

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.16 - phase: 0
(212 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
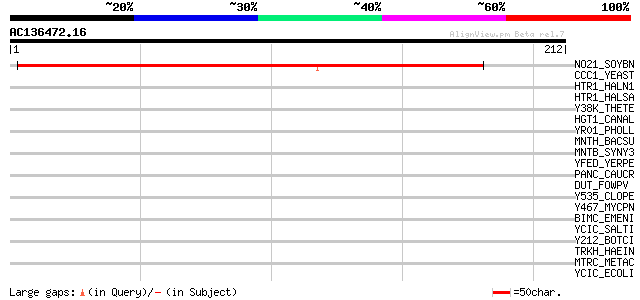
Score E
Sequences producing significant alignments: (bits) Value
NO21_SOYBN (P16313) Nodulin 21 (N-21) 183 3e-46
CCC1_YEAST (P47818) CCC1 protein 41 0.002
HTR1_HALN1 (P33741) Sensory rhodopsin I transducer (HTR-I) (Meth... 37 0.034
HTR1_HALSA (P33955) Sensory rhodopsin I transducer (HTR-I) (Meth... 36 0.057
Y38K_THETE (P05715) Hypothetical 38 kDa protein in 23S RNA operon 33 0.49
HGT1_CANAL (O74713) High-affinity glucose transporter 33 0.63
YR01_PHOLL (Q7N3L4) Hypothetical UPF0059 protein plu2701 32 1.1
MNTH_BACSU (P96593) Manganese transport protein mntH 32 1.1
MNTB_SYNY3 (Q55282) Manganese transport system membrane protein ... 32 1.1
YFED_YERPE (Q56955) Chelated iron transport system membrane prot... 32 1.4
PANC_CAUCR (Q9A6C8) Pantoate--beta-alanine ligase (EC 6.3.2.1) (... 31 1.8
DUT_FOWPV (Q9J5G5) Deoxyuridine 5'-triphosphate nucleotidohydrol... 31 1.8
Y535_CLOPE (Q8XN03) Hypothetical UPF0059 protein CPE0535 30 3.1
Y467_MYCPN (P75110) Hypothetical ABC transporter ATP-binding pro... 30 3.1
BIMC_EMENI (P17120) Kinesin-like protein bimC 30 4.1
YCIC_SALTI (Q8Z7E3) Hypothetical UPF0259 protein yciC 30 5.4
Y212_BOTCI (P56739) Hypothetical 21.2 kDa protein 30 5.4
TRKH_HAEIN (P44843) Trk system potassium uptake protein trkH 30 5.4
MTRC_METAC (Q8TU01) Tetrahydromethanopterin S-methyltransferase ... 30 5.4
YCIC_ECOLI (P21365) Hypothetical UPF0259 protein yciC 29 7.0
>NO21_SOYBN (P16313) Nodulin 21 (N-21)
Length = 206
Score = 183 bits (464), Expect = 3e-46
Identities = 96/189 (50%), Positives = 125/189 (65%), Gaps = 11/189 (5%)
Query: 4 IQHHHAATLARSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMM 63
+ H+H + + + + E++ DY QRAQWLRAA+LGANDGL+S ASLMM
Sbjct: 17 VPHNHVGAVLLTIPTIKIDGKQTLATEDHTSIDYLQRAQWLRAAILGANDGLVSVASLMM 76
Query: 64 GVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNIS------- 116
GVGAV +D K M+L G AGLVAGAC MAIGEFV+VY+QY++E QMKR N+S
Sbjct: 77 GVGAVKRDAKAMLLAGFAGLVAGACGMAIGEFVAVYTQYEVEVGQMKRDMNMSVGGERDL 136
Query: 117 ----QKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGL 172
++ LPNP A ASA+ F++GA VPLL AAF+++Y+ R+ VVV + LAL FG
Sbjct: 137 EMEMERRTLPNPLQATLASALCFSIGALVPLLSAAFIENYRTRIIVVVAMSCLALVVFGW 196
Query: 173 LSAVLGKAP 181
+ A LGK P
Sbjct: 197 VGAKLGKTP 205
>CCC1_YEAST (P47818) CCC1 protein
Length = 322
Score = 41.2 bits (95), Expect = 0.002
Identities = 21/65 (32%), Positives = 39/65 (59%), Gaps = 1/65 (1%)
Query: 48 VLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFA 107
++G +DGL +L G+ ++ D K +I G A L++GA SM +G ++ S+ D A
Sbjct: 105 IIGLSDGLTVPFALTAGLSSLG-DAKLVITGGFAELISGAISMGLGGYLGAKSESDYYHA 163
Query: 108 QMKRQ 112
++K++
Sbjct: 164 EVKKE 168
>HTR1_HALN1 (P33741) Sensory rhodopsin I transducer (HTR-I)
(Methyl-accepting phototaxis protein I) (MPP-I)
Length = 535
Score = 37.0 bits (84), Expect = 0.034
Identities = 16/31 (51%), Positives = 23/31 (73%)
Query: 54 GLLSTASLMMGVGAVTKDVKTMILTGIAGLV 84
G ++TA L++GVG T DV + I+ GIAGL+
Sbjct: 16 GYIATAGLLVGVGVTTNDVPSTIVAGIAGLL 46
>HTR1_HALSA (P33955) Sensory rhodopsin I transducer (HTR-I)
(Methyl-accepting phototaxis protein I) (MPP-I)
Length = 535
Score = 36.2 bits (82), Expect = 0.057
Identities = 22/65 (33%), Positives = 36/65 (54%), Gaps = 2/65 (3%)
Query: 54 GLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQG 113
G ++TA L++GVG T +V + I+ GIAGL+ S+ GE V + + + G
Sbjct: 16 GYIATAGLLVGVGVTTNEVPSTIVAGIAGLLT-LGSINAGETVGRIKEIGAQ-TERVANG 73
Query: 114 NISQK 118
N+ Q+
Sbjct: 74 NLEQE 78
>Y38K_THETE (P05715) Hypothetical 38 kDa protein in 23S RNA operon
Length = 373
Score = 33.1 bits (74), Expect = 0.49
Identities = 27/68 (39%), Positives = 38/68 (55%), Gaps = 13/68 (19%)
Query: 143 LLGAAF---VKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSL 199
L G AF VK Y+ +LG+ G+VSL GFGL SAV PL+ S +G + +L
Sbjct: 111 LYGIAFNLAVKWYQDKLGLATGLVSL---GFGLGSAVAN--PLIAS-----VGNYREATL 160
Query: 200 TFGLTKLV 207
G+ +L+
Sbjct: 161 AIGVVELL 168
>HGT1_CANAL (O74713) High-affinity glucose transporter
Length = 545
Score = 32.7 bits (73), Expect = 0.63
Identities = 22/76 (28%), Positives = 36/76 (46%), Gaps = 1/76 (1%)
Query: 129 FASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVG-VVSLALFGFGLLSAVLGKAPLVKSSL 187
F ++ AF ++GAA + R +++G ++S GFG A + A L +
Sbjct: 96 FGRRLSLLTCAFFWMVGAAIQSSVQNRAQLIIGRIISGIGVGFGSAVAPVYGAELAPRKI 155
Query: 188 RVLIGGWLAMSLTFGL 203
R LIGG +T G+
Sbjct: 156 RGLIGGMFQFFVTLGI 171
>YR01_PHOLL (Q7N3L4) Hypothetical UPF0059 protein plu2701
Length = 193
Score = 32.0 bits (71), Expect = 1.1
Identities = 38/164 (23%), Positives = 69/164 (41%), Gaps = 21/164 (12%)
Query: 59 ASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVY-SQYDIEF----------- 106
AS+ G + I TG+ +A AC+ IG + +Y SQY IE+
Sbjct: 20 ASICKGAALHRPHFREAIRTGLIFGLAEACTPLIGWSLGLYASQYIIEWDHWVAFTLLFI 79
Query: 107 ---AQMKRQGNISQKDKLPNPY----YAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVV 159
+ Q+ K +P +AIA ++ A +G AF++ V +
Sbjct: 80 LGCRMITESFKTKQEKKCESPCRHNSIVLITTAIATSLDAMAIGIGLAFLEVNIVHTAMA 139
Query: 160 VGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGL 203
+G++++ + G+L L K + LIGG + +++ F +
Sbjct: 140 IGMMTMIMATLGMLIGRYIGPRLGKRA--ELIGGLILIAIGFNI 181
>MNTH_BACSU (P96593) Manganese transport protein mntH
Length = 425
Score = 32.0 bits (71), Expect = 1.1
Identities = 24/66 (36%), Positives = 35/66 (52%), Gaps = 5/66 (7%)
Query: 130 ASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRV 189
A+A+ F G FV L AF + G +V +S ALFG GLL A L + + S V
Sbjct: 273 AAALFFKNGLFVEDLDVAFQQ-----FGHLVSPMSAALFGIGLLVAGLSSSSVGTLSGDV 327
Query: 190 LIGGWL 195
++ G++
Sbjct: 328 IMQGFI 333
>MNTB_SYNY3 (Q55282) Manganese transport system membrane protein
mntB
Length = 306
Score = 32.0 bits (71), Expect = 1.1
Identities = 26/76 (34%), Positives = 35/76 (45%), Gaps = 7/76 (9%)
Query: 128 AFASAIAFAVGAFVPLLGAAFVKDY---KVRL--GVVVGVVSLALFGFGLLSAVLGKAPL 182
A+A I FA+GAF GA Y K RL V+G+V F GL+ ++ K P
Sbjct: 67 AYALNIPFAIGAFTFGFGATVAIGYVKSKTRLKEDAVIGIVFTGFFALGLV--LVTKIPS 124
Query: 183 VKSSLRVLIGGWLAMS 198
+L G L +S
Sbjct: 125 NVDLFHILFGNVLGIS 140
>YFED_YERPE (Q56955) Chelated iron transport system membrane protein
yfeD
Length = 297
Score = 31.6 bits (70), Expect = 1.4
Identities = 23/78 (29%), Positives = 36/78 (45%), Gaps = 7/78 (8%)
Query: 128 AFASAIAFAVGAFVPLLGAAFVKDY-----KVRLGVVVGVVSLALFGFGLLSAVLGKAPL 182
AF I +GAFV + A Y +V+ V+G+V +F FGL+ + +
Sbjct: 61 AFWLGIPLVIGAFVSGIFCAVATGYLKENSRVKEDTVMGIVFSGMFAFGLV--LFSRIDT 118
Query: 183 VKSSLRVLIGGWLAMSLT 200
+ +L G L +SLT
Sbjct: 119 DQHLSHILFGNMLGISLT 136
>PANC_CAUCR (Q9A6C8) Pantoate--beta-alanine ligase (EC 6.3.2.1)
(Pantothenate synthetase) (Pantoate activating enzyme)
Length = 285
Score = 31.2 bits (69), Expect = 1.8
Identities = 26/100 (26%), Positives = 43/100 (43%), Gaps = 16/100 (16%)
Query: 123 NPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAV------ 176
N Y +A A A A+ + AA + + V +L GFG + V
Sbjct: 191 NAYLSAEERAAAVALPTAMKAAAAAVAQGGPIEDAERSAVAALQAAGFGQVDYVEIREAS 250
Query: 177 ----LGKAPLVKSSLRVLIGGWLAMSLTFGLTKLVNHVVV 212
LG P+ ++S R+L+ WL G T+L++++ V
Sbjct: 251 DLSRLGPGPIGEASGRILVAAWL------GKTRLIDNMAV 284
>DUT_FOWPV (Q9J5G5) Deoxyuridine 5'-triphosphate nucleotidohydrolase
(EC 3.6.1.23) (dUTPase) (dUTP pyrophosphatase)
Length = 145
Score = 31.2 bits (69), Expect = 1.8
Identities = 22/77 (28%), Positives = 33/77 (42%), Gaps = 12/77 (15%)
Query: 98 VYSQYDIEFAQMKRQGNISQKD---KLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKV 154
+YS YD M ++ + + D K+PN YY A A F+ + DY+
Sbjct: 28 LYSAYDYVIEPMNKE--LIKTDIILKIPNGYYGRIAPRSGLAYNYFIDVGAGVIDSDYRG 85
Query: 155 RLGVVVGVVSLALFGFG 171
+GV+ LF FG
Sbjct: 86 NVGVL-------LFNFG 95
>Y535_CLOPE (Q8XN03) Hypothetical UPF0059 protein CPE0535
Length = 213
Score = 30.4 bits (67), Expect = 3.1
Identities = 29/95 (30%), Positives = 46/95 (47%), Gaps = 16/95 (16%)
Query: 105 EFAQMKRQGNISQKDKLPNPYYAAFASAI-AFAVGAFVPLLGAAFVKDYKVRLGVVVGVV 163
EFA MKR+ +S K N A A++I A AVG LG + V+ +++G++
Sbjct: 112 EFANMKRKEELSAK----NLTVLAIATSIDALAVGVSFAFLGISIVQTI-----IIIGII 162
Query: 164 SLALFGFG-LLSAVLGK-----APLVKSSLRVLIG 192
+ L G ++ LG A +V + +LIG
Sbjct: 163 TFVLCFLGVIIGEKLGDIFKNYAEIVGGVILILIG 197
>Y467_MYCPN (P75110) Hypothetical ABC transporter ATP-binding
protein MG467 homolog (K05_orf339)
Length = 339
Score = 30.4 bits (67), Expect = 3.1
Identities = 28/91 (30%), Positives = 48/91 (51%), Gaps = 3/91 (3%)
Query: 110 KRQGNISQ-KDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALF 168
K+ G ISQ K+KL PY + V + + +D K ++ ++ V SL +
Sbjct: 83 KQGGLISQWKEKLEAPYRTPYVEQKQGMVISIDKMWKHVHGEDSKEQIAILSDV-SLEI- 140
Query: 169 GFGLLSAVLGKAPLVKSSLRVLIGGWLAMSL 199
G+G + +LG + K++L LIGG+ ++SL
Sbjct: 141 GYGEIVIILGPSGSGKTTLLNLIGGYDSISL 171
>BIMC_EMENI (P17120) Kinesin-like protein bimC
Length = 1184
Score = 30.0 bits (66), Expect = 4.1
Identities = 31/109 (28%), Positives = 43/109 (39%), Gaps = 7/109 (6%)
Query: 5 QHHHAATLARSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMG 64
Q HA L + KV + E D ++ + L AA A S M
Sbjct: 617 QTRHAKLLETTSVKVNEFIATEISNIERTRSDLSEYNRSLDAACNNAKAETSSAHEDMNN 676
Query: 65 VGAVTKD----VKTMILTGIAGLVAGACSMA---IGEFVSVYSQYDIEF 106
V KD VK+ + G+ GL A A ++ IGEF ++SQ F
Sbjct: 677 VLEEIKDLREEVKSKVGEGLNGLSAAAARISEEVIGEFTQLHSQLHTSF 725
>YCIC_SALTI (Q8Z7E3) Hypothetical UPF0259 protein yciC
Length = 247
Score = 29.6 bits (65), Expect = 5.4
Identities = 23/79 (29%), Positives = 39/79 (49%), Gaps = 3/79 (3%)
Query: 136 AVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAP--LVKSSLRVLIGG 193
A+GA P L F+ + L V +G++ + + G +++ VL AP LV+ + V
Sbjct: 118 AIGASAPALPKLFILIFLTTLLVQIGIMLIVVPGI-IMAIVLALAPVMLVEEKMGVFAAM 176
Query: 194 WLAMSLTFGLTKLVNHVVV 212
+M L + KLV V+
Sbjct: 177 RSSMRLAWANMKLVAPAVI 195
>Y212_BOTCI (P56739) Hypothetical 21.2 kDa protein
Length = 183
Score = 29.6 bits (65), Expect = 5.4
Identities = 12/26 (46%), Positives = 20/26 (76%)
Query: 19 VYNMDVEKKGEEENDEKDYTQRAQWL 44
V+ V++KGE ++E+D+TQRA W+
Sbjct: 137 VHPNTVDEKGELLDEERDHTQRAIWM 162
>TRKH_HAEIN (P44843) Trk system potassium uptake protein trkH
Length = 487
Score = 29.6 bits (65), Expect = 5.4
Identities = 16/47 (34%), Positives = 30/47 (63%), Gaps = 7/47 (14%)
Query: 158 VVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLT 204
+ +G+VSL+L+G+GL+S + + K +L++ +MS+T G T
Sbjct: 287 IFIGIVSLSLYGYGLVSDI--NEAVTKGALQL-----TSMSMTAGYT 326
>MTRC_METAC (Q8TU01) Tetrahydromethanopterin S-methyltransferase
subunit C (EC 2.1.1.86)
(N5-methyltetrahydromethanopterin--coenzyme M
methyltransferase subunit C)
Length = 267
Score = 29.6 bits (65), Expect = 5.4
Identities = 20/85 (23%), Positives = 38/85 (44%), Gaps = 7/85 (8%)
Query: 127 AAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSS 186
A F ++ G V + AA + G V+GV++ + G G+ + +
Sbjct: 87 ALFGLSVGGIAGPIVSFIAAAII-------GAVIGVLANKVIGMGIPIMEQAMVEIAGAG 139
Query: 187 LRVLIGGWLAMSLTFGLTKLVNHVV 211
V+IG + ++ TF ++V +VV
Sbjct: 140 TLVIIGLSVVIAGTFDYAEVVEYVV 164
>YCIC_ECOLI (P21365) Hypothetical UPF0259 protein yciC
Length = 247
Score = 29.3 bits (64), Expect = 7.0
Identities = 22/79 (27%), Positives = 41/79 (51%), Gaps = 3/79 (3%)
Query: 136 AVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAP--LVKSSLRVLIGG 193
A+GA P+L F+ + L V +G++ + + G +++ +L AP LV+ + V
Sbjct: 118 AIGASAPILPKLFILIFLTTLLVQIGIMLVVVPGI-IMAILLALAPVMLVQDKMGVFASM 176
Query: 194 WLAMSLTFGLTKLVNHVVV 212
+M LT+ +LV V+
Sbjct: 177 RSSMRLTWANMRLVAPAVL 195
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.135 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,938,099
Number of Sequences: 164201
Number of extensions: 819852
Number of successful extensions: 2931
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 2922
Number of HSP's gapped (non-prelim): 32
length of query: 212
length of database: 59,974,054
effective HSP length: 106
effective length of query: 106
effective length of database: 42,568,748
effective search space: 4512287288
effective search space used: 4512287288
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC136472.16


