

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.12 - phase: 0 /pseudo
(346 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
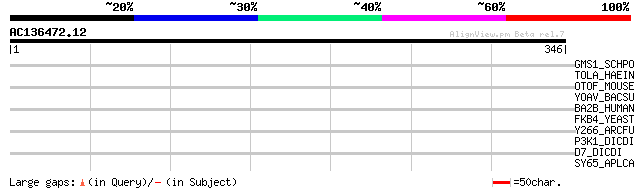
Score E
Sequences producing significant alignments: (bits) Value
GMS1_SCHPO (P87041) UDP-galactose transporter (Golgi UDP-Gal tra... 35 0.21
TOLA_HAEIN (P44678) TolA protein 35 0.28
OTOF_MOUSE (Q9ESF1) Otoferlin (Fer-1 like protein 2) 35 0.28
YOAV_BACSU (O34416) Hypothetical transport protein yoaV 34 0.47
BA2B_HUMAN (Q9UIF8) Bromodomain adjacent to zinc finger domain 2... 33 0.81
FKB4_YEAST (Q06205) FK506-binding protein 4 (EC 5.2.1.8) (Peptid... 32 2.4
Y266_ARCFU (O29973) Hypothetical transport protein AF0266 31 5.2
P3K1_DICDI (P54673) Phosphatidylinositol 3-kinase 1 (EC 2.7.1.13... 30 6.8
D7_DICDI (P54682) cAMP-inducible prespore protein D7 precursor 30 6.8
SY65_APLCA (P41823) Synaptotagmin (p65) 30 8.9
>GMS1_SCHPO (P87041) UDP-galactose transporter (Golgi UDP-Gal
transporter)
Length = 353
Score = 35.4 bits (80), Expect = 0.21
Identities = 16/88 (18%), Positives = 42/88 (47%)
Query: 211 YYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSII 270
Y S+ + I++ +++ CV + + T + +++ +LA +++ K+ L +I
Sbjct: 251 YNSIVWLAILLQAGGGIIVALCVAFADNIMKNFSTSISIIISSLASVYLMDFKISLTFLI 310
Query: 271 GAVLIVCGLYAVVWGKSKEMKKKNQLVP 298
G +L++ + +SK + +P
Sbjct: 311 GVMLVIAATFLYTKPESKPSPSRGTYIP 338
>TOLA_HAEIN (P44678) TolA protein
Length = 372
Score = 35.0 bits (79), Expect = 0.28
Identities = 28/114 (24%), Positives = 52/114 (45%), Gaps = 15/114 (13%)
Query: 242 SVFTPLMLLMVALAGCTILNEKLHL----------GSIIGAVLIVCGLYAVVWGKSKEMK 291
+ F +LL L G IL+ H G +IGAV++ G A WG+ ++ +
Sbjct: 11 NAFAISILLHFILFGLLILSSLYHTVEIMGGGEGEGDVIGAVIVDTGTAAQEWGRIQQ-Q 69
Query: 292 KKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDNEDSQTDEHEQQSQ 345
KK Q K P +PV + + E ++ + + ++ + E + E ++Q +
Sbjct: 70 KKGQADKQKRP---EPV-VEEKPPEPNQEEIKHQQEVQRQEELKRQQEQQRQQE 119
>OTOF_MOUSE (Q9ESF1) Otoferlin (Fer-1 like protein 2)
Length = 1997
Score = 35.0 bits (79), Expect = 0.28
Identities = 18/59 (30%), Positives = 33/59 (55%)
Query: 288 KEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDNEDSQTDEHEQQSQE 346
KE KKK + PS+ P E +P E ++ K ++++ + ++ +E S TD E++ E
Sbjct: 1302 KERKKKKKKGPSEEPEEEEPDESMLDWWSKYFASIDTMKEQLRQHETSGTDLEEKEEME 1360
>YOAV_BACSU (O34416) Hypothetical transport protein yoaV
Length = 292
Score = 34.3 bits (77), Expect = 0.47
Identities = 18/71 (25%), Positives = 38/71 (53%)
Query: 210 AYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSI 269
A +S+ + G++ +G V W ++ AS+ + ++ G L+E++ + I
Sbjct: 211 AVWSLLFNGLLSTGFTFVVWFWVLNQIQASKASMALMFVPVLALFFGWLQLHEQITINII 270
Query: 270 IGAVLIVCGLY 280
+GA+LI CG++
Sbjct: 271 LGALLICCGIF 281
>BA2B_HUMAN (Q9UIF8) Bromodomain adjacent to zinc finger domain 2B
(hWALp4)
Length = 1972
Score = 33.5 bits (75), Expect = 0.81
Identities = 18/65 (27%), Positives = 30/65 (45%), Gaps = 4/65 (6%)
Query: 282 VVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDNEDSQTDEHE 341
++ + KE N + P K H P + +V S++ ++ KD+EDS DE E
Sbjct: 362 ILHSQGKEKAVSNNVNPVKTQHHSHPAKSLVEQFRGTDSDIPSS----KDSEDSNEDEEE 417
Query: 342 QQSQE 346
+E
Sbjct: 418 DDEEE 422
>FKB4_YEAST (Q06205) FK506-binding protein 4 (EC 5.2.1.8)
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase)
Length = 392
Score = 32.0 bits (71), Expect = 2.4
Identities = 17/49 (34%), Positives = 26/49 (52%)
Query: 288 KEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDNEDSQ 336
K+ KKKN SK HE D E + +K + + + VKD+E+S+
Sbjct: 230 KDEKKKNNKKDSKRKHEEDEEESAKPAEKKQTTKKDKKAEKVKDSEESK 278
>Y266_ARCFU (O29973) Hypothetical transport protein AF0266
Length = 276
Score = 30.8 bits (68), Expect = 5.2
Identities = 17/43 (39%), Positives = 27/43 (62%)
Query: 241 ASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLIVCGLYAVV 283
ASVF + ++ LAG +L E L L ++ G+ L++ G+Y VV
Sbjct: 231 ASVFLLAIPVVSLLAGNILLAEPLTLRTVAGSGLVLLGIYIVV 273
>P3K1_DICDI (P54673) Phosphatidylinositol 3-kinase 1 (EC 2.7.1.137)
(PI3-kinase) (PtdIns-3-kinase) (PI3K)
Length = 1570
Score = 30.4 bits (67), Expect = 6.8
Identities = 17/59 (28%), Positives = 31/59 (51%), Gaps = 5/59 (8%)
Query: 293 KNQLVPSKNPHEFDPVEIVVRSIEKDKS-----NLNNNNQMVKDNEDSQTDEHEQQSQE 346
KN S N + + +EI+ K+ + N NNNN + K +DS+ ++++ +QE
Sbjct: 34 KNNSTSSNNNNNHNNIEIIGIDNNKNNNKNNNDNNNNNNNIDKKRKDSKNKQNQEINQE 92
>D7_DICDI (P54682) cAMP-inducible prespore protein D7 precursor
Length = 850
Score = 30.4 bits (67), Expect = 6.8
Identities = 15/53 (28%), Positives = 32/53 (60%), Gaps = 1/53 (1%)
Query: 293 KNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDNEDSQTDEHEQQSQ 345
+N+ +K +E D +E+++ E+++ + N N Q + +NE+ Q + +QQ Q
Sbjct: 687 ENEQDAAKLQNEMDQLELLLEENEEEQFS-NKNLQQLNENENQQQQQGQQQVQ 738
>SY65_APLCA (P41823) Synaptotagmin (p65)
Length = 426
Score = 30.0 bits (66), Expect = 8.9
Identities = 21/74 (28%), Positives = 40/74 (53%), Gaps = 2/74 (2%)
Query: 269 IIGAVL-IVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQ 327
I G++L +VC +Y V ++ KKK K + V+++ S K+K +L+
Sbjct: 77 IAGSLLFLVCCVYCVCRRSCRKRKKKEGKKGLKGAVDLKSVQLLGNSY-KEKPDLDELPV 135
Query: 328 MVKDNEDSQTDEHE 341
++DNED+++ + E
Sbjct: 136 NMEDNEDAESTKSE 149
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.334 0.145 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,266,930
Number of Sequences: 164201
Number of extensions: 1573197
Number of successful extensions: 7002
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 6983
Number of HSP's gapped (non-prelim): 21
length of query: 346
length of database: 59,974,054
effective HSP length: 111
effective length of query: 235
effective length of database: 41,747,743
effective search space: 9810719605
effective search space used: 9810719605
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 66 (30.0 bits)
Medicago: description of AC136472.12


