYGGALAIAYALKYQQFLRGLIVSSGLS 143
Query: 156 QSPY 159
PY
Sbjct: 144 SVPY 147
>LCAT_MOUSE (P16301) Phosphatidylcholine-sterol acyltransferase
precursor (EC 2.3.1.43) (Lecithin-cholesterol
acyltransferase) (Phospholipid-cholesterol
acyltransferase)
Length = 438
Score = 34.3 bits (77), Expect = 0.51
Identities = 23/65 (35%), Positives = 32/65 (48%), Gaps = 5/65 (7%)
Query: 111 IEEIYWGSGKPVMLLGHSKGGIDAAAAL---SLYWSDLKGKVAGLALVQSPYGGTPIASD 167
+EE+Y GKPV L+GHS G + L W D + G + +P+GG+ A
Sbjct: 188 VEEMYAAYGKPVFLIGHSLGCLHVLHFLLRQPQSWKD--HFIDGFISLGAPWGGSIKAMR 245
Query: 168 ILREG 172
IL G
Sbjct: 246 ILASG 250
>DHA_BACSU (Q08352) Alanine dehydrogenase (EC 1.4.1.1) (Stage V
sporulation protein N)
Length = 378
Score = 33.9 bits (76), Expect = 0.66
Identities = 28/101 (27%), Positives = 45/101 (43%), Gaps = 6/101 (5%)
Query: 208 DFIMKHKLPLDIPLISFRSEASITPSVLATMTQIA-HAELPRLILPKFGSKVSDQFVESG 266
+ +MK K PL + FR VL T +A EL + + K + ++ + V G
Sbjct: 68 EMVMKVKEPLPEEYVYFRKGL-----VLFTYLHLAAEPELAQALKDKGVTAIAYETVSEG 122
Query: 267 RQVPVMVPVSAAMAAFALHLQLRYGEKSDGVVTCRDAEVPG 307
R +P++ P+S A + ++ EK G A VPG
Sbjct: 123 RTLPLLTPMSEVAGRMAAQIGAQFLEKPKGGKGILLAGVPG 163
>LCAT_RAT (P18424) Phosphatidylcholine-sterol acyltransferase
precursor (EC 2.3.1.43) (Lecithin-cholesterol
acyltransferase) (Phospholipid-cholesterol
acyltransferase)
Length = 440
Score = 32.3 bits (72), Expect = 1.9
Identities = 22/65 (33%), Positives = 31/65 (46%), Gaps = 5/65 (7%)
Query: 111 IEEIYWGSGKPVMLLGHSKGGIDAAAAL---SLYWSDLKGKVAGLALVQSPYGGTPIASD 167
+EE+Y GKPV L+GHS G + L W D + G + +P+GG+
Sbjct: 188 VEEMYAAYGKPVFLIGHSLGCLHVLHFLLRQPQSWKD--HFIDGFISLGAPWGGSIKPMR 245
Query: 168 ILREG 172
IL G
Sbjct: 246 ILASG 250
>LIP_VIBCH (P15493) Lipase precursor (EC 3.1.1.3) (Triacylglycerol
lipase)
Length = 312
Score = 32.0 bits (71), Expect = 2.5
Identities = 28/121 (23%), Positives = 51/121 (42%), Gaps = 5/121 (4%)
Query: 61 LLIPGLFSNH---GPLYFVATKRFFSKMGLACHIAKVHSEASVEHNAMEIKQYIEEIYWG 117
+L+ GLF G YF + ++ G ++A+V + S E ++ +E +
Sbjct: 39 VLVHGLFGFDTLAGMDYFHGIPQSLTRDGAQVYVAQVSATNSSERRGEQLLAQVESLLAV 98
Query: 118 SG-KPVMLLGHSKGGIDAAAALSLYWSDLKGKVAGLALVQSPYGGTPIASDILREGQIGD 176
+G K V L+GHS GG S+ DL V + V + ++ G + +
Sbjct: 99 TGAKKVNLIGHSHGGPTIRYVASVR-PDLVASVTSIGGVHKGSAVADLVRGVIPSGSVSE 157
Query: 177 K 177
+
Sbjct: 158 Q 158
>YFC5_YEAST (P43571) Hypothetical 117.8 kDa protein in STE2-FRS2
intergenic region
Length = 1029
Score = 31.2 bits (69), Expect = 4.3
Identities = 14/46 (30%), Positives = 28/46 (60%), Gaps = 1/46 (2%)
Query: 122 VMLLGHSKGGIDAAAALSLYWSDLKGKVAGLALVQSPYGGTPIASD 167
V+++GHS GGI + L+L + + G ++ + + SP+ +P+ D
Sbjct: 230 VIIVGHSMGGIVSRVMLTLK-NHVPGSISTILTLSSPHAASPVTFD 274
>TGL2_YEAST (P54857) Lipase 2 (EC 3.1.1.3) (Triacylglycerol lipase)
Length = 326
Score = 30.8 bits (68), Expect = 5.6
Identities = 24/104 (23%), Positives = 46/104 (44%), Gaps = 12/104 (11%)
Query: 74 YFVATKRFFSKMGLACHIAKVHSEASVEHNAMEI-KQYIEEIYWGSGK----PVMLLGHS 128
Y++ K+F G KV S+E AM + Q +E+ K + L+ HS
Sbjct: 85 YWIGVKKFLQSKGCTVITTKVPGFGSIEERAMALDAQLQKEVKKIESKDKRHSLNLIAHS 144
Query: 129 KGGIDAAAALSLYWSDLKGK---VAGLALVQSPYGGTPIASDIL 169
GG+D + ++K + + L + +P+ G+ +A ++
Sbjct: 145 MGGLDCRYLI----CNIKNRNYDILSLTTISTPHRGSEMADYVV 184
>DHA_OCEIH (Q8CX61) Alanine dehydrogenase (EC 1.4.1.1)
Length = 376
Score = 30.8 bits (68), Expect = 5.6
Identities = 24/90 (26%), Positives = 41/90 (44%), Gaps = 6/90 (6%)
Query: 208 DFIMKHKLPLDIPLISFRSEASITPSVLATMTQIAHA-ELPRLILPKFGSKVSDQFVESG 266
D IMK K PL FR +L T +A A EL + ++ + ++ + +
Sbjct: 69 DMIMKVKEPLSSEYKYFRKGL-----ILFTYLHLAAAPELTKALVDSEVTAIAYETITVN 123
Query: 267 RQVPVMVPVSAAMAAFALHLQLRYGEKSDG 296
+P++ P+S A + +Y EKS+G
Sbjct: 124 GTLPLLTPMSEVAGRMATQIGAQYLEKSEG 153
>LCAT_TATKG (O35840) Phosphatidylcholine-sterol acyltransferase (EC
2.3.1.43) (Lecithin-cholesterol acyltransferase)
(Phospholipid-cholesterol acyltransferase) (Fragment)
Length = 293
Score = 30.4 bits (67), Expect = 7.3
Identities = 13/20 (65%), Positives = 15/20 (75%)
Query: 111 IEEIYWGSGKPVMLLGHSKG 130
IEE+Y GKPV L+GHS G
Sbjct: 106 IEEMYAAYGKPVFLIGHSLG 125
>K6PF_DEIRA (Q9RWN1) 6-phosphofructokinase (EC 2.7.1.11)
(Phosphofructokinase) (Phosphohexokinase)
Length = 329
Score = 30.4 bits (67), Expect = 7.3
Identities = 23/83 (27%), Positives = 43/83 (51%), Gaps = 6/83 (7%)
Query: 130 GGIDAAAALSLYWSDLKGKVAGLALVQSPYGGTPIASDILREGQIGDKETRRILE----L 185
GG +A A + L+ +V+ L +Q GGTP++SD + ++G+ +++ +
Sbjct: 235 GGAEAVAQAIHAGTGLETRVSILGHIQR--GGTPVSSDRILASRLGEAAVYALMDGKKGV 292
Query: 186 IICKIIKGDIRALEDLTYEKRKD 208
++ +I G T+EKRKD
Sbjct: 293 MVGRIGGGIAYTPLHETWEKRKD 315
>BLP1_PSEAE (Q03170) Beta-lactamase PSE-1 precursor (EC 3.5.2.6)
(Beta-lactamase CARB-2) (Carbenicillinase 2)
Length = 288
Score = 30.4 bits (67), Expect = 7.3
Identities = 40/152 (26%), Positives = 62/152 (40%), Gaps = 17/152 (11%)
Query: 154 LVQSPYGGTPIASDILREGQIGDKETRRILELIICKIIKGDIRALEDLTYEKR-----KD 208
++ S GG +D LR QIGDKETR L+ I + +G + L D T K
Sbjct: 132 IILSAVGGPKGVTDFLR--QIGDKETR--LDRIEPDLNEGKLGDLRDTTTPKAIASTLNK 187
Query: 209 FIMKHKLP--LDIPLISFRSEASITPSVLATMTQIAHAELPRLILPKFGSKVSDQFVESG 266
F+ L L S+ +T ++L ++ R FG++ V S
Sbjct: 188 FLFGSALSEMNQKKLESWMVNNQVTGNLLRSVLPAGWNIADRSGAGGFGARSITAVVWSE 247
Query: 267 RQVPVMVPVSAAMAAFALHLQLRYGEKSDGVV 298
Q P++V + A Q E++D +V
Sbjct: 248 HQAPIIVSIYLAQT------QASMAERNDAIV 273
>LYA3_MOUSE (Q8VEB4) 1-O-acylceramide synthase precursor (EC
2.3.1.-) (ACS) (Lysosomal phospholipase A2) (LCAT-like
lysophospholipase) (LLPL)
Length = 412
Score = 30.0 bits (66), Expect = 9.6
Identities = 19/69 (27%), Positives = 32/69 (45%), Gaps = 1/69 (1%)
Query: 105 MEIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGK-VAGLALVQSPYGGTP 163
+ +++ IEE+Y G PV+L+ HS G + L K K + + +P+GG
Sbjct: 175 LALREMIEEMYQMYGGPVVLVAHSMGNVYMLYFLQRQPQVWKDKYIHAFVSLGAPWGGVA 234
Query: 164 IASDILREG 172
+L G
Sbjct: 235 KTLRVLASG 243
>LYA3_HUMAN (Q8NCC3) 1-O-acylceramide synthase precursor (EC
2.3.1.-) (ACS) (Lysosomal phospholipase A2) (LCAT-like
lysophospholipase) (LLPL) (UNQ341/PRO540)
Length = 412
Score = 30.0 bits (66), Expect = 9.6
Identities = 19/69 (27%), Positives = 32/69 (45%), Gaps = 1/69 (1%)
Query: 105 MEIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGK-VAGLALVQSPYGGTP 163
+ +++ IEE+Y G PV+L+ HS G + L K K + + +P+GG
Sbjct: 175 LALREMIEEMYQLYGGPVVLVAHSMGNMYTLYFLQRQPQAWKDKYIRAFVSLGAPWGGVA 234
Query: 164 IASDILREG 172
+L G
Sbjct: 235 KTLRVLASG 243
>LCAT_RABIT (P53761) Phosphatidylcholine-sterol acyltransferase
precursor (EC 2.3.1.43) (Lecithin-cholesterol
acyltransferase) (Phospholipid-cholesterol
acyltransferase)
Length = 440
Score = 30.0 bits (66), Expect = 9.6
Identities = 17/53 (32%), Positives = 28/53 (52%), Gaps = 1/53 (1%)
Query: 111 IEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGK-VAGLALVQSPYGGT 162
+EE++ GKPV L+GHS G + L K + + G + +P+GG+
Sbjct: 188 VEEMHAAYGKPVFLIGHSLGCLHLLYFLLRQPQSWKDRFIDGFISLGAPWGGS 240
>LCAT_ELIQU (O35573) Phosphatidylcholine-sterol acyltransferase (EC
2.3.1.43) (Lecithin-cholesterol acyltransferase)
(Phospholipid-cholesterol acyltransferase) (Fragment)
Length = 299
Score = 30.0 bits (66), Expect = 9.6
Identities = 13/30 (43%), Positives = 19/30 (63%)
Query: 101 EHNAMEIKQYIEEIYWGSGKPVMLLGHSKG 130
E +++ +EE+Y GKPV L+GHS G
Sbjct: 100 EEYYLKLAGLVEEMYATYGKPVFLIGHSLG 129
>BLC3_PSEAE (P37322) Beta-lactamase CARB-3 precursor (EC 3.5.2.6)
(Carbenicillinase 3)
Length = 288
Score = 30.0 bits (66), Expect = 9.6
Identities = 41/156 (26%), Positives = 64/156 (40%), Gaps = 25/156 (16%)
Query: 154 LVQSPYGGTPIASDILREGQIGDKETRRILELIICKIIKGDIRALEDLTYEKRKDFIMKH 213
++ S GG +D LR QIGDKETR L+ I + +G + L D T K +
Sbjct: 132 IILSAVGGPKGVTDFLR--QIGDKETR--LDRIEPDLNEGKLGDLRDTTTPKAIASTLNK 187
Query: 214 KL-----------PLDIPLISFRSEASITPSVLATMTQIAHAELPRLILPKFGSKVSDQF 262
L L+ +++ + ++ SVL IA R FG++
Sbjct: 188 LLFGSALSEMNQKKLESWMVNNQVTGNLLRSVLPAGWNIA----DRSGAGGFGARSITAV 243
Query: 263 VESGRQVPVMVPVSAAMAAFALHLQLRYGEKSDGVV 298
V S Q P++V + A Q E++D +V
Sbjct: 244 VWSEHQAPIIVSIYLAQT------QASMAERNDAIV 273
>BCHP_RHOCA (P26172) Geranylgeranyl hydrogenase
Length = 391
Score = 30.0 bits (66), Expect = 9.6
Identities = 28/128 (21%), Positives = 55/128 (42%), Gaps = 12/128 (9%)
Query: 116 WGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGKVAGLALVQSPYGGTPIASDILREGQIG 175
W +GK V+L G + G + ++ +Y++ G+ A A + G P + R G +
Sbjct: 256 WDNGKDVVLSGDAAGVVAPSSGEGIYYAHAGGRYAAEAAMAFLKSGKPADLKLARAGFMK 315
Query: 176 DKET-RRILELIICKIIKGDIRA-----------LEDLTYEKRKDFIMKHKLPLDIPLIS 223
+ T ++L ++ K D R ++ +T+E + M PL I
Sbjct: 316 EHGTVFKVLRMMQDKYYHSDDRRERFVSLCHDVDVQRMTFESYMNKKMTKFQPLKNLKIG 375
Query: 224 FRSEASIT 231
F++ A ++
Sbjct: 376 FKNLAHLS 383
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,741,633
Number of Sequences: 164201
Number of extensions: 1753191
Number of successful extensions: 3990
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 3982
Number of HSP's gapped (non-prelim): 21
length of query: 363
length of database: 59,974,054
effective HSP length: 111
effective length of query: 252
effective length of database: 41,747,743
effective search space: 10520431236
effective search space used: 10520431236
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC137546.13


