

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
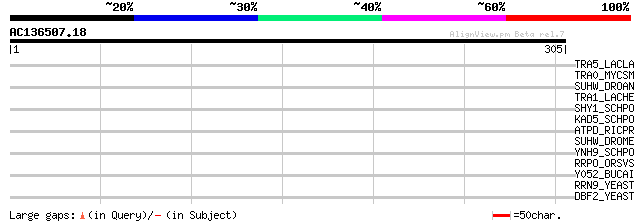
Query= AC136507.18 - phase: 1 /pseudo
(305 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TRA5_LACLA (P35881) Transposase for insertion sequence element I... 37 0.047
TRA0_MYCSM (P35883) Transposase for insertion sequence element I... 37 0.061
SUHW_DROAN (Q08875) Suppressor of hairy wing protein 32 1.5
TRA1_LACHE (P35880) Transposase for insertion sequence element I... 32 2.0
SHY1_SCHPO (Q9Y810) Shy1 protein (SURF1-like protein) 32 2.6
KAD5_SCHPO (Q09831) Probable serine/threonine-protein kinase C4G... 32 2.6
ATPD_RICPR (Q9ZCF2) ATP synthase delta chain (EC 3.6.3.14) 32 2.6
SUHW_DROME (P08970) Suppressor of hairy wing protein 31 4.4
YNH9_SCHPO (Q96WW0) Hypothetical WD-repeat protein C32H8.09 in c... 30 5.7
RRPO_ORSVS (Q84133) RNA-directed RNA polymerase (EC 2.7.7.48) (1... 30 5.7
Y052_BUCAI (P57160) Hypothetical protein BU052 30 7.4
RRN9_YEAST (P53437) RNA polymerase I specific transcription init... 30 9.7
DBF2_YEAST (P22204) Cell cycle protein kinase DBF2 (EC 2.7.1.37) 30 9.7
>TRA5_LACLA (P35881) Transposase for insertion sequence element
IS905
Length = 391
Score = 37.4 bits (85), Expect = 0.047
Identities = 30/147 (20%), Positives = 66/147 (44%), Gaps = 10/147 (6%)
Query: 134 LISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTY----KTNIYR 189
+I ++ H Y+ + + + T E++ H S++ N+ +VL +D TY + + +
Sbjct: 120 IIERMYGHHYSPATISNISKATQENVATFHERSLEA--NY-SVLFLDGTYLPLRRGTVSK 176
Query: 190 MPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSL 249
+ +G+T +G+ +E N W +L KL + + ++VTD L
Sbjct: 177 ECIHIALGITPEGQKAVLGYEIAPNEN--NASWST-LLDKLQNQGIQQVSLVVTDGFKGL 233
Query: 250 MKAVAHVFPESYAMNCYFHVQANVKQR 276
+ ++ +P + C H+ N+ +
Sbjct: 234 EQIISQAYPLAKQQRCLIHISRNLASK 260
>TRA0_MYCSM (P35883) Transposase for insertion sequence element
IS6120
Length = 323
Score = 37.0 bits (84), Expect = 0.061
Identities = 18/48 (37%), Positives = 24/48 (49%)
Query: 226 MLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANV 273
+LR M P + + D + KAV VFP + C+FH QANV
Sbjct: 113 LLRDCKRRGMTAPVLAIGDGALGFWKAVREVFPATKEQRCWFHKQANV 160
>SUHW_DROAN (Q08875) Suppressor of hairy wing protein
Length = 886
Score = 32.3 bits (72), Expect = 1.5
Identities = 26/124 (20%), Positives = 47/124 (36%), Gaps = 10/124 (8%)
Query: 7 VCERSGSHIVPKKKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALE 66
+C++ S +V KK + TG + C + K+ + G H E +
Sbjct: 446 LCDKKLSALVALKKHRRYHTGEKPYSCTVCNQAFAVKEVLNRHMKRHTGERPHKCEECGK 505
Query: 67 GHILAGRLKEDDKKIVRDLT---------KNKMLPRNILIHLKNQRPH-CMTNVKQVYNE 116
I A +L+ K +R K L R++ H + +RP+ T K+ +
Sbjct: 506 SFIQATQLRTHSKTHIRPFACDMCEEKFKTEKQLERHVKEHSRQKRPYFSCTECKRHFRN 565
Query: 117 RQQI 120
Q+
Sbjct: 566 TAQL 569
>TRA1_LACHE (P35880) Transposase for insertion sequence element
IS1201
Length = 369
Score = 32.0 bits (71), Expect = 2.0
Identities = 29/144 (20%), Positives = 62/144 (42%), Gaps = 10/144 (6%)
Query: 134 LISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTY----KTNIYR 189
LI K+ Y+ + + I + H KL + F V +D+TY + R
Sbjct: 120 LIEKMYGSHYSPAQVSNISKQMIPKVEAYHKR--KLSDKFFCVY-LDATYVPLRRETFER 176
Query: 190 MPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSL 249
++ +G+ + + +E E VW ++L+ + S + ++ ++D + +
Sbjct: 177 EAVYIAIGIKPNGHKEVIDYCIAPNENIE--VWT-ELLQSMKSRGLEQVELFLSDGVVGM 233
Query: 250 MKAVAHVFPESYAMNCYFHVQANV 273
A+A +P+++ C HV N+
Sbjct: 234 KTALAKTYPQAHFQRCLVHVMRNI 257
>SHY1_SCHPO (Q9Y810) Shy1 protein (SURF1-like protein)
Length = 290
Score = 31.6 bits (70), Expect = 2.6
Identities = 25/96 (26%), Positives = 44/96 (45%), Gaps = 7/96 (7%)
Query: 3 MFRLVCERSGSHIVPKKKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPME 62
+FR + G+ + PKK G S F + +Q K+ +EW++ I+N + +
Sbjct: 19 VFRYLSTLEGTTVRPKKNKFLVGLLSAVPIVTFALGTWQVKR-REWKMGIINTLTERLQQ 77
Query: 63 PALEGHILAGRLKEDDKKIVRDLTKNKMLPRNILIH 98
PA+ +L + E D K L ++L R + H
Sbjct: 78 PAI---LLPKTVTEQDTK---KLEWTRVLLRGVFCH 107
>KAD5_SCHPO (Q09831) Probable serine/threonine-protein kinase
C4G8.05 (EC 2.7.1.37)
Length = 566
Score = 31.6 bits (70), Expect = 2.6
Identities = 25/96 (26%), Positives = 44/96 (45%), Gaps = 6/96 (6%)
Query: 149 TQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVG 208
T ++ ++DIF H +L + TV TY +N ++ E VG +S + + +G
Sbjct: 144 TSVQREKLKDIFSPHGKEKELAHIKKTVATRARTYSSNSIKICDVE-VGPSSFEKVFLLG 202
Query: 209 FG-----FMTHEKEENFVWVLKMLRKLLSSKMNMPK 239
G ++ EK+ + +K+L K K N K
Sbjct: 203 KGDVGRVYLVREKKSGKFYAMKVLSKQEMIKRNKSK 238
>ATPD_RICPR (Q9ZCF2) ATP synthase delta chain (EC 3.6.3.14)
Length = 183
Score = 31.6 bits (70), Expect = 2.6
Identities = 12/36 (33%), Positives = 24/36 (66%)
Query: 81 IVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQVYNE 116
+V+ NK++ +L+ +KN R H ++N+ +VYN+
Sbjct: 65 LVKTTNFNKIVNNFLLLLIKNSRTHILSNIVEVYNK 100
>SUHW_DROME (P08970) Suppressor of hairy wing protein
Length = 941
Score = 30.8 bits (68), Expect = 4.4
Identities = 22/107 (20%), Positives = 40/107 (36%), Gaps = 9/107 (8%)
Query: 7 VCERSGSHIVPKKKPKHAGTGSRKCGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALE 66
+C++ S +V KK + TG + C + K+ + G H + +
Sbjct: 445 LCDKKFSALVALKKHRRYHTGEKPYSCTVCNQAFAVKEVLNRHMKRHTGERPHKCDECGK 504
Query: 67 GHILAGRLKEDDKKIVR---------DLTKNKMLPRNILIHLKNQRP 104
I A +L+ K +R K L R++ H + +RP
Sbjct: 505 SFIQATQLRTHSKTHIRPFPCEQCDEKFKTEKQLERHVKTHSRTKRP 551
>YNH9_SCHPO (Q96WW0) Hypothetical WD-repeat protein C32H8.09 in
chromosome II
Length = 483
Score = 30.4 bits (67), Expect = 5.7
Identities = 16/78 (20%), Positives = 38/78 (48%), Gaps = 6/78 (7%)
Query: 120 IWKANRGDKKPLQYLISKLEEH------KYTYYSRTQLESNTIEDIFWAHPTSIKLFNNF 173
+WK N G +K Q ++ +++ ++ YY+ Q + + + I W+ + L+++F
Sbjct: 61 LWKPNLGAEKCHQICVASVDKVFVLDIVQHDYYASIQCDQDPLSSISWSPSGELLLWSSF 120
Query: 174 PTVLVMDSTYKTNIYRMP 191
+ + + S Y +P
Sbjct: 121 DSKITVWSLNTQKGYLLP 138
>RRPO_ORSVS (Q84133) RNA-directed RNA polymerase (EC 2.7.7.48) (183
kDa protein) [Contains: Methyltransferase/RNA helicase
(MT/HEL) (126 kDa protein)]
Length = 1612
Score = 30.4 bits (67), Expect = 5.7
Identities = 22/96 (22%), Positives = 40/96 (40%), Gaps = 10/96 (10%)
Query: 210 GFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHV-FPESYAMNCYFH 268
G M H NF+ L R+ + V + + VA+ +P+ + C +H
Sbjct: 898 GLMLHTGCVNFLVALSHCREAM--------VFGDAEQIPFINRVANFPYPKHFRYTCLYH 949
Query: 269 VQAN-VKQRCVLDCKYPLGFKKDGKEVSNRDVVKKI 303
+ + RC D + + K DGK + DV++ +
Sbjct: 950 REVRRLSLRCPADVTHFMNSKYDGKVLCTNDVIRSV 985
>Y052_BUCAI (P57160) Hypothetical protein BU052
Length = 144
Score = 30.0 bits (66), Expect = 7.4
Identities = 13/39 (33%), Positives = 21/39 (53%)
Query: 156 IEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFE 194
+++IF I+++ FP +LV+ TYK N FE
Sbjct: 78 LKNIFLGKIKEIEIYKMFPIILVLSDTYKVNACIKKFFE 116
>RRN9_YEAST (P53437) RNA polymerase I specific transcription
initiation factor RRN9
Length = 365
Score = 29.6 bits (65), Expect = 9.7
Identities = 39/160 (24%), Positives = 70/160 (43%), Gaps = 18/160 (11%)
Query: 82 VRDLTKNKMLPRNILIHLKN--QRPHCMTNVKQVYNERQQIW----KANRGDKKPLQYLI 135
V +L + RNIL+ L + + H + ++ RQ +AN+ D +P+Q
Sbjct: 165 VDELNIPNEISRNILVKLDSLFEGLHDKIAKENEFDVRQDKHSNNIRANQIDDEPMQ--- 221
Query: 136 SKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEV 195
KYTY+ ED+ + S++L+N+ P YK +R+P ++
Sbjct: 222 -ANRRIKYTYHDLVSRGCEMNEDMTDIYMKSLELYNDIP------EKYKKRKFRLPK-QI 273
Query: 196 VGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLR-KLLSSK 234
+ S + + E+F+ V K+L+ K L+SK
Sbjct: 274 LKKYHQPKKTSSYLKELLSKTREDFIPVEKLLKDKRLTSK 313
>DBF2_YEAST (P22204) Cell cycle protein kinase DBF2 (EC 2.7.1.37)
Length = 572
Score = 29.6 bits (65), Expect = 9.7
Identities = 15/42 (35%), Positives = 26/42 (61%), Gaps = 2/42 (4%)
Query: 208 GFG--FMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDM 247
G+G ++ +K+ V LK+L K L K+N K ++T+RD+
Sbjct: 187 GYGQVYLARKKDTKEVCALKILNKKLLFKLNETKHVLTERDI 228
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,786,562
Number of Sequences: 164201
Number of extensions: 1438958
Number of successful extensions: 3199
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 3195
Number of HSP's gapped (non-prelim): 13
length of query: 305
length of database: 59,974,054
effective HSP length: 110
effective length of query: 195
effective length of database: 41,911,944
effective search space: 8172829080
effective search space used: 8172829080
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 65 (29.6 bits)
Medicago: description of AC136507.18


