

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137079.13 + phase: 0
(357 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
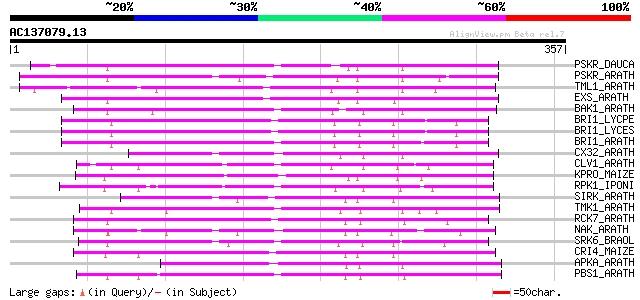
Sequences producing significant alignments: (bits) Value
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 145 2e-34
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 135 2e-31
TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor 134 3e-31
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 134 3e-31
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 134 5e-31
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 131 3e-30
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 130 5e-30
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 124 3e-28
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 121 2e-27
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 120 4e-27
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 116 1e-25
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 115 2e-25
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 112 2e-24
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 111 2e-24
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 111 3e-24
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 110 7e-24
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 109 1e-23
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 108 2e-23
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 108 3e-23
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 107 6e-23
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 145 bits (366), Expect = 2e-34
Identities = 103/327 (31%), Positives = 170/327 (51%), Gaps = 38/327 (11%)
Query: 14 PMKKATSEGRLKGGDNNNSELVFFVED-HERFKLEDLLRATADLRSENF-----WSSLFK 67
P KKA ++ G + S ++F +D + L+D+L++T+ N + ++K
Sbjct: 703 PEKKADADEIELG---SRSVVLFHNKDSNNELSLDDILKSTSSFNQANIIGCGGFGLVYK 759
Query: 68 VKFENNVEYAVKRLKNLQVSCD-EFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQ 126
+ + A+KRL D EF+ ++ +S+ +H N++ L+GY + K +KL+IY Y
Sbjct: 760 ATLPDGTKVAIKRLSGDTGQMDREFQAEVETLSRAQHPNLVHLLGYCNYKNDKLLIYSYM 819
Query: 127 SNGSVLNLLNDYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNI 186
NGS+ L++ + WK RL IA G A GLA++++ E I H ++K SNI
Sbjct: 820 DNGSLDYWLHEKVDGPPSLDWKTRLRIARGAAEGLAYLHQSCEP----HILHRDIKSSNI 875
Query: 187 LLDDKNEALISEHGLSKFFEPDRGTFFSSH---------GYTAPE----KSLTEKGDVYS 233
LL D A +++ GL++ P + +H GY PE T KGDVYS
Sbjct: 876 LLSDTFVAHLADFGLARLILP-----YDTHVTTDLVGTLGYIPPEYGQASVATYKGDVYS 930
Query: 234 FGVILLELLTG-QSIEVSR----IDLVRWVRSMVREEWTGEVFDKEVRENDH-QGAFSLL 287
FGV+LLELLTG + ++V + DL+ WV M E+ E+FD + + DH + +L
Sbjct: 931 FGVVLLELLTGRRPMDVCKPRGSRDLISWVLQMKTEKRESEIFDPFIYDKDHAEEMLLVL 990
Query: 288 NIALMCVSRSQENRPNFGEILETIEGV 314
IA C+ + + RP +++ +E +
Sbjct: 991 EIACRCLGENPKTRPTTQQLVSWLENI 1017
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 135 bits (340), Expect = 2e-31
Identities = 100/333 (30%), Positives = 170/333 (51%), Gaps = 34/333 (10%)
Query: 7 KKNNLDSPMKKATSEGRLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENF----- 61
+ +D ++++ S R + G+ + +V F + + +DLL +T N
Sbjct: 685 RSGEVDPEIEESESMNRKELGEIGSKLVVLFQSNDKELSYDDLLDSTNSFDQANIIGCGG 744
Query: 62 WSSLFKVKFENNVEYAVKRLKNLQVSCD-EFREILKQISKVKHQNILSLVGYRSTKEEKL 120
+ ++K + + A+K+L + EF ++ +S+ +H N++ L G+ K ++L
Sbjct: 745 FGMVYKATLPDGKKVAIKKLSGDCGQIEREFEAEVETLSRAQHPNLVLLRGFCFYKNDRL 804
Query: 121 IIYKYQSNGSVLNLLNDYIARRKDFP----WKLRLNIACGIARGLAFIYKKLEEGEVNSI 176
+IY Y NGS L+ ++ R D P WK RL IA G A+GL +++ EG I
Sbjct: 805 LIYSYMENGS----LDYWLHERNDGPALLKWKTRLRIAQGAAKGLLYLH----EGCDPHI 856
Query: 177 PHGNLKLSNILLDDKNEALISEHGLSKFFEPDR----GTFFSSHGYTAPE----KSLTEK 228
H ++K SNILLD+ + +++ GL++ P + GY PE T K
Sbjct: 857 LHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVSTDLVGTLGYIPPEYGQASVATYK 916
Query: 229 GDVYSFGVILLELLTG-QSIEVSR----IDLVRWVRSMVREEWTGEVFDKEV--RENDHQ 281
GDVYSFGV+LLELLT + +++ + DL+ WV M E EVFD + +END +
Sbjct: 917 GDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISWVVKMKHESRASEVFDPLIYSKENDKE 976
Query: 282 GAFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
F +L IA +C+S + + RP +++ ++ V
Sbjct: 977 -MFRVLEIACLCLSENPKQRPTTQQLVSWLDDV 1008
>TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor
Length = 674
Score = 134 bits (338), Expect = 3e-31
Identities = 98/330 (29%), Positives = 172/330 (51%), Gaps = 31/330 (9%)
Query: 7 KKNNLDSP--MKKATSEGRLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSS 64
+K++++S +++ E + + +LV F + E L+D+L AT + + + +
Sbjct: 328 RKSSIESEDDLEEGDEEDEIGEKEGGEGKLVVF-QGGENLTLDDVLNATGQVMEKTSYGT 386
Query: 65 LFKVKFENNVEYAVKRLKNLQVSCDEFRE---ILKQISKVKHQNILSLVG-YRSTKEEKL 120
++K K + A++ L+ + +C + +++Q+ +++H+N++ L Y+ + EKL
Sbjct: 387 VYKAKLSDGGNIALRLLR--EGTCKDRSSCLPVIRQLGRIRHENLVPLRAFYQGKRGEKL 444
Query: 121 IIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGN 180
+IY Y N S+ +LL++ R+ W R IA GIARGLA+ L G+ I HGN
Sbjct: 445 LIYDYLPNISLHDLLHESKPRKPALNWARRHKIALGIARGLAY----LHTGQEVPIIHGN 500
Query: 181 LKLSNILLDDKNEALISEHGLSKFF----EPDRGTFFSSHGYTAPE----KSLTEKGDVY 232
++ N+L+DD A ++E GL K + + S GY APE K + DVY
Sbjct: 501 IRSKNVLVDDFFFARLTEFGLDKIMVQAVADEIVSQAKSDGYKAPELHKMKKCNPRSDVY 560
Query: 233 SFGVILLELLTGQSIEVSR------IDLVRWVRSMVREEWTGEVFD----KEVRENDHQG 282
+FG++LLE+L G+ S +DL V++ V EE T EVFD K +R +G
Sbjct: 561 AFGILLLEILMGKKPGKSGRNGNEFVDLPSLVKAAVLEETTMEVFDLEAMKGIRSPMEEG 620
Query: 283 AFSLLNIALMCVSRSQENRPNFGEILETIE 312
L +A+ C + RP+ E+++ +E
Sbjct: 621 LVHALKLAMGCCAPVTTVRPSMEEVVKQLE 650
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 134 bits (338), Expect = 3e-31
Identities = 96/302 (31%), Positives = 149/302 (48%), Gaps = 25/302 (8%)
Query: 34 LVFFVEDHERFKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVSC 88
+ F + + +L D++ AT +N + +++K AVK+L +
Sbjct: 895 IAMFEQPLLKVRLGDIVEATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQG 954
Query: 89 D-EFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPW 147
+ EF ++ + KVKH N++SL+GY S EEKL++Y+Y NGS+ + L + + W
Sbjct: 955 NREFMAEMETLGKVKHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDW 1014
Query: 148 KLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEP 207
RL IA G ARGLAF L G + I H ++K SNILLD E +++ GL++
Sbjct: 1015 SKRLKIAVGAARGLAF----LHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISA 1070
Query: 208 DRG----TFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQS------IEVSRID 253
+ GY PE T KGDVYSFGVILLEL+TG+ E +
Sbjct: 1071 CESHVSTVIAGTFGYIPPEYGQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGGN 1130
Query: 254 LVRWVRSMVREEWTGEVFDK-EVRENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIE 312
LV W + + +V D V LL IA++C++ + RPN ++L+ ++
Sbjct: 1131 LVGWAIQKINQGKAVDVIDPLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKALK 1190
Query: 313 GV 314
+
Sbjct: 1191 EI 1192
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 134 bits (336), Expect = 5e-31
Identities = 93/299 (31%), Positives = 159/299 (53%), Gaps = 35/299 (11%)
Query: 42 ERFKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVSCDE--FREI 94
+RF L +L A+ + ++N + ++K + + AVKRLK + E F+
Sbjct: 275 KRFSLRELQVASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERTQGGELQFQTE 334
Query: 95 LKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIA 154
++ IS H+N+L L G+ T E+L++Y Y +NGSV + L + + W R IA
Sbjct: 335 VEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPPLDWPKRQRIA 394
Query: 155 CGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFE-------- 206
G ARGLA+++ + I H ++K +NILLD++ EA++ + GL+K +
Sbjct: 395 LGSARGLAYLHDHCDP----KIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTT 450
Query: 207 PDRGTFFSSHGYTAPEKSLT----EKGDVYSFGVILLELLTGQ-SIEVSR------IDLV 255
RGT G+ APE T EK DV+ +GV+LLEL+TGQ + +++R + L+
Sbjct: 451 AVRGTI----GHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLL 506
Query: 256 RWVRSMVREEWTGEVFDKEVREN-DHQGAFSLLNIALMCVSRSQENRPNFGEILETIEG 313
WV+ +++E+ + D +++ N + L+ +AL+C S RP E++ +EG
Sbjct: 507 DWVKGLLKEKKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRMLEG 565
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 131 bits (329), Expect = 3e-30
Identities = 92/297 (30%), Positives = 152/297 (50%), Gaps = 27/297 (9%)
Query: 34 LVFFVEDHERFKLEDLLRATADLRSENFWSS-----LFKVKFENNVEYAVKRLKNLQVSC 88
L F + + DLL AT +++ S ++K + ++ A+K+L ++
Sbjct: 866 LAAFEKPLRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQG 925
Query: 89 D-EFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPW 147
D EF ++ I K+KH+N++ L+GY EE+L++Y+Y GS+ ++L+D W
Sbjct: 926 DREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNW 985
Query: 148 KLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEP 207
R IA G ARGLAF++ + I H ++K SN+LLD+ EA +S+ G+++
Sbjct: 986 PARRKIAIGAARGLAFLHHNC----IPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSA 1041
Query: 208 -----DRGTFFSSHGYTAPEK----SLTEKGDVYSFGVILLELLTGQ----SIEVSRIDL 254
T + GY PE + KGDVYS+GV+LLELLTG+ S + +L
Sbjct: 1042 MDTHLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNL 1101
Query: 255 VRWVRSMVREEWTGEVFDKEVRENDHQGAFSL---LNIALMCVSRSQENRPNFGEIL 308
V WV+ + + T +VFD+E+ + D L L +A C+ RP +++
Sbjct: 1102 VGWVKLHAKGKIT-DVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVM 1157
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 130 bits (327), Expect = 5e-30
Identities = 92/297 (30%), Positives = 152/297 (50%), Gaps = 27/297 (9%)
Query: 34 LVFFVEDHERFKLEDLLRATADLRSENFWSS-----LFKVKFENNVEYAVKRLKNLQVSC 88
L F + + DLL AT +++ S ++K + ++ A+K+L ++
Sbjct: 866 LAAFEKPLRKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQG 925
Query: 89 D-EFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPW 147
D EF ++ I K+KH+N++ L+GY EE+L++Y+Y GS+ ++L+D W
Sbjct: 926 DREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNW 985
Query: 148 KLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEP 207
R IA G ARGLAF++ + I H ++K SN+LLD+ EA +S+ G+++
Sbjct: 986 PARRKIAIGAARGLAFLHHNC----IPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSA 1041
Query: 208 -----DRGTFFSSHGYTAPEK----SLTEKGDVYSFGVILLELLTGQ----SIEVSRIDL 254
T + GY PE + KGDVYS+GV+LLELLTG+ S + +L
Sbjct: 1042 MDTHLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNL 1101
Query: 255 VRWVRSMVREEWTGEVFDKEVRENDHQGAFSL---LNIALMCVSRSQENRPNFGEIL 308
V WV+ + + T +VFD+E+ + D L L +A C+ RP +++
Sbjct: 1102 VGWVKLHAKGKIT-DVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVM 1157
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 124 bits (312), Expect = 3e-28
Identities = 89/297 (29%), Positives = 149/297 (49%), Gaps = 27/297 (9%)
Query: 34 LVFFVEDHERFKLEDLLRATADLRSENFWSS-----LFKVKFENNVEYAVKRLKNLQVSC 88
L F + + DLL+AT +++ S ++K ++ A+K+L ++
Sbjct: 861 LAAFEKPLRKLTFADLLQATNGFHNDSLIGSGGFGDVYKAILKDGSAVAIKKLIHVSGQG 920
Query: 89 D-EFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPW 147
D EF ++ I K+KH+N++ L+GY +E+L++Y++ GS+ ++L+D W
Sbjct: 921 DREFMAEMETIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNW 980
Query: 148 KLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEP 207
R IA G ARGLAF++ I H ++K SN+LLD+ EA +S+ G+++
Sbjct: 981 STRRKIAIGSARGLAFLHHNCSP----HIIHRDMKSSNVLLDENLEARVSDFGMARLMSA 1036
Query: 208 -----DRGTFFSSHGYTAPEK----SLTEKGDVYSFGVILLELLTGQ----SIEVSRIDL 254
T + GY PE + KGDVYS+GV+LLELLTG+ S + +L
Sbjct: 1037 MDTHLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDSPDFGDNNL 1096
Query: 255 VRWVRSMVREEWTGEVFDKEVRENDHQGAFSL---LNIALMCVSRSQENRPNFGEIL 308
V WV+ + +VFD E+ + D L L +A+ C+ RP +++
Sbjct: 1097 VGWVKQHAKLR-ISDVFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVM 1152
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 121 bits (304), Expect = 2e-27
Identities = 81/256 (31%), Positives = 136/256 (52%), Gaps = 27/256 (10%)
Query: 77 AVKRLKNLQVS-CDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLL 135
A+KRL + V E+R + + + H+N++ L+GY +E L++Y++ GS L
Sbjct: 122 AIKRLNSESVQGFAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGS----L 177
Query: 136 NDYIARRKD-FPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEA 194
++ RR D FPW LR+ I G ARGLAF++ E + + + K SNILLD +A
Sbjct: 178 ESHLFRRNDPFPWDLRIKIVIGAARGLAFLHSLQRE-----VIYRDFKASNILLDSNYDA 232
Query: 195 LISEHGLSKFFEPDRGT-----FFSSHGYTAPEKSLT----EKGDVYSFGVILLELLTGQ 245
+S+ GL+K D + ++GY APE T K DV++FGV+LLE++TG
Sbjct: 233 KLSDFGLAKLGPADEKSHVTTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGL 292
Query: 246 SIEVSR-----IDLVRWVR-SMVREEWTGEVFDKEVR-ENDHQGAFSLLNIALMCVSRSQ 298
+ ++ LV W+R + + ++ DK ++ + + A + I L C+
Sbjct: 293 TAHNTKRPRGQESLVDWLRPELSNKHRVKQIMDKGIKGQYTTKVATEMARITLSCIEPDP 352
Query: 299 ENRPNFGEILETIEGV 314
+NRP+ E++E +E +
Sbjct: 353 KNRPHMKEVVEVLEHI 368
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 120 bits (302), Expect = 4e-27
Identities = 90/295 (30%), Positives = 146/295 (48%), Gaps = 37/295 (12%)
Query: 44 FKLEDLLRATADLRSENFWSS-----LFKVKFENNVEYAVKRL--KNLQVSCDEFREILK 96
FK ED+L L+ EN +++ NNV+ A+KRL + S F ++
Sbjct: 683 FKSEDVLEC---LKEENIIGKGGAGIVYRGSMPNNVDVAIKRLVGRGTGRSDHGFTAEIQ 739
Query: 97 QISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACG 156
+ +++H++I+ L+GY + K+ L++Y+Y NGS+ LL+ ++ W+ R +A
Sbjct: 740 TLGRIRHRHIVRLLGYVANKDTNLLLYEYMPNGSLGELLHG--SKGGHLQWETRHRVAVE 797
Query: 157 IARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFF-----EPDRGT 211
A+GL +++ I H ++K +NILLD EA +++ GL+KF +
Sbjct: 798 AAKGLCYLHHDCSP----LILHRDVKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSS 853
Query: 212 FFSSHGYTAPEKSLT----EKGDVYSFGVILLELLTGQSIE---VSRIDLVRWVRSMVRE 264
S+GY APE + T EK DVYSFGV+LLEL+ G+ +D+VRWVR+ E
Sbjct: 854 IAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIAGKKPVGEFGEGVDIVRWVRN-TEE 912
Query: 265 EWTG--------EVFDKEVRENDHQGAFSLLNIALMCVSRSQENRPNFGEILETI 311
E T + D + + IA+MCV RP E++ +
Sbjct: 913 EITQPSDAAIVVAIVDPRLTGYPLTSVIHVFKIAMMCVEEEAAARPTMREVVHML 967
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1
precursor (EC 2.7.1.37)
Length = 817
Score = 116 bits (290), Expect = 1e-25
Identities = 88/295 (29%), Positives = 148/295 (49%), Gaps = 31/295 (10%)
Query: 43 RFKLEDLLRATADLRSE---NFWSSLFKVKFENNVEYAVKRLKNLQVSCDEFREILKQIS 99
R+ +L++AT + E +++K E++ AVK+L+N++ + F+ L I
Sbjct: 523 RYSYRELVKATRKFKVELGRGESGTVYKGVLEDDRHVAVKKLENVRQGKEVFQAELSVIG 582
Query: 100 KVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGIAR 159
++ H N++ + G+ S +L++ +Y NGS+ N+L W+ R NIA G+A+
Sbjct: 583 RINHMNLVRIWGFCSEGSHRLLVSEYVENGSLANILFSE-GGNILLDWEGRFNIALGVAK 641
Query: 160 GLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSSH--- 216
GLA+++ + E + H ++K NILLD E I++ GL K T SH
Sbjct: 642 GLAYLHHECLEWVI----HCDVKPENILLDQAFEPKITDFGLVKLLNRGGSTQNVSHVRG 697
Query: 217 --GYTAPE----KSLTEKGDVYSFGVILLELLTGQSIE--VSRIDLVR-WVRSMVR---- 263
GY APE +T K DVYS+GV+LLELLTG + V D V +R +VR
Sbjct: 698 TLGYIAPEWVSSLPITAKVDVYSYGVVLLELLTGTRVSELVGGTDEVHSMLRKLVRMLSA 757
Query: 264 ------EEWTGEVFDKEV-RENDHQGAFSLLNIALMCVSRSQENRPNFGEILETI 311
+ W D ++ R ++ A +L+ +A+ C+ + RP ++T+
Sbjct: 758 KLEGEEQSWIDGYLDSKLNRPVNYVQARTLIKLAVSCLEEDRSKRPTMEHAVQTL 812
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 115 bits (287), Expect = 2e-25
Identities = 94/310 (30%), Positives = 156/310 (50%), Gaps = 41/310 (13%)
Query: 33 ELVFFVEDHERFKLEDLLRATADLRSE-----NFWSSLFKVKFENNVEYAVKRL-----K 82
E+ ++ + L +L AT +L + +++K + YAVK+L K
Sbjct: 793 EIAISAQEGDGSLLNKVLEATENLNDKYVIGKGAHGTIYKATLSPDKVYAVKKLVFTGIK 852
Query: 83 NLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARR 142
N VS REI + I KV+H+N++ L + KE LI+Y Y NGS+ ++L++
Sbjct: 853 NGSVSM--VREI-ETIGKVRHRNLIKLEEFWLRKEYGLILYTYMENGSLHDILHE-TNPP 908
Query: 143 KDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLS 202
K W R NIA G A GLA+++ + +I H ++K NILLD E IS+ G++
Sbjct: 909 KPLDWSTRHNIAVGTAHGLAYLHFDCDP----AIVHRDIKPMNILLDSDLEPHISDFGIA 964
Query: 203 KFFEPD-----RGTFFSSHGYTAPEKSLT----EKGDVYSFGVILLELLT-GQSIEVS-- 250
K + T + GY APE + T + DVYS+GV+LLEL+T ++++ S
Sbjct: 965 KLLDQSATSIPSNTVQGTIGYMAPENAFTTVKSRESDVYSYGVVLLELITRKKALDPSFN 1024
Query: 251 -RIDLVRWVRSMVREEWTGEV--------FDKEVRENDHQGAFSLLNIALMCVSRSQENR 301
D+V WVRS+ + TGE+ D+ + + + L++AL C + + R
Sbjct: 1025 GETDIVGWVRSVWTQ--TGEIQKIVDPSLLDELIDSSVMEQVTEALSLALRCAEKEVDKR 1082
Query: 302 PNFGEILETI 311
P ++++ +
Sbjct: 1083 PTMRDVVKQL 1092
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 112 bits (279), Expect = 2e-24
Identities = 81/261 (31%), Positives = 136/261 (52%), Gaps = 25/261 (9%)
Query: 72 NNVEYAVKRLKNLQVS-CDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGS 130
N + AVK L EFR + + +V H N+ SLVGY + ++IY+Y +N +
Sbjct: 594 NGEQVAVKVLSEESAQGYKEFRAEVDLLMRVHHTNLTSLVGYCNEINHMVLIYEYMANEN 653
Query: 131 VLNLLNDYIARRKDF--PWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILL 188
L DY+A ++ F W+ RL I+ A+GL +++ G I H ++K +NILL
Sbjct: 654 ----LGDYLAGKRSFILSWEERLKISLDAAQGLEYLHN----GCKPPIVHRDVKPTNILL 705
Query: 189 DDKNEALISEHGLSKFFEPDRGTFFS-----SHGYTAPE----KSLTEKGDVYSFGVILL 239
++K +A +++ GLS+ F + S S GY PE + + EK DVYS GV+LL
Sbjct: 706 NEKLQAKMADFGLSRSFSVEGSGQISTVVAGSIGYLDPEYYSTRQMNEKSDVYSLGVVLL 765
Query: 240 ELLTGQ----SIEVSRIDLVRWVRSMVREEWTGEVFDKEVREN-DHQGAFSLLNIALMCV 294
E++TGQ S + ++ + VRS++ + D+ +RE D A+ + IAL C
Sbjct: 766 EVITGQPAIASSKTEKVHISDHVRSILANGDIRGIVDQRLRERYDVGSAWKMSEIALACT 825
Query: 295 SRSQENRPNFGEILETIEGVM 315
+ RP +++ ++ ++
Sbjct: 826 EHTSAQRPTMSQVVMELKQIV 846
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 111 bits (278), Expect = 2e-24
Identities = 86/296 (29%), Positives = 152/296 (51%), Gaps = 30/296 (10%)
Query: 46 LEDLLRATADLRSENFWSS-----LFKVKFENNVEYAVKRLKNLQVSCDEFREILKQIS- 99
++ L T + S+N S ++K + + + AVKR++N ++ F E +I+
Sbjct: 578 IQVLRSVTNNFSSDNILGSGGFGVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIAV 637
Query: 100 --KVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIAR-RKDFPWKLRLNIACG 156
KV+H+++++L+GY EKL++Y+Y G++ L ++ K WK RL +A
Sbjct: 638 LTKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQRLTLALD 697
Query: 157 IARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGT----F 212
+ARG+ +++ + S H +LK SNILL D A +++ GL + +G+
Sbjct: 698 VARGVEYLHGLAHQ----SFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIETRI 753
Query: 213 FSSHGYTAPEKS----LTEKGDVYSFGVILLELLTG-QSIEVSR----IDLVRWVRSMV- 262
+ GY APE + +T K DVYSFGVIL+EL+TG +S++ S+ I LV W + M
Sbjct: 754 AGTFGYLAPEYAVTGRVTTKVDVYSFGVILMELITGRKSLDESQPEESIHLVSWFKRMYI 813
Query: 263 -REEWTGEVFDK--EVRENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGVM 315
+E + D ++ E ++ +A C +R RP+ G + + ++
Sbjct: 814 NKEASFKKAIDTTIDLDEETLASVHTVAELAGHCCAREPYQRPDMGHAVNILSSLV 869
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 111 bits (277), Expect = 3e-24
Identities = 82/290 (28%), Positives = 150/290 (51%), Gaps = 27/290 (9%)
Query: 42 ERFKLEDLLRATADLRSENF-----WSSLFKVKFEN-NVEYAVKRL-KNLQVSCDEFREI 94
+ F ++L AT + RS+ F + +FK E + A+K+L +N EF
Sbjct: 89 QTFTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNGVQGIREFVVE 148
Query: 95 LKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIA 154
+ +S H N++ L+G+ + +++L++Y+Y GS+ + L+ + +K W R+ IA
Sbjct: 149 VLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRMKIA 208
Query: 155 CGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFS 214
G ARGL +++ ++ + + +LK SNILL + + +S+ GL+K T S
Sbjct: 209 AGAARGLEYLHDRM----TPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVS 264
Query: 215 -----SHGYTAPEKS----LTEKGDVYSFGVILLELLTG-QSIEVSRI----DLVRWVRS 260
++GY AP+ + LT K D+YSFGV+LLEL+TG ++I+ ++ +LV W R
Sbjct: 265 TRVMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNTKTRKDQNLVGWARP 324
Query: 261 MVREEWTGEVFDKEVRENDH--QGAFSLLNIALMCVSRSQENRPNFGEIL 308
+ ++ + + + +G + L I+ MCV RP +++
Sbjct: 325 LFKDRRNFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDVV 374
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 110 bits (274), Expect = 7e-24
Identities = 98/313 (31%), Positives = 152/313 (48%), Gaps = 57/313 (18%)
Query: 42 ERFKLEDLLRATADLRSENF------------W---SSLFKVKFENNVEYAVKRLKNLQV 86
+ F L +L AT + R ++ W SSL K + AVKRL Q
Sbjct: 54 KNFSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRLN--QE 111
Query: 87 SCDEFREILKQIS---KVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRK 143
RE L +I+ ++ H N++ L+GY +E +L++Y++ + GS L +++ RR
Sbjct: 112 GFQGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGS----LENHLFRRG 167
Query: 144 DF----PWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEH 199
F W R+ +A G ARGLAF++ + + + + K SNILLD A +S+
Sbjct: 168 TFYQPLSWNTRVRMALGAARGLAFLHNAQPQ-----VIYRDFKASNILLDSNYNAKLSDF 222
Query: 200 GLSKFFEPDRGTFFS-----SHGYTAPE----KSLTEKGDVYSFGVILLELLTG-----Q 245
GL++ + S + GY APE L+ K DVYSFGV+LLELL+G +
Sbjct: 223 GLARDGPMGDNSHVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDK 282
Query: 246 SIEVSRIDLVRWVRSMVREEWTGEVFDKEVRENDHQGAFSLLN------IALMCVSRSQE 299
+ V +LV W R + T + V + QG +SL +AL C+S +
Sbjct: 283 NQPVGEHNLVDWARPYL----TNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDAK 338
Query: 300 NRPNFGEILETIE 312
+RP EI++T+E
Sbjct: 339 SRPTMNEIVKTME 351
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 109 bits (272), Expect = 1e-23
Identities = 86/296 (29%), Positives = 151/296 (50%), Gaps = 41/296 (13%)
Query: 45 KLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVS-CDEFREILKQI 98
++E +++AT + S N + ++K + + E AVKRL V DEF + I
Sbjct: 517 EMETVVKATENFSSCNKLGQGGFGIVYKGRLLDGKEIAVKRLSKTSVQGTDEFMNEVTLI 576
Query: 99 SKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYI---ARRKDFPWKLRLNIAC 155
++++H N++ ++G +EK++IY+Y N S L+ Y+ RR W R +I
Sbjct: 577 ARLQHINLVQVLGCCIEGDEKMLIYEYLENLS----LDSYLFGKTRRSKLNWNERFDITN 632
Query: 156 GIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRG----- 210
G+ARGL ++++ I H +LK+SNILLD IS+ G+++ FE D
Sbjct: 633 GVARGLLYLHQDSRF----RIIHRDLKVSNILLDKNMIPKISDFGMARIFERDETEANTM 688
Query: 211 TFFSSHGYTAPEKSL----TEKGDVYSFGVILLELLTGQ------SIEVSRIDLVRWVRS 260
++GY +PE ++ +EK DV+SFGVI+LE+++G+ +++ DL+ +V S
Sbjct: 689 KVVGTYGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYEN-DLLSYVWS 747
Query: 261 MVREEWTGEVFDKEVREN--------DHQGAFSLLNIALMCVSRSQENRPNFGEIL 308
+E E+ D + ++ Q + I L+CV E+RP ++
Sbjct: 748 RWKEGRALEIVDPVIVDSLSSQPSIFQPQEVLKCIQIGLLCVQELAEHRPAMSSVV 803
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 108 bits (270), Expect = 2e-23
Identities = 80/293 (27%), Positives = 147/293 (49%), Gaps = 26/293 (8%)
Query: 42 ERFKLEDLLRATADLRSEN-----FWSSLFKVKFENNVEYAVKRL---KNLQVSCDEFRE 93
+ F E+L +AT ++ +S +FK + AVKR +++ S EF
Sbjct: 491 QEFSYEELEQATGGFSEDSQVGKGSFSCVFKGILRDGTVVAVKRAIKASDVKKSSKEFHN 550
Query: 94 ILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIAR-RKDFPWKLRLN 152
L +S++ H ++L+L+GY E+L++Y++ ++GS+ L+ +K W R+
Sbjct: 551 ELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLKKRLNWARRVT 610
Query: 153 IACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTF 212
IA ARG+ +++ + H ++K SNIL+D+ + A +++ GLS D GT
Sbjct: 611 IAVQAARGIEYLHGY----ACPPVIHRDIKSSNILIDEDHNARVADFGLSILGPADSGTP 666
Query: 213 FS-----SHGYTAPE----KSLTEKGDVYSFGVILLELLTGQ---SIEVSRIDLVRWVRS 260
S + GY PE LT K DVYSFGV+LLE+L+G+ ++ ++V W
Sbjct: 667 LSELPAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQFEEGNIVEWAVP 726
Query: 261 MVREEWTGEVFDKEVR-ENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIE 312
+++ + D + +D + + ++A CV ++RP+ ++ +E
Sbjct: 727 LIKAGDIFAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPSMDKVTTALE 779
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 108 bits (269), Expect = 3e-23
Identities = 73/235 (31%), Positives = 126/235 (53%), Gaps = 21/235 (8%)
Query: 98 ISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGI 157
+ + H++++ L+GY E +L++Y++ GS+ N L + WKLRL +A G
Sbjct: 126 LGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGLYFQPLSWKLRLKVALGA 185
Query: 158 ARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSS-- 215
A+GLAF++ + + + K SNILLD + A +S+ GL+K + S+
Sbjct: 186 AKGLAFLHSS-----ETRVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPIGDKSHVSTRV 240
Query: 216 ---HGYTAPEK----SLTEKGDVYSFGVILLELLTG-QSIEVSR----IDLVRWVRS-MV 262
HGY APE LT K DVYSFGV+LLELL+G ++++ +R +LV W + +V
Sbjct: 241 MGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDKNRPSGERNLVEWAKPYLV 300
Query: 263 REEWTGEVFDKEVREN-DHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGVMN 316
+ V D +++ + A + ++L C++ + RPN E++ +E + +
Sbjct: 301 NKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKLRPNMSEVVSHLEHIQS 355
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 107 bits (266), Expect = 6e-23
Identities = 81/297 (27%), Positives = 150/297 (50%), Gaps = 29/297 (9%)
Query: 44 FKLEDLLRATADLRSENF-----WSSLFKVKFENNVEY-AVKRL--KNLQVSCDEFREIL 95
F +L AT + + F + ++K + ++ + AVK+L LQ + + E+L
Sbjct: 74 FAFRELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQLDRNGLQGNREFLVEVL 133
Query: 96 KQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIAC 155
+S + H N+++L+GY + +++L++Y++ GS+ + L+D ++ W +R+ IA
Sbjct: 134 -MLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAA 192
Query: 156 GIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFS- 214
G A+GL F++ K + + + K SNILLD+ +S+ GL+K + S
Sbjct: 193 GAAKGLEFLHDKANP----PVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVST 248
Query: 215 ----SHGYTAPEKS----LTEKGDVYSFGVILLELLTGQSIEVSRI-----DLVRWVRSM 261
++GY APE + LT K DVYSFGV+ LEL+TG+ S + +LV W R +
Sbjct: 249 RVMGTYGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAWARPL 308
Query: 262 VREEWTG-EVFDKEVREN-DHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGVMN 316
+ ++ D ++ + + L +A MC+ RP +++ + + N
Sbjct: 309 FNDRRKFIKLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADVVTALSYLAN 365
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.135 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,793,442
Number of Sequences: 164201
Number of extensions: 1718961
Number of successful extensions: 6794
Number of sequences better than 10.0: 1181
Number of HSP's better than 10.0 without gapping: 108
Number of HSP's successfully gapped in prelim test: 1073
Number of HSP's that attempted gapping in prelim test: 5702
Number of HSP's gapped (non-prelim): 1312
length of query: 357
length of database: 59,974,054
effective HSP length: 111
effective length of query: 246
effective length of database: 41,747,743
effective search space: 10269944778
effective search space used: 10269944778
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 66 (30.0 bits)
Medicago: description of AC137079.13


