

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.11 - phase: 2 /pseudo
(192 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
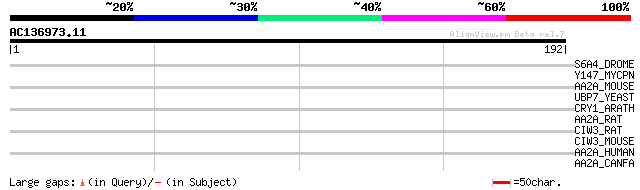
Score E
Sequences producing significant alignments: (bits) Value
S6A4_DROME (P51905) Sodium-dependent serotonin transporter (5HT ... 32 1.2
Y147_MYCPN (P75585) Hypothetical protein MG147 homolog (VXpSPT7_... 31 2.0
AA2A_MOUSE (Q60613) Adenosine A2a receptor 30 2.6
UBP7_YEAST (P40453) Ubiquitin carboxyl-terminal hydrolase 7 (EC ... 30 3.4
CRY1_ARATH (Q43125) Cryptochrome 1 apoprotein (Blue light photor... 30 4.5
AA2A_RAT (P30543) Adenosine A2a receptor 30 4.5
CIW3_RAT (O54912) Potassium channel subfamily K member 3 (Acid-s... 29 5.9
CIW3_MOUSE (O35111) Potassium channel subfamily K member 3 (Acid... 29 5.9
AA2A_HUMAN (P29274) Adenosine A2a receptor 29 5.9
AA2A_CANFA (P11617) Adenosine A2a receptor 29 5.9
>S6A4_DROME (P51905) Sodium-dependent serotonin transporter (5HT
transporter) (5HTT) (Cocaine-sensitive serotonin
transporter) (dSERT1)
Length = 622
Score = 31.6 bits (70), Expect = 1.2
Identities = 14/48 (29%), Positives = 23/48 (47%), Gaps = 9/48 (18%)
Query: 64 WTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTF 111
W CW + P+ LLT +FS + +L + YYY +W++
Sbjct: 528 WRICWTYISPVFLLTIFIFSIMGYKEMLGEE---------YYYPDWSY 566
>Y147_MYCPN (P75585) Hypothetical protein MG147 homolog
(VXpSPT7_orf377)
Length = 377
Score = 30.8 bits (68), Expect = 2.0
Identities = 23/65 (35%), Positives = 30/65 (45%), Gaps = 3/65 (4%)
Query: 74 LVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEW--TFTLVMIYFALGTTVSAYGCWK 131
L LLT L F S + + D H + F TE TF + + FAL + GCWK
Sbjct: 31 LTLLTYGLVLFSSSGRIDHGD-HFHLRERFVLTTEELVTFVVASVVFALTVALFGLGCWK 89
Query: 132 VLNKP 136
+L P
Sbjct: 90 LLQGP 94
>AA2A_MOUSE (Q60613) Adenosine A2a receptor
Length = 410
Score = 30.4 bits (67), Expect = 2.6
Identities = 26/98 (26%), Positives = 45/98 (45%), Gaps = 4/98 (4%)
Query: 2 YDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVSTS 61
Y V AI ++ILG + V W +S Q+ +N FV SL A+ + ++
Sbjct: 6 YIMVELAIAVLAILGNVLVCWAVWINSNLQNVTNFFVVSLAAADIAVGVLAIPFAITIST 65
Query: 62 QLWTSC----WRGVHPLVLLTTRLFSFVSLAMLLYLDI 95
+C + LVL + +FS +++A+ Y+ I
Sbjct: 66 GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAI 103
>UBP7_YEAST (P40453) Ubiquitin carboxyl-terminal hydrolase 7 (EC
3.1.2.15) (Ubiquitin thiolesterase 7) (Ubiquitin-specific
processing protease 7) (Deubiquitinating enzyme 7)
Length = 1071
Score = 30.0 bits (66), Expect = 3.4
Identities = 25/100 (25%), Positives = 43/100 (43%), Gaps = 11/100 (11%)
Query: 76 LLTTRLFSFVSLAMLLYLDIHEYDASIFYY------YTEWTFTLVMIYFALGTTVSAYGC 129
L T + +FV+L +L + + S FYY T T+ L++ Y
Sbjct: 946 LTTVKTINFVTLPKILVIHL-----SRFYYDLTKKNNTVVTYPLILNIILKNNDTMKYKL 1000
Query: 130 WKVLNKPPPLQNGEMTEFLRRDLETKGSIFTFQSRYAEEE 169
+ V+N L +G T + +DLE +I + Y ++E
Sbjct: 1001 FGVVNHTGTLISGHYTSLVNKDLEHNVNIGRSKWYYFDDE 1040
>CRY1_ARATH (Q43125) Cryptochrome 1 apoprotein (Blue light
photoreceptor)
Length = 681
Score = 29.6 bits (65), Expect = 4.5
Identities = 14/47 (29%), Positives = 23/47 (48%), Gaps = 8/47 (17%)
Query: 20 VLWTNEGSSTSQSDSNIFVESL--------LVANSPSSDNRVAIGHV 58
V W NEG+ + N+F++S+ + N P S R +GH+
Sbjct: 273 VAWANEGNEAGEESVNLFLKSIGLREYSRYISFNHPYSHERPLLGHL 319
>AA2A_RAT (P30543) Adenosine A2a receptor
Length = 410
Score = 29.6 bits (65), Expect = 4.5
Identities = 26/98 (26%), Positives = 45/98 (45%), Gaps = 4/98 (4%)
Query: 2 YDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVSTS 61
Y V AI ++ILG + V W +S Q+ +N FV SL A+ + ++
Sbjct: 6 YITVELAIAVLAILGNVLVCWAVWINSNLQNVTNFFVVSLAAADIAVGVLAIPFAITIST 65
Query: 62 QLWTSC----WRGVHPLVLLTTRLFSFVSLAMLLYLDI 95
+C + LVL + +FS +++A+ Y+ I
Sbjct: 66 GFCAACHGCLFFACFVLVLTQSSIFSLLAIAIDRYIAI 103
>CIW3_RAT (O54912) Potassium channel subfamily K member 3
(Acid-sensitive potassium channel protein TASK-1)
(TWIK-related acid-sensitive K+ channel 1) (Two pore
potassium channel KT3.1)
Length = 411
Score = 29.3 bits (64), Expect = 5.9
Identities = 19/60 (31%), Positives = 24/60 (39%), Gaps = 5/60 (8%)
Query: 81 LFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFALGTTVSAYGCWKVLNKPPPLQ 140
L FVS L + A+ F YY WTF Y + T +G + L K LQ
Sbjct: 161 LIGFVSCISTLCIG-----AAAFSYYERWTFFQAYYYCFITLTTIGFGDYVALQKDQALQ 215
>CIW3_MOUSE (O35111) Potassium channel subfamily K member 3
(Acid-sensitive potassium channel protein TASK-1)
(TWIK-related acid-sensitive K+ channel 1) (Cardiac
two-pore background K+ channel) (cTBAK-1) (Two pore
potassium channel KT3.1)
Length = 409
Score = 29.3 bits (64), Expect = 5.9
Identities = 19/60 (31%), Positives = 24/60 (39%), Gaps = 5/60 (8%)
Query: 81 LFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFALGTTVSAYGCWKVLNKPPPLQ 140
L FVS L + A+ F YY WTF Y + T +G + L K LQ
Sbjct: 161 LIGFVSCISTLCIG-----AAAFSYYERWTFFQAYYYCFITLTTIGFGDYVALQKDQALQ 215
>AA2A_HUMAN (P29274) Adenosine A2a receptor
Length = 412
Score = 29.3 bits (64), Expect = 5.9
Identities = 26/98 (26%), Positives = 45/98 (45%), Gaps = 4/98 (4%)
Query: 2 YDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVSTS 61
Y V AI ++ILG + V W +S Q+ +N FV SL A+ + ++
Sbjct: 9 YITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST 68
Query: 62 QLWTSC----WRGVHPLVLLTTRLFSFVSLAMLLYLDI 95
+C + LVL + +FS +++A+ Y+ I
Sbjct: 69 GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAI 106
>AA2A_CANFA (P11617) Adenosine A2a receptor
Length = 412
Score = 29.3 bits (64), Expect = 5.9
Identities = 26/98 (26%), Positives = 45/98 (45%), Gaps = 4/98 (4%)
Query: 2 YDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVSTS 61
Y V AI ++ILG + V W +S Q+ +N FV SL A+ + ++
Sbjct: 9 YITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST 68
Query: 62 QLWTSC----WRGVHPLVLLTTRLFSFVSLAMLLYLDI 95
+C + LVL + +FS +++A+ Y+ I
Sbjct: 69 GFCAACHNCLFFACFVLVLTQSSIFSLLAIAIDRYIAI 106
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,554,710
Number of Sequences: 164201
Number of extensions: 794398
Number of successful extensions: 2470
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 2467
Number of HSP's gapped (non-prelim): 10
length of query: 192
length of database: 59,974,054
effective HSP length: 104
effective length of query: 88
effective length of database: 42,897,150
effective search space: 3774949200
effective search space used: 3774949200
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC136973.11


