

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136507.1 - phase: 0
(236 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
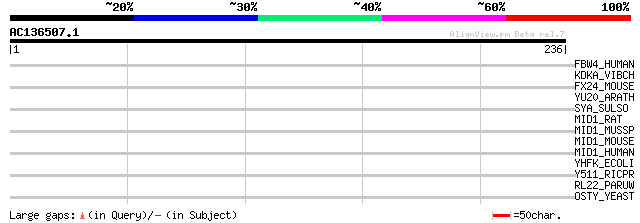
Score E
Sequences producing significant alignments: (bits) Value
FBW4_HUMAN (P57775) F-box/WD-repeat protein 4 (F-box and WD-40 d... 32 1.0
KDKA_VIBCH (Q9KVB9) 3-deoxy-D-manno-octulosonic acid kinase (EC ... 32 1.3
FX24_MOUSE (Q9D417) F-box only protein 24 31 2.2
YU20_ARATH (Q9ZPV5) Hypothetical UPF0120 protein At2g18220 30 3.8
SYA_SULSO (P96041) Alanyl-tRNA synthetase (EC 6.1.1.7) (Alanine-... 30 5.0
MID1_RAT (P82458) Midline 1 protein (Tripartite motif-containing... 30 6.5
MID1_MUSSP (P82457) Midline 1 protein (Tripartite motif-containi... 30 6.5
MID1_MOUSE (O70583) Midline 1 protein (EC 6.3.2.-) (Tripartite m... 30 6.5
MID1_HUMAN (O15344) Midline 1 protein (EC 6.3.2.-) (Tripartite m... 30 6.5
YHFK_ECOLI (P45537) Hypothetical protein yhfK 29 8.5
Y511_RICPR (Q9ZD36) Hypothetical protein RP511 29 8.5
RL22_PARUW (Q6ME58) 50S ribosomal protein L22 29 8.5
OSTY_YEAST (Q03723) Dolichyl-diphosphooligosaccharide--protein g... 29 8.5
>FBW4_HUMAN (P57775) F-box/WD-repeat protein 4 (F-box and WD-40
domain protein 4) (Dactylin)
Length = 412
Score = 32.3 bits (72), Expect = 1.0
Identities = 32/113 (28%), Positives = 50/113 (43%), Gaps = 14/113 (12%)
Query: 89 LPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGR--------VIGS 140
LPE +L I L +L ++A VC LR D LW R + ++ S
Sbjct: 31 LPEELLLLICSYLDMRALGRLAQVCRWLRRFTSCDLLWRRIARASLNSGFTRLGTDLMTS 90
Query: 141 VAYREWKWHVATKKNIGSLRHGKPRSLIKFMSLYWPFSWMRMKVDAAYSNSAN 193
V +E + V+ +G R G L+K+ P WM+++ D+ Y + AN
Sbjct: 91 VPVKE-RVKVSQNWRLGRCREG---ILLKWRCSQMP--WMQLEDDSLYISQAN 137
>KDKA_VIBCH (Q9KVB9) 3-deoxy-D-manno-octulosonic acid kinase (EC
2.7.1.-) (KDO kinase)
Length = 251
Score = 32.0 bits (71), Expect = 1.3
Identities = 18/72 (25%), Positives = 32/72 (44%), Gaps = 7/72 (9%)
Query: 114 HSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGKPRSLIKFM-- 171
H +C + W+++ G+V+GS R W V ++ G+LRH + L +
Sbjct: 36 HEDPSQCCNPEFWQQN-----GKVLGSATGRGTTWFVQLQQTQGALRHYRRGGLFGKLVA 90
Query: 172 SLYWPFSWMRMK 183
YW W + +
Sbjct: 91 DSYWFTGWEKTR 102
>FX24_MOUSE (Q9D417) F-box only protein 24
Length = 589
Score = 31.2 bits (69), Expect = 2.2
Identities = 19/65 (29%), Positives = 30/65 (45%), Gaps = 1/65 (1%)
Query: 84 LSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIG-SVA 142
+SV P ++E I+ LP L + CH E C ++ +W R ++ R+ S
Sbjct: 24 ISVQLFPPELVEHIVSFLPVKDLVALGQTCHYFHEVCDAEGVWRRICRRLSPRIRDQSSG 83
Query: 143 YREWK 147
R WK
Sbjct: 84 ARPWK 88
>YU20_ARATH (Q9ZPV5) Hypothetical UPF0120 protein At2g18220
Length = 779
Score = 30.4 bits (67), Expect = 3.8
Identities = 26/100 (26%), Positives = 41/100 (41%), Gaps = 4/100 (4%)
Query: 20 LLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRMSLVKKTLVSKVENVEEEVDS 79
L++ L W+ V L F L+SFCK T R K L+S++E E V+
Sbjct: 527 LVEHLSQWSCSVAFFELSFIPTIRLRSFCK---STKAERFRKEMKQLISQIEANSEFVNK 583
Query: 80 LQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRER 119
+ + L +L E LE + + +R+R
Sbjct: 584 KRALIKFLP-NDLAAESFLEDEKKAGKTPLLQYAEIIRQR 622
>SYA_SULSO (P96041) Alanyl-tRNA synthetase (EC 6.1.1.7)
(Alanine--tRNA ligase) (AlaRS)
Length = 900
Score = 30.0 bits (66), Expect = 5.0
Identities = 15/52 (28%), Positives = 28/52 (53%)
Query: 54 TLVSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSS 105
T+ +V K L ++NVEEE+D + +LD + ++ + + L PS+
Sbjct: 298 TIKIHRQMVSKELGIDIKNVEEELDRAAKVFQILDHTKTIMLMLADGLVPSN 349
>MID1_RAT (P82458) Midline 1 protein (Tripartite motif-containing
protein 18)
Length = 667
Score = 29.6 bits (65), Expect = 6.5
Identities = 29/104 (27%), Positives = 49/104 (46%), Gaps = 10/104 (9%)
Query: 20 LLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRMSLVKKTLVSKVENVEEEVDS 79
L++P+P + +R + L H+D + +C + + + LV + +V + E D
Sbjct: 161 LIEPIPD--SHIRGLMCLEHEDEKVNMYCVTDDQLICALCKLVGRHRDHQVAALSERYDK 218
Query: 80 LQQNLSVLDLPELV-----LECILEKLPPSSLCQMAGVCHSLRE 118
L+QNL +L L+ LE +L KL CQ V S +E
Sbjct: 219 LKQNLE-SNLTNLIKRNTELETLLAKL--IQTCQHVEVNASRQE 259
>MID1_MUSSP (P82457) Midline 1 protein (Tripartite motif-containing
protein 18)
Length = 667
Score = 29.6 bits (65), Expect = 6.5
Identities = 29/104 (27%), Positives = 49/104 (46%), Gaps = 10/104 (9%)
Query: 20 LLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRMSLVKKTLVSKVENVEEEVDS 79
L++P+P + +R + L H+D + +C + + + LV + +V + E D
Sbjct: 161 LIEPIPD--SHIRGLMCLEHEDEKVNMYCVTDDQLICALCKLVGRHRDHQVAALSERYDK 218
Query: 80 LQQNLSVLDLPELV-----LECILEKLPPSSLCQMAGVCHSLRE 118
L+QNL +L L+ LE +L KL CQ V S +E
Sbjct: 219 LKQNLE-SNLTNLIKRNTELETLLAKL--IQTCQHVEVNASRQE 259
>MID1_MOUSE (O70583) Midline 1 protein (EC 6.3.2.-) (Tripartite
motif-containing protein 18) (Midin)
Length = 680
Score = 29.6 bits (65), Expect = 6.5
Identities = 29/104 (27%), Positives = 49/104 (46%), Gaps = 10/104 (9%)
Query: 20 LLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRMSLVKKTLVSKVENVEEEVDS 79
L++P+P + +R + L H+D + +C + + + LV + +V + E D
Sbjct: 161 LIEPIPD--SHIRGLMCLEHEDEKVNMYCVTDDQLICALCKLVGRHRDHQVAALSERYDK 218
Query: 80 LQQNLSVLDLPELV-----LECILEKLPPSSLCQMAGVCHSLRE 118
L+QNL +L L+ LE +L KL CQ V S +E
Sbjct: 219 LKQNLE-SNLTNLIKRNTELETLLAKL--IQTCQHVEVNASRQE 259
>MID1_HUMAN (O15344) Midline 1 protein (EC 6.3.2.-) (Tripartite
motif-containing protein 18) (Putative transcription
factor XPRF) (Midin)
Length = 667
Score = 29.6 bits (65), Expect = 6.5
Identities = 29/104 (27%), Positives = 49/104 (46%), Gaps = 10/104 (9%)
Query: 20 LLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRMSLVKKTLVSKVENVEEEVDS 79
L++P+P + +R + L H+D + +C + + + LV + +V + E D
Sbjct: 161 LIEPIPD--SHIRGLMCLEHEDEKVNMYCVTDDQLICALCKLVGRHRDHQVAALSERYDK 218
Query: 80 LQQNLSVLDLPELV-----LECILEKLPPSSLCQMAGVCHSLRE 118
L+QNL +L L+ LE +L KL CQ V S +E
Sbjct: 219 LKQNLE-SNLTNLIKRNTELETLLAKL--IQTCQHVEVNASRQE 259
>YHFK_ECOLI (P45537) Hypothetical protein yhfK
Length = 700
Score = 29.3 bits (64), Expect = 8.5
Identities = 12/35 (34%), Positives = 20/35 (56%)
Query: 176 PFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLA 210
P +W RM+V+ A++ N N + +F + YLA
Sbjct: 544 PLAWQRMRVNQAHNTLYNSLNQAMQEPAFNSHYLA 578
>Y511_RICPR (Q9ZD36) Hypothetical protein RP511
Length = 950
Score = 29.3 bits (64), Expect = 8.5
Identities = 17/59 (28%), Positives = 30/59 (50%), Gaps = 4/59 (6%)
Query: 44 LKSFCKLFRKTLVSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLP 102
LK F KLF K + S +K+ ++E +S + +L +LD ++ +L K+P
Sbjct: 237 LKKFNKLFYKQEIMLTSFLKEVKAQSKPFLQEHFESYKIDLKILD----IIPTLLNKIP 291
>RL22_PARUW (Q6ME58) 50S ribosomal protein L22
Length = 111
Score = 29.3 bits (64), Expect = 8.5
Identities = 13/29 (44%), Positives = 21/29 (71%)
Query: 61 LVKKTLVSKVENVEEEVDSLQQNLSVLDL 89
L+KKTL S V N E ++D ++NL V+++
Sbjct: 47 LLKKTLDSAVANAETQLDMRRENLKVVEV 75
>OSTY_YEAST (Q03723) Dolichyl-diphosphooligosaccharide--protein
glycosyltransferase 37 kDa subunit (EC 2.4.1.119)
(Oligosaccharyl transferase 37 kDa subunit) (OTASE 37
kDa subunit)
Length = 332
Score = 29.3 bits (64), Expect = 8.5
Identities = 14/43 (32%), Positives = 24/43 (55%), Gaps = 4/43 (9%)
Query: 4 YFITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLFHKDFCLKS 46
YF+ C+ F+F+K ++ LP TN+ +L S++ L S
Sbjct: 192 YFVACMVVFIFIKKVI----LPKVTNKWKLFSMILSLGILLPS 230
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,411,244
Number of Sequences: 164201
Number of extensions: 1070505
Number of successful extensions: 3526
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 3520
Number of HSP's gapped (non-prelim): 15
length of query: 236
length of database: 59,974,054
effective HSP length: 107
effective length of query: 129
effective length of database: 42,404,547
effective search space: 5470186563
effective search space used: 5470186563
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC136507.1


