

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137077.10 + phase: 0
(247 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
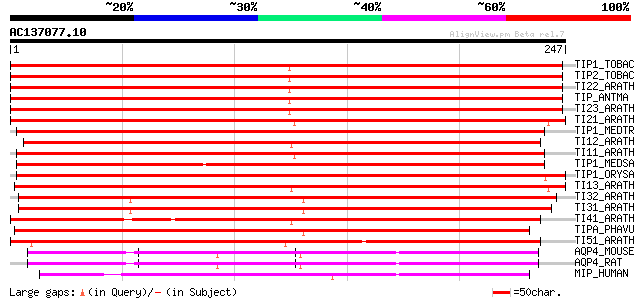
Score E
Sequences producing significant alignments: (bits) Value
TIP1_TOBAC (P21653) Probable aquaporin TIP-type RB7-5A (Tonoplas... 410 e-114
TIP2_TOBAC (P24422) Probable aquaporin TIP-type RB7-18C (Tonopla... 408 e-114
TI22_ARATH (Q41975) Probable aquaporin TIP2.2 (Tonoplast intrins... 405 e-113
TIP_ANTMA (P33560) Probable aquaporin TIP-type (Tonoplast intrin... 401 e-112
TI23_ARATH (Q9FGL2) Probable aquaporin TIP2.3 (Tonoplast intrins... 390 e-108
TI21_ARATH (Q41951) Aquaporin TIP2.1 (Tonoplast intrinsic protei... 375 e-104
TIP1_MEDTR (Q9FY14) Probable aquaporin TIP-type (MtAQP1) 307 2e-83
TI12_ARATH (Q41963) Aquaporin TIP1.2 (Tonoplast intrinsic protei... 305 9e-83
TI11_ARATH (P25818) Aquaporin TIP1.1 (Tonoplast intrinsic protei... 302 4e-82
TIP1_MEDSA (P42067) Probable aquaporin TIP-type (Membrane channe... 299 5e-81
TIP1_ORYSA (P50156) Probable aquaporin TIP-type 1 (Tonoplast int... 290 2e-78
TI13_ARATH (O82598) Putative aquaporin TIP1.3 (Tonoplast intrins... 278 7e-75
TI32_ARATH (O22588) Probable aquaporin TIP3.2 (Tonoplast intrins... 260 2e-69
TI31_ARATH (P26587) Aquaporin TIP3.1 (Tonoplast intrinsic protei... 258 1e-68
TI41_ARATH (O82316) Probable aquaporin TIP4.1 (Tonoplast intrins... 256 3e-68
TIPA_PHAVU (P23958) Probable aquaporin TIP-type alpha (Tonoplast... 238 8e-63
TI51_ARATH (Q9STX9) Putative aquaporin TIP5.1 (Tonoplast intrins... 209 4e-54
AQP4_MOUSE (P55088) Aquaporin 4 (WCH4) (Mercurial-insensitive wa... 164 2e-40
AQP4_RAT (P47863) Aquaporin 4 (WCH4) (Mercurial-insensitive wate... 157 3e-38
MIP_HUMAN (P30301) Lens fiber major intrinsic protein (MIP26) (M... 155 1e-37
>TIP1_TOBAC (P21653) Probable aquaporin TIP-type RB7-5A (Tonoplast
intrinsic protein, root-specific RB7-5A) (TobRB7)
(RT-TIP)
Length = 250
Score = 410 bits (1055), Expect = e-114
Identities = 206/247 (83%), Positives = 223/247 (89%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MV+I FG+ DSFS SLKAY++EFIATL+FVFAGVGSAIAYN LT+DAALDPAGLVAVA
Sbjct: 1 MVRIAFGSIGDSFSVGSLKAYVAEFIATLLFVFAGVGSAIAYNKLTADAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV+IAANISGGHLNPAVT GLA+GGNITILTG FYWIAQLLGS VA LLL
Sbjct: 61 VAHAFALFVGVSIAANISGGHLNPAVTLGLAVGGNITILTGFFYWIAQLLGSTVACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTHGVAAGLN + G+V EIIITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 YVTNGLAVPTHGVAAGLNGLQGVVMEIIITFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVV+G+F+ NWIYW GPLIGGGLAG IYGDVFIG +
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVAGDFSQNWIYWAGPLIGGGLAGFIYGDVFIGCH 240
Query: 240 TPAPASE 246
TP P SE
Sbjct: 241 TPLPTSE 247
>TIP2_TOBAC (P24422) Probable aquaporin TIP-type RB7-18C (Tonoplast
intrinsic protein, root-specific RB7-18C) (TobRB7)
(RT-TIP)
Length = 250
Score = 408 bits (1049), Expect = e-114
Identities = 206/247 (83%), Positives = 221/247 (89%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MV I FG+ DSFS SLKAY++EFIATL+FVFAGVGSAIAYN LT+DAALDPAGLVAVA
Sbjct: 1 MVLIAFGSIGDSFSVGSLKAYVAEFIATLLFVFAGVGSAIAYNKLTADAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV+IAANISGGHLNPAVT GLA+GGNITILTG FYWIAQLLGS VA LLL
Sbjct: 61 VAHAFALFVGVSIAANISGGHLNPAVTLGLAVGGNITILTGFFYWIAQLLGSTVACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTHGVAAGLN G+V EIIITF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 YVTNGLAVPTHGVAAGLNGFQGVVMEIIITFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVV+G+F+ NWIYW GPLIGGGLAG IYGDVFIG +
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVAGDFSQNWIYWAGPLIGGGLAGFIYGDVFIGCH 240
Query: 240 TPAPASE 246
TP P SE
Sbjct: 241 TPLPTSE 247
>TI22_ARATH (Q41975) Probable aquaporin TIP2.2 (Tonoplast intrinsic
protein 2.2)
Length = 250
Score = 405 bits (1041), Expect = e-113
Identities = 204/247 (82%), Positives = 223/247 (89%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVKI G+ DSFS ASLKAYLSEFIATL+FVFAGVGSA+A+ LTSDAALDPAGLVAVA
Sbjct: 1 MVKIEIGSVGDSFSVASLKAYLSEFIATLLFVFAGVGSALAFAKLTSDAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV+IAANISGGHLNPAVT GLA+GGNIT++TG FYWIAQ LGSIVA LLL
Sbjct: 61 VAHAFALFVGVSIAANISGGHLNPAVTLGLAVGGNITVITGFFYWIAQCLGSIVACLLLV 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT +SVPTHGVAAGL I G+V EI++TF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTNGESVPTHGVAAGLGAIEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVVSG+F+ WIYWVGPL+GG LAGLIYGDVFIGSY
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVSGDFSQIWIYWVGPLVGGALAGLIYGDVFIGSY 240
Query: 240 TPAPASE 246
PAP +E
Sbjct: 241 APAPTTE 247
>TIP_ANTMA (P33560) Probable aquaporin TIP-type (Tonoplast intrinsic
protein DiP) (Dark intrinsic protein)
Length = 250
Score = 401 bits (1030), Expect = e-112
Identities = 201/247 (81%), Positives = 220/247 (88%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVKI FG+ DSFS AS+KAY++EFIATL+FVFAGVGSAIAYN LTSDAALDPAGLVAVA
Sbjct: 1 MVKIAFGSIGDSFSVASIKAYVAEFIATLLFVFAGVGSAIAYNKLTSDAALDPAGLVAVA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAFALFVGV++AAN+SGGHLNPAVT GLA+GGNITILTGLFYWIAQ LGS VA LLL
Sbjct: 61 VAHAFALFVGVSMAANVSGGHLNPAVTLGLAVGGNITILTGLFYWIAQCLGSTVACLLLK 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT SVPTHGVAAG++ I G+V EIIITF LVYTVYATAADPKKGS+G IAPIAIGF+
Sbjct: 121 FVTNGLSVPTHGVAAGMDAIQGVVMEIIITFALVYTVYATAADPKKGSLGVIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAV SG+F+ NWIYW GPLIGG LAG IYGDVFI ++
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVASGDFSQNWIYWAGPLIGGALAGFIYGDVFITAH 240
Query: 240 TPAPASE 246
P P SE
Sbjct: 241 APLPTSE 247
>TI23_ARATH (Q9FGL2) Probable aquaporin TIP2.3 (Tonoplast intrinsic
protein 2.3)
Length = 250
Score = 390 bits (1003), Expect = e-108
Identities = 196/247 (79%), Positives = 217/247 (87%), Gaps = 1/247 (0%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
MVKI G+ DSFS +SLKAYLSEFIATL+FVFAGVGSA+A+ LTSD ALDPAGLVA+A
Sbjct: 1 MVKIEVGSVGDSFSVSSLKAYLSEFIATLLFVFAGVGSAVAFAKLTSDGALDPAGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
+AHAFALFVGV+IAANISGGHLNPAVT GLAIGGNIT++TG FYWIAQ LGSIVA LLL
Sbjct: 61 IAHAFALFVGVSIAANISGGHLNPAVTLGLAIGGNITLITGFFYWIAQCLGSIVACLLLV 120
Query: 121 YVT-AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
+VT KSVPTHGV+AGL + G+V EI++TF LVYTVYATAADPKKGS+GTIAPIAIGF+
Sbjct: 121 FVTNGKSVPTHGVSAGLGAVEGVVMEIVVTFALVYTVYATAADPKKGSLGTIAPIAIGFI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSY 239
VGANILAAGPFSGGSMNPARSFGPAVVSG+ + WIYWVGPL+GG LAGLIYGDVFIGSY
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVVSGDLSQIWIYWVGPLVGGALAGLIYGDVFIGSY 240
Query: 240 TPAPASE 246
E
Sbjct: 241 EAVETRE 247
>TI21_ARATH (Q41951) Aquaporin TIP2.1 (Tonoplast intrinsic protein
2.1) (Delta-tonoplast intrinsic protein) (Delta-TIP)
Length = 250
Score = 375 bits (964), Expect = e-104
Identities = 187/250 (74%), Positives = 215/250 (85%), Gaps = 3/250 (1%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
M + FG+FDDSFS ASL+AYL+EFI+TL+FVFAGVGSAIAY LTSDAALD GLVA+A
Sbjct: 1 MAGVAFGSFDDSFSLASLRAYLAEFISTLLFVFAGVGSAIAYAKLTSDAALDTPGLVAIA 60
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
V H FALFV VAI ANISGGH+NPAVTFGLA+GG IT++TG+FYWIAQLLGS A LL
Sbjct: 61 VCHGFALFVAVAIGANISGGHVNPAVTFGLAVGGQITVITGVFYWIAQLLGSTAACFLLK 120
Query: 121 YVTAK-SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
YVT +VPTH VAAGL I G+V EIIITF LVYTVYATAADPKKGS+GTIAP+AIG +
Sbjct: 121 YVTGGLAVPTHSVAAGLGSIEGVVMEIIITFALVYTVYATAADPKKGSLGTIAPLAIGLI 180
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS- 238
VGANILAAGPFSGGSMNPARSFGPAV +G+F+ +W+YWVGPLIGGGLAGLIYG+VF+GS
Sbjct: 181 VGANILAAGPFSGGSMNPARSFGPAVAAGDFSGHWVYWVGPLIGGGLAGLIYGNVFMGSS 240
Query: 239 -YTPAPASEF 247
+ P +++F
Sbjct: 241 EHVPLASADF 250
>TIP1_MEDTR (Q9FY14) Probable aquaporin TIP-type (MtAQP1)
Length = 250
Score = 307 bits (786), Expect = 2e-83
Identities = 153/235 (65%), Positives = 187/235 (79%)
Query: 4 IGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAH 63
I GT ++ +LKA L+EFI+T IFVFAG GS IAYN LT+D A PAGL++ ++AH
Sbjct: 6 IAVGTPQEATHPDTLKAGLAEFISTFIFVFAGSGSGIAYNKLTNDGAATPAGLISASIAH 65
Query: 64 AFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVT 123
AFALFV V++ ANISGGH+NPAVTFG +GGNIT+L G+ Y IAQLLGSIVAS LL +VT
Sbjct: 66 AFALFVAVSVGANISGGHVNPAVTFGAFVGGNITLLRGIVYIIAQLLGSIVASALLVFVT 125
Query: 124 AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGAN 183
A SVP G++ G+ LV EI++TFGLVYTVYATA DPKKG+IG IAPIAIGF+VGAN
Sbjct: 126 ASSVPAFGLSEGVGVGPALVLEIVMTFGLVYTVYATAVDPKKGNIGIIAPIAIGFIVGAN 185
Query: 184 ILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS 238
IL G F+G SMNPA SFGPAVVS +++++W+YW GPLIGGG+AGL+Y +FI S
Sbjct: 186 ILVGGAFTGASMNPAVSFGPAVVSWSWSNHWVYWAGPLIGGGIAGLVYEVLFINS 240
>TI12_ARATH (Q41963) Aquaporin TIP1.2 (Tonoplast intrinsic protein
1.2) (Gamma-tonoplast intrinsic protein 2) (Gamma-TIP2)
(Salt-stress induced tonoplast intrinsic protein)
Length = 253
Score = 305 bits (780), Expect = 9e-83
Identities = 147/231 (63%), Positives = 185/231 (79%), Gaps = 1/231 (0%)
Query: 7 GTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFA 66
G ++ + +L+A L+EFI+TLIFVFAG GS IA+N +T + A P+GLVA A+AHAF
Sbjct: 10 GVQEEVYHPNALRAALAEFISTLIFVFAGSGSGIAFNKITDNGATTPSGLVAAALAHAFG 69
Query: 67 LFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTA-K 125
LFV V++ ANISGGH+NPAVTFG+ +GGNIT+L G+ YWIAQLLGS+ A LL++ T +
Sbjct: 70 LFVAVSVGANISGGHVNPAVTFGVLLGGNITLLRGILYWIAQLLGSVAACFLLSFATGGE 129
Query: 126 SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGANIL 185
+P G++AG+ + LVFEI++TFGLVYTVYATA DPK GS+GTIAPIAIGF+VGANIL
Sbjct: 130 PIPAFGLSAGVGSLNALVFEIVMTFGLVYTVYATAVDPKNGSLGTIAPIAIGFIVGANIL 189
Query: 186 AAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFI 236
A G FSG SMNPA +FGPAVVS + ++W+YW GPLIGGGLAG+IY VFI
Sbjct: 190 AGGAFSGASMNPAVAFGPAVVSWTWTNHWVYWAGPLIGGGLAGIIYDFVFI 240
>TI11_ARATH (P25818) Aquaporin TIP1.1 (Tonoplast intrinsic protein
1.1) (Gamma-tonoplast intrinsic protein) (Gamma-TIP)
(Aquaporin-TIP) (Tonoplast intrinsic protein,
root-specific RB7)
Length = 251
Score = 302 bits (774), Expect = 4e-82
Identities = 150/236 (63%), Positives = 184/236 (77%), Gaps = 1/236 (0%)
Query: 4 IGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAH 63
I G D++ +LKA L+EFI+TLIFV AG GS +A+N LT + A P+GLVA AVAH
Sbjct: 6 IAIGRPDEATRPDALKAALAEFISTLIFVVAGSGSGMAFNKLTENGATTPSGLVAAAVAH 65
Query: 64 AFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVT 123
AF LFV V++ ANISGGH+NPAVTFG IGGNIT+L G+ YWIAQLLGS+VA L+L + T
Sbjct: 66 AFGLFVAVSVGANISGGHVNPAVTFGAFIGGNITLLRGILYWIAQLLGSVVACLILKFAT 125
Query: 124 AK-SVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGA 182
+VP G++AG+ + VFEI++TFGLVYTVYATA DPK GS+GTIAPIAIGF+VGA
Sbjct: 126 GGLAVPAFGLSAGVGVLNAFVFEIVMTFGLVYTVYATAIDPKNGSLGTIAPIAIGFIVGA 185
Query: 183 NILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS 238
NILA G FSG SMNPA +FGPAVVS + ++W+YW GPL+GGG+AGLIY FI +
Sbjct: 186 NILAGGAFSGASMNPAVAFGPAVVSWTWTNHWVYWAGPLVGGGIAGLIYEVFFINT 241
Score = 35.4 bits (80), Expect = 0.13
Identities = 26/65 (40%), Positives = 33/65 (50%), Gaps = 9/65 (13%)
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFA--DNWIYWVGPLIGGGLAGLIYGDVFIG 237
VGANI SGG +NPA +FG A + GN +YW+ L+G +A LI G
Sbjct: 75 VGANI------SGGHVNPAVTFG-AFIGGNITLLRGILYWIAQLLGSVVACLILKFATGG 127
Query: 238 SYTPA 242
PA
Sbjct: 128 LAVPA 132
>TIP1_MEDSA (P42067) Probable aquaporin TIP-type (Membrane channel
protein 1) (MsMCP1)
Length = 249
Score = 299 bits (765), Expect = 5e-81
Identities = 151/235 (64%), Positives = 185/235 (78%), Gaps = 1/235 (0%)
Query: 4 IGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAH 63
I GT ++ +LKA L+EFI+T IFVFAG GS IAYN LT+D A PAGL++ ++AH
Sbjct: 6 IAVGTPQEATHPDTLKAGLAEFISTFIFVFAGSGSGIAYNKLTNDGAATPAGLISASIAH 65
Query: 64 AFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVT 123
AFALFV V++ ANISGGH+NPAV FG +GGNIT+L G+ Y IAQLLGSIV S LL +VT
Sbjct: 66 AFALFVAVSVGANISGGHVNPAV-FGAFVGGNITLLRGIVYIIAQLLGSIVCSALLVFVT 124
Query: 124 AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGAN 183
A SVP G++ G+ LV EI++TFGLVYTVYATA DPKKG+IG IAPIAIGF+VGAN
Sbjct: 125 ASSVPAFGLSEGVGVGPALVLEIVMTFGLVYTVYATAVDPKKGNIGIIAPIAIGFIVGAN 184
Query: 184 ILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS 238
IL G F+G SMNPA SFGPAVVS +++++W+YW GPLIGGG+AGL+Y +FI S
Sbjct: 185 ILVGGAFTGASMNPAVSFGPAVVSWSWSNHWVYWAGPLIGGGIAGLVYEVLFINS 239
>TIP1_ORYSA (P50156) Probable aquaporin TIP-type 1 (Tonoplast
intrinsic protein gamma) (Gamma TIP)
Length = 250
Score = 290 bits (743), Expect = 2e-78
Identities = 147/245 (60%), Positives = 180/245 (73%), Gaps = 1/245 (0%)
Query: 4 IGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAH 63
I G+ + + +LKA L+EFI+TLIFVFAG GS +A++ LT A PAGL+A AVAH
Sbjct: 6 IAVGSHQEVYHPGALKAALAEFISTLIFVFAGQGSGMAFSKLTGGGATTPAGLIAAAVAH 65
Query: 64 AFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVT 123
AFALFV V++ ANISGGH+NPAVTFG +GGNIT+ GL YWIAQLLGS VA LL + T
Sbjct: 66 AFALFVAVSVGANISGGHVNPAVTFGAFVGGNITLFRGLLYWIAQLLGSTVACFLLRFST 125
Query: 124 AKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGAN 183
G++ LV EI++TFGLVYTVYATA DPKKGS+GTIAPIAIGF+VGAN
Sbjct: 126 GGLATGTFGLTGVSVWEALVLEIVMTFGLVYTVYATAVDPKKGSLGTIAPIAIGFIVGAN 185
Query: 184 ILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIG-SYTPA 242
IL G F G SMNPA SFGPA+VS ++ W+YWVGPLIGGGLAG+IY +FI ++
Sbjct: 186 ILVGGAFDGASMNPAVSFGPALVSWSWESQWVYWVGPLIGGGLAGVIYEVLFISHTHEQL 245
Query: 243 PASEF 247
P +++
Sbjct: 246 PTTDY 250
>TI13_ARATH (O82598) Putative aquaporin TIP1.3 (Tonoplast intrinsic
protein 1.3) (Gamma-tonoplast intrinsic protein 3)
(Gamma-TIP3)
Length = 252
Score = 278 bits (712), Expect = 7e-75
Identities = 137/248 (55%), Positives = 178/248 (71%), Gaps = 3/248 (1%)
Query: 3 KIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVA 62
+I GT ++ +++A +EF + +IFVFAG GS +AY LT D PAGLVA +++
Sbjct: 5 RIAIGTPGEASRPDAIRAAFAEFFSMVIFVFAGQGSGMAYGKLTGDGPATPAGLVAASLS 64
Query: 63 HAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYV 122
HAFALFV V++ AN+SGGH+NPAVTFG IGGNIT+L + YWIAQLLG++VA LLL
Sbjct: 65 HAFALFVAVSVGANVSGGHVNPAVTFGAFIGGNITLLRAILYWIAQLLGAVVACLLLKVS 124
Query: 123 TA-KSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVG 181
T ++ G+ P +VFEI++TFGLVYTVYATA DPKKG IG IAP+AIG +VG
Sbjct: 125 TGGMETAAFSLSYGVTPWNAVVFEIVMTFGLVYTVYATAVDPKKGDIGIIAPLAIGLIVG 184
Query: 182 ANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS--Y 239
ANIL G F G SMNPA SFGPAVVS + ++W+YWVGP IG +A ++Y +FIGS +
Sbjct: 185 ANILVGGAFDGASMNPAVSFGPAVVSWIWTNHWVYWVGPFIGAAIAAIVYDTIFIGSNGH 244
Query: 240 TPAPASEF 247
P P+++F
Sbjct: 245 EPLPSNDF 252
>TI32_ARATH (O22588) Probable aquaporin TIP3.2 (Tonoplast intrinsic
protein 3.2) (Beta-tonoplast intrinsic protein)
(Beta-TIP)
Length = 267
Score = 260 bits (665), Expect = 2e-69
Identities = 132/245 (53%), Positives = 172/245 (69%), Gaps = 6/245 (2%)
Query: 5 GFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALD-----PAGLVAV 59
GFG D++ S++A L+EF++T +FVFAG GS +A + L D A P GLV V
Sbjct: 10 GFGRADEATHPDSIRATLAEFLSTFVFVFAGEGSILALDKLYWDTAAHTGTNTPGGLVLV 69
Query: 60 AVAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLL 119
A+AHA ALF V+ A N+SGGH+NPAVTF IGG I+++ ++YW+AQL+G+I+A LLL
Sbjct: 70 ALAHALALFAAVSAAINVSGGHVNPAVTFAALIGGRISVIRAIYYWVAQLIGAILACLLL 129
Query: 120 NYVTAKSVPT-HGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGF 178
T P VA+G++ + GL+ EII+TF LVY VY+TA DPK+GSIG IAP+AIG
Sbjct: 130 RLATNGLRPVGFHVASGVSELHGLLMEIILTFALVYVVYSTAIDPKRGSIGIIAPLAIGL 189
Query: 179 VVGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS 238
+VGANIL GPF G SMNPAR+FGPA+V ++++WIYWVGP IGG LA LIY + I S
Sbjct: 190 IVGANILVGGPFDGASMNPARAFGPALVGWRWSNHWIYWVGPFIGGALAALIYEYMIIPS 249
Query: 239 YTPAP 243
P
Sbjct: 250 VNEPP 254
>TI31_ARATH (P26587) Aquaporin TIP3.1 (Tonoplast intrinsic protein
3.1) (Alpha-tonoplast intrinsic protein) (Alpha-TIP)
Length = 268
Score = 258 bits (658), Expect = 1e-68
Identities = 130/243 (53%), Positives = 170/243 (69%), Gaps = 6/243 (2%)
Query: 5 GFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALD-----PAGLVAV 59
GFG D++ S++A L+EF++T +FVFA GS ++ + L + A P GL+ V
Sbjct: 10 GFGRADEATHPDSIRATLAEFLSTFVFVFAAEGSILSLDKLYWEHAAHAGTNTPGGLILV 69
Query: 60 AVAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLL 119
A+AHAFALF V+ A N+SGGH+NPAVTFG +GG +T + ++YWIAQLLG+I+A LLL
Sbjct: 70 ALAHAFALFAAVSAAINVSGGHVNPAVTFGALVGGRVTAIRAIYYWIAQLLGAILACLLL 129
Query: 120 NYVTAKSVPT-HGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGF 178
T P +A+G+ + GLV EII+TFGLVY VY+T DPK+GS+G IAP+AIG
Sbjct: 130 RLTTNGMRPVGFRLASGVGAVNGLVLEIILTFGLVYVVYSTLIDPKRGSLGIIAPLAIGL 189
Query: 179 VVGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGS 238
+VGANIL GPFSG SMNPAR+FGPA+V + D+WIYWVGP IG LA LIY + I +
Sbjct: 190 IVGANILVGGPFSGASMNPARAFGPALVGWRWHDHWIYWVGPFIGSALAALIYEYMVIPT 249
Query: 239 YTP 241
P
Sbjct: 250 EPP 252
Score = 39.3 bits (90), Expect = 0.009
Identities = 29/95 (30%), Positives = 43/95 (44%), Gaps = 8/95 (8%)
Query: 51 LDPA-GLVAVAVAHAFALFVG--VAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIA 107
+DP G + + A L VG + + SG +NPA FG A+ G YW+
Sbjct: 172 IDPKRGSLGIIAPLAIGLIVGANILVGGPFSGASMNPARAFGPALVG-WRWHDHWIYWVG 230
Query: 108 QLLGSIVASLLLNYVTAKSVP----THGVAAGLNP 138
+GS +A+L+ Y+ + P HGV L P
Sbjct: 231 PFIGSALAALIYEYMVIPTEPPTHHAHGVHQPLAP 265
>TI41_ARATH (O82316) Probable aquaporin TIP4.1 (Tonoplast intrinsic
protein 4.1) (Epsilon-tonoplast intrinsic protein)
(Epsilon-TIP)
Length = 249
Score = 256 bits (655), Expect = 3e-68
Identities = 132/237 (55%), Positives = 168/237 (70%), Gaps = 5/237 (2%)
Query: 1 MVKIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVA 60
M KI G ++ +KA + EFI T +FVFAGVGSA+A + L + + GL AVA
Sbjct: 1 MKKIELGHHSEAAKPDCIKALIVEFITTFLFVFAGVGSAMATDSLVGNTLV---GLFAVA 57
Query: 61 VAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLN 120
VAHAF + V ++ A +ISGGHLNPAVT GL +GG+I++ YWI QLL S A LL+
Sbjct: 58 VAHAFVVAVMIS-AGHISGGHLNPAVTLGLLLGGHISVFRAFLYWIDQLLASSAACFLLS 116
Query: 121 YVTA-KSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFV 179
Y+T P H +A+G++ G+++EII+TF L++TVYAT DPKKGS+ P+ GFV
Sbjct: 117 YLTGGMGTPVHTLASGVSYTQGIIWEIILTFSLLFTVYATIVDPKKGSLDGFGPLLTGFV 176
Query: 180 VGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFI 236
VGANILA G FSG SMNPARSFGPA+VSGN+ D+W+YWVGPLIGGGLAG IY +V I
Sbjct: 177 VGANILAGGAFSGASMNPARSFGPALVSGNWTDHWVYWVGPLIGGGLAGFIYENVLI 233
>TIPA_PHAVU (P23958) Probable aquaporin TIP-type alpha (Tonoplast
intrinsic protein alpha) (Alpha TIP)
Length = 256
Score = 238 bits (608), Expect = 8e-63
Identities = 115/230 (50%), Positives = 162/230 (70%), Gaps = 1/230 (0%)
Query: 3 KIGFGTFDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVA 62
+ FG D++ S++A L+EF +T IFVFAG GS +A + D+A L+A+A+A
Sbjct: 5 RYSFGRTDEATHPDSMRASLAEFASTFIFVFAGEGSGLALVKIYQDSAFSAGELLALALA 64
Query: 63 HAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYV 122
HAFALF V+ + ++SGGH+NPAV+FG IGG I+++ ++YWIAQLLGSIVA+L+L V
Sbjct: 65 HAFALFAAVSASMHVSGGHVNPAVSFGALIGGRISVIRAVYYWIAQLLGSIVAALVLRLV 124
Query: 123 TAKSVPT-HGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVG 181
T P+ V+ G+ + E+++TFGL+YTVY TA DPK+G++ IAP+AIG +VG
Sbjct: 125 TNNMRPSGFHVSPGVGVGHMFILEVVMTFGLMYTVYGTAIDPKRGAVSYIAPLAIGLIVG 184
Query: 182 ANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIY 231
ANIL GPF G MNPA +FGP++V + +WI+WVGPL+G LA L+Y
Sbjct: 185 ANILVGGPFDGACMNPALAFGPSLVGWQWHQHWIFWVGPLLGAALAALVY 234
Score = 35.0 bits (79), Expect = 0.17
Identities = 23/75 (30%), Positives = 38/75 (50%), Gaps = 7/75 (9%)
Query: 60 AVAHAFALFVGVAIAANI------SGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSI 113
AV++ L +G+ + ANI G +NPA+ FG ++ G +F W+ LLG+
Sbjct: 170 AVSYIAPLAIGLIVGANILVGGPFDGACMNPALAFGPSLVGWQWHQHWIF-WVGPLLGAA 228
Query: 114 VASLLLNYVTAKSVP 128
+A+L+ Y P
Sbjct: 229 LAALVYEYAVIPIEP 243
>TI51_ARATH (Q9STX9) Putative aquaporin TIP5.1 (Tonoplast intrinsic
protein 5.1)
Length = 256
Score = 209 bits (533), Expect = 4e-54
Identities = 108/238 (45%), Positives = 156/238 (65%), Gaps = 3/238 (1%)
Query: 1 MVKIGFGT-FDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAV 59
M+ F + F S +L+ Y+SEFI+T FV A VGS ++ L + P G++
Sbjct: 4 MIPTSFSSKFQGVLSMNALRCYVSEFISTFFFVLAAVGSVMSSRKLMAGDVSGPFGVLIP 63
Query: 60 AVAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLL 119
A+A+A AL V I+ N+SGGH+NPAVTF +A+ G I++ T +FYW +Q++ S++A L+L
Sbjct: 64 AIANALALSSSVYISWNVSGGHVNPAVTFAMAVAGRISVPTAMFYWTSQMIASVMACLVL 123
Query: 120 NY-VTAKSVPTHGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGF 178
V + VP + +A + V E ++ F LVYTV+ TA+DP++G + PI IGF
Sbjct: 124 KVTVMEQHVPIYKIAGEMTGFGASVLEGVLAFVLVYTVF-TASDPRRGLPLAVGPIFIGF 182
Query: 179 VVGANILAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFI 236
V GAN+LAAGPFSGGSMNPA +FG A+V G+F + +YWVGPL+GG A L+Y +V +
Sbjct: 183 VAGANVLAAGPFSGGSMNPACAFGSAMVYGSFKNQAVYWVGPLLGGATAALVYDNVVV 240
Score = 38.9 bits (89), Expect = 0.012
Identities = 29/89 (32%), Positives = 39/89 (43%), Gaps = 4/89 (4%)
Query: 50 ALDPAGLVAVAVAHAFALFVG---VAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWI 106
A DP + +AV F FV V A SGG +NPA FG A+ + YW+
Sbjct: 164 ASDPRRGLPLAVGPIFIGFVAGANVLAAGPFSGGSMNPACAFGSAMVYG-SFKNQAVYWV 222
Query: 107 AQLLGSIVASLLLNYVTAKSVPTHGVAAG 135
LLG A+L+ + V G + G
Sbjct: 223 GPLLGGATAALVYDNVVVPVEDDRGSSTG 251
>AQP4_MOUSE (P55088) Aquaporin 4 (WCH4) (Mercurial-insensitive water
channel) (MIWC)
Length = 323
Score = 164 bits (415), Expect = 2e-40
Identities = 98/231 (42%), Positives = 138/231 (59%), Gaps = 8/231 (3%)
Query: 9 FDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFALF 68
F ++ A KA +EF+ATLIFV GVGS I + + +D +V +++ ++
Sbjct: 26 FKGVWTQAFWKAVSAEFLATLIFVLLGVGSTINWGGSENPLPVD---MVLISLCFGLSIA 82
Query: 69 VGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTAKSVP 128
V +ISGGH+NPAVT + I+I +FY IAQ LG+I+ + +L VT SV
Sbjct: 83 TMVQCFGHISGGHINPAVTVAMVCTRKISIAKSVFYIIAQCLGAIIGAGILYLVTPPSVV 142
Query: 129 ----THGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGANI 184
V L GL+ E+IITF LV+T++A+ + G+IA +AIGF V
Sbjct: 143 GGLGVTTVHGNLTAGHGLLVELIITFQLVFTIFASCDSKRTDVTGSIA-LAIGFSVAIGH 201
Query: 185 LAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVF 235
L A ++G SMNPARSFGPAV+ GN+A++WIYWVGP++G LAG +Y VF
Sbjct: 202 LFAINYTGASMNPARSFGPAVIMGNWANHWIYWVGPIMGAVLAGALYEYVF 252
Score = 42.7 bits (99), Expect = 8e-04
Identities = 24/71 (33%), Positives = 39/71 (54%), Gaps = 3/71 (4%)
Query: 58 AVAVAHAFALFVGVAIAANISGGHLNPAVTFGLA-IGGNITILTGLFYWIAQLLGSIVAS 116
++A+A F++ +G A N +G +NPA +FG A I GN YW+ ++G+++A
Sbjct: 188 SIALAIGFSVAIGHLFAINYTGASMNPARSFGPAVIMGNWA--NHWIYWVGPIMGAVLAG 245
Query: 117 LLLNYVTAKSV 127
L YV V
Sbjct: 246 ALYEYVFCPDV 256
>AQP4_RAT (P47863) Aquaporin 4 (WCH4) (Mercurial-insensitive water
channel) (MIWC)
Length = 323
Score = 157 bits (396), Expect = 3e-38
Identities = 94/231 (40%), Positives = 134/231 (57%), Gaps = 8/231 (3%)
Query: 9 FDDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFALF 68
F ++ A KA +EF+A LIFV VGS I + + +D +V +++ ++
Sbjct: 26 FKGVWTQAFWKAVTAEFLAMLIFVLLSVGSTINWGGSENPLPVD---MVLISLCFGLSIA 82
Query: 69 VGVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTAKSVP 128
V +ISGGH+NPAVT + I+I +FY AQ LG+I+ + +L VT SV
Sbjct: 83 TMVQCFGHISGGHINPAVTVAMVCTRKISIAKSVFYITAQCLGAIIGAGILYLVTPPSVV 142
Query: 129 ----THGVAAGLNPIAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGANI 184
V L GL+ E+IITF LV+T++A+ + G++A +AIGF V
Sbjct: 143 GGLGVTTVHGNLTAGHGLLVELIITFQLVFTIFASCDSKRTDVTGSVA-LAIGFSVAIGH 201
Query: 185 LAAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVF 235
L A ++G SMNPARSFGPAV+ GN+ ++WIYWVGP+IG LAG +Y VF
Sbjct: 202 LFAINYTGASMNPARSFGPAVIMGNWENHWIYWVGPIIGAVLAGALYEYVF 252
Score = 42.7 bits (99), Expect = 8e-04
Identities = 25/71 (35%), Positives = 39/71 (54%), Gaps = 3/71 (4%)
Query: 58 AVAVAHAFALFVGVAIAANISGGHLNPAVTFGLA-IGGNITILTGLFYWIAQLLGSIVAS 116
+VA+A F++ +G A N +G +NPA +FG A I GN YW+ ++G+++A
Sbjct: 188 SVALAIGFSVAIGHLFAINYTGASMNPARSFGPAVIMGNWE--NHWIYWVGPIIGAVLAG 245
Query: 117 LLLNYVTAKSV 127
L YV V
Sbjct: 246 ALYEYVFCPDV 256
>MIP_HUMAN (P30301) Lens fiber major intrinsic protein (MIP26)
(MP26) (Aquaporin 0)
Length = 263
Score = 155 bits (391), Expect = 1e-37
Identities = 84/222 (37%), Positives = 130/222 (57%), Gaps = 12/222 (5%)
Query: 14 SAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFALFVGVAI 73
SA+ +A +EF ATL +VF G+GS++ + A P ++ VA+A AL V
Sbjct: 6 SASFWRAIFAEFFATLFYVFFGLGSSLRW-------APGPLHVLQVAMAFGLALATLVQS 58
Query: 74 AANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTAKSVPTHGVA 133
+ISG H+NPAVTF +G +++L Y AQLLG++ + +L VT +V +
Sbjct: 59 VGHISGAHVNPAVTFAFLVGSQMSLLRAFCYMAAQLLGAVAGAAVLYSVTPPAVRGNLAL 118
Query: 134 AGLNPIAGL----VFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGANILAAGP 189
L+P + EI +T V ++AT + + G +G++A +A+GF + L
Sbjct: 119 NTLHPAVSVGQATTVEIFLTLQFVLCIFATYDERRNGQLGSVA-LAVGFSLALGHLFGMY 177
Query: 190 FSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIY 231
++G MNPARSF PA+++GNF ++W+YWVGP+IGGGL L+Y
Sbjct: 178 YTGAGMNPARSFAPAILTGNFTNHWVYWVGPIIGGGLGSLLY 219
Score = 35.8 bits (81), Expect = 0.099
Identities = 24/72 (33%), Positives = 38/72 (52%), Gaps = 11/72 (15%)
Query: 56 LVAVAVAHAFALFVGVAIAANISGGHLNPAVTFGLAIGGNITILTGLF-----YWIAQLL 110
L +VA+A F+L +G +G +NPA +F AI LTG F YW+ ++
Sbjct: 157 LGSVALAVGFSLALGHLFGMYYTGAGMNPARSFAPAI------LTGNFTNHWVYWVGPII 210
Query: 111 GSIVASLLLNYV 122
G + SLL +++
Sbjct: 211 GGGLGSLLYDFL 222
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.141 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,309,071
Number of Sequences: 164201
Number of extensions: 1256627
Number of successful extensions: 4391
Number of sequences better than 10.0: 213
Number of HSP's better than 10.0 without gapping: 155
Number of HSP's successfully gapped in prelim test: 58
Number of HSP's that attempted gapping in prelim test: 3679
Number of HSP's gapped (non-prelim): 260
length of query: 247
length of database: 59,974,054
effective HSP length: 107
effective length of query: 140
effective length of database: 42,404,547
effective search space: 5936636580
effective search space used: 5936636580
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC137077.10


