

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136449.11 - phase: 0 /pseudo
(982 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
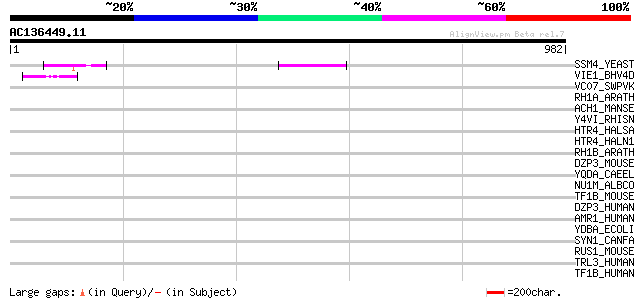
Score E
Sequences producing significant alignments: (bits) Value
SSM4_YEAST (P40318) SSM4 protein 89 4e-17
VIE1_BHV4D (P27426) 32.7 kDa immediate early protein IE1 47 2e-04
VC07_SWPVK (P32225) Hypothetical protein C7 44 0.002
RH1A_ARATH (Q9SUS4) RING-H2 zinc finger protein RHA1a 44 0.002
ACH1_MANSE (P91766) Acetylcholine receptor protein, alpha-like c... 39 0.052
Y4VI_RHISN (Q53217) Putative short-chain type dehydrogenase/redu... 38 0.15
HTR4_HALSA (Q48317) Halobacterial transducer protein IV 38 0.15
HTR4_HALN1 (Q9HP84) Halobacterial transducer protein IV 38 0.15
RH1B_ARATH (Q9SUS5) RING-H2 zinc finger protein RHA1b 37 0.20
DZP3_MOUSE (Q7TPV2) Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) ... 37 0.20
YQDA_CAEEL (Q09268) Hypothetical RING finger protein C32D5.10 in... 37 0.34
NU1M_ALBCO (P48897) NADH-ubiquinone oxidoreductase chain 1 (EC 1... 37 0.34
TF1B_MOUSE (Q62318) Transcription intermediary factor 1-beta (TI... 36 0.44
DZP3_HUMAN (Q86Y13) Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) ... 36 0.44
AMR1_HUMAN (Q9Y4X0) AMME syndrome candidate gene 1 protein 36 0.44
YDBA_ECOLI (P33666) Hypothetical protein ydbA 35 0.75
SYN1_CANFA (O62732) Synapsin-1 (Synapsin I) (Fragment) 35 0.98
RUS1_MOUSE (Q8BG26) RUN and SH3 domain containing protein 1 35 1.3
TRL3_HUMAN (Q9HCF6) Long transient receptor potential channel 3 ... 34 1.7
TF1B_HUMAN (Q13263) Transcription intermediary factor 1-beta (TI... 34 1.7
>SSM4_YEAST (P40318) SSM4 protein
Length = 1319
Score = 89.4 bits (220), Expect = 4e-17
Identities = 41/119 (34%), Positives = 63/119 (52%), Gaps = 14/119 (11%)
Query: 60 DDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNAR--------Q 111
+DD CRICR DNPL +PC C GSIK++H+ CLL+W+ N +
Sbjct: 30 NDDAPSGATCRICRGEATEDNPLFHPCKCRGSIKYMHESCLLEWVASKNIDISKPGADVK 89
Query: 112 CEVCKHPFSFSPVYAENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFI 170
C++C +P F +YAEN P ++PF + + +L FF + L++ L + +I
Sbjct: 90 CDICHYPIQFKTIYAENMPEKIPFS------LLLSKSILTFFEKARLALTIGLAAVLYI 142
Score = 65.9 bits (159), Expect = 5e-10
Identities = 38/124 (30%), Positives = 62/124 (49%), Gaps = 4/124 (3%)
Query: 476 KVAFLLVIELGVFPLMCGWWLDV---CTIQMFGKTMVHRAQFFSASPLASSLAHWVVGIV 532
KV L IEL FP++ G LD C I M+ + P S +W +G +
Sbjct: 625 KVFTLFFIELAGFPILAGVMLDFSLFCPILASNSRMLWVPSICAIWPPFSLFVYWTIGTL 684
Query: 533 YMLQISIFVSLLR-GVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVYGS 591
YM + ++ ++R ++R GVL+F+R P DPN D + +P+ R+ LS+ +Y
Sbjct: 685 YMYWFAKYIGMIRKNIIRPGVLFFIRSPEDPNIKILHDSLIHPMSIQLSRLCLSMFIYAI 744
Query: 592 LMVM 595
+V+
Sbjct: 745 FIVL 748
>VIE1_BHV4D (P27426) 32.7 kDa immediate early protein IE1
Length = 285
Score = 47.4 bits (111), Expect = 2e-04
Identities = 24/97 (24%), Positives = 47/97 (47%), Gaps = 6/97 (6%)
Query: 23 SSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPL 82
S P +S +P + + E + D++ + C ICR+ G++
Sbjct: 90 SKQPKHTSKNPTKGEIQYFPVEKCKDIHRVENQSSIDEEGKQ----CWICRD-GESLPEA 144
Query: 83 RYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPF 119
RY C C G +++ H++CL W++ S ++C+ C+ P+
Sbjct: 145 RY-CNCYGDLQYCHEECLKTWISMSGEKKCKFCQTPY 180
>VC07_SWPVK (P32225) Hypothetical protein C7
Length = 155
Score = 44.3 bits (103), Expect = 0.002
Identities = 27/105 (25%), Positives = 47/105 (44%), Gaps = 19/105 (18%)
Query: 66 EDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVY 125
+ VC IC++ + C C K VH +C+ +W+ +S R C++C ++ V
Sbjct: 2 DPVCWICKDDYSIEKNY---CNCKNEYKVVHDECMKKWIQYSRERSCKLCNKEYNIISV- 57
Query: 126 AENAPARLPFQEFVVGMAMKACHVLQFFVRLSFVLSVWLLIIPFI 170
R PF ++V ++K C + S +L L + FI
Sbjct: 58 ------RKPFSQWV--FSIKDC-------KKSAILYATLFLCTFI 87
>RH1A_ARATH (Q9SUS4) RING-H2 zinc finger protein RHA1a
Length = 159
Score = 44.3 bits (103), Expect = 0.002
Identities = 22/84 (26%), Positives = 39/84 (46%), Gaps = 3/84 (3%)
Query: 45 AVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWL 104
+ + A+ P ++ D D ED C +C + ++D+ +R C H CL +W+
Sbjct: 62 SASLANELIPVVRFSDLPTDPEDCCTVCLSDFESDDKVRQLPKCG---HVFHHYCLDRWI 118
Query: 105 NHSNARQCEVCKHPFSFSPVYAEN 128
N +C VC+H F Y ++
Sbjct: 119 VDYNKMKCPVCRHRFLPKEKYTQS 142
>ACH1_MANSE (P91766) Acetylcholine receptor protein, alpha-like
chain precursor (MARA1)
Length = 516
Score = 39.3 bits (90), Expect = 0.052
Identities = 20/75 (26%), Positives = 38/75 (50%), Gaps = 6/75 (8%)
Query: 721 CYVIWTAVAGVRYSIEQIRKRRTSVLLNQIWKWCSIVVKSSALLSIWIFVIPVLI---GL 777
C + A+ G+ Y +Q RK S + + WK+ ++V+ L WIF + V++ G+
Sbjct: 426 CPELHKAIDGINYIADQTRKEEESTRVKEDWKYVAMVLDRPFL---WIFTLAVVVGSAGI 482
Query: 778 LFELLVIVPMRVPVD 792
+ + + R P+D
Sbjct: 483 ILQAPTLYDERAPID 497
>Y4VI_RHISN (Q53217) Putative short-chain type
dehydrogenase/reductase y4vI (EC 1.-.-.-)
Length = 548
Score = 37.7 bits (86), Expect = 0.15
Identities = 35/124 (28%), Positives = 55/124 (44%), Gaps = 16/124 (12%)
Query: 248 GARVARRPAGQANRNVNGDANGEDAVAAQGVAGAGQVIRRNAENVAARWEMQAARLEAHV 307
GA +ARR A + V D +GE+AV G+ G + RR V E V
Sbjct: 290 GAAIARRFAENGDTVVIADGDGEEAVKLAGLLGDKHLSRRVDRTV-----------ETEV 338
Query: 308 EQMFDGLDDADGAEDVPFDELVGMQVPPTDAS----LSLANITLKNALTAVQNLSTATQE 363
+F+ L + G DV + + + VP T+ S + ++ L A T V+ + +
Sbjct: 339 VSLFEELRERFGHLDVFVNGMNEILVPNTEESPEVLKRILDVNLTGAFTCVRE-AAISMR 397
Query: 364 SGSI 367
SGS+
Sbjct: 398 SGSV 401
>HTR4_HALSA (Q48317) Halobacterial transducer protein IV
Length = 810
Score = 37.7 bits (86), Expect = 0.15
Identities = 47/209 (22%), Positives = 79/209 (37%), Gaps = 26/209 (12%)
Query: 206 GFLLSASIVFIFFGGTSLRDYFRHLREIGGQDAE-------------REDEVDRNGARVA 252
GF+L A + + GGT R+ ++ + AE R DE+ + A +
Sbjct: 327 GFILVALVGVVLVGGTIGRNTAAAVQSLSAAAAEIEAGNYDVDVASSRRDEIGQLFASIG 386
Query: 253 RRPAGQANRNVNGDANGEDAVAAQGVAGA----GQVIRRNAENVAARWEMQAARLEAHVE 308
+ +A E A AQ A A + R AE+ A E AA LEA E
Sbjct: 387 SMRDALVTQIDEAEAAREQATEAQQDAEAERERAEDARERAEDAKADAEALAAELEAQAE 446
Query: 309 QMFD---GLDDADGAEDVPFDELVGMQVPPTDASLSLANITLKNALTAVQNLSTATQESG 365
+ D D D +P D+ + AS + ++ + +Q + A
Sbjct: 447 RYSDVMAACADGDLTRRMPADDTDNEAMAAIAASFNEMLAQWEHTIIDIQEFADA----- 501
Query: 366 SIGQIAEMLKVNASELSEMSNNITASVSD 394
+ +E +V A++ S ++ SV +
Sbjct: 502 -VATASEEAEVGAADAERASGQVSESVQE 529
>HTR4_HALN1 (Q9HP84) Halobacterial transducer protein IV
Length = 810
Score = 37.7 bits (86), Expect = 0.15
Identities = 47/209 (22%), Positives = 79/209 (37%), Gaps = 26/209 (12%)
Query: 206 GFLLSASIVFIFFGGTSLRDYFRHLREIGGQDAE-------------REDEVDRNGARVA 252
GF+L A + + GGT R+ ++ + AE R DE+ + A +
Sbjct: 327 GFILVALVGVVLVGGTIGRNTAAAVQSLSAAAAEIEAGNYDVDVASSRRDEIGQLFASIG 386
Query: 253 RRPAGQANRNVNGDANGEDAVAAQGVAGA----GQVIRRNAENVAARWEMQAARLEAHVE 308
+ +A E A AQ A A + R AE+ A E AA LEA E
Sbjct: 387 SMRDALVTQIDEAEAAREQATEAQQDAEAERERAEDARERAEDAKADAEALAAELEAQAE 446
Query: 309 QMFD---GLDDADGAEDVPFDELVGMQVPPTDASLSLANITLKNALTAVQNLSTATQESG 365
+ D D D +P D+ + AS + ++ + +Q + A
Sbjct: 447 RYSDVMAACADGDLTRRMPADDTDNEAMAAIAASFNEMLAQWEHTIIDIQEFADA----- 501
Query: 366 SIGQIAEMLKVNASELSEMSNNITASVSD 394
+ +E +V A++ S ++ SV +
Sbjct: 502 -VATASEEAEVGAADAERASGQVSESVQE 529
>RH1B_ARATH (Q9SUS5) RING-H2 zinc finger protein RHA1b
Length = 157
Score = 37.4 bits (85), Expect = 0.20
Identities = 21/79 (26%), Positives = 37/79 (46%), Gaps = 7/79 (8%)
Query: 45 AVATASTAP----PSAKYDDDDEDEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCL 100
A++T++T P ++ D D ED C +C + +D+ +R C H CL
Sbjct: 57 ALSTSATLANELIPVVRFSDLLTDPEDCCTVCLSDFVSDDKIRQLPKCG---HVFHHRCL 113
Query: 101 LQWLNHSNARQCEVCKHPF 119
+W+ N C +C++ F
Sbjct: 114 DRWIVDCNKITCPICRNRF 132
>DZP3_MOUSE (Q7TPV2) Ubiquitin ligase protein DZIP3 (EC 6.3.2.-)
(DAZ-interacting protein 3 homolog)
Length = 1204
Score = 37.4 bits (85), Expect = 0.20
Identities = 29/109 (26%), Positives = 45/109 (40%), Gaps = 12/109 (11%)
Query: 8 PPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEED 67
P + +A +++SPS ++S P + + A S K D++E+EE+
Sbjct: 1088 PKKSESEEKSAQDGNNASPSHTASQPNAPQDPKS-----AQGSATWEGDKDMDNEEEEEE 1142
Query: 68 VCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCK 116
C IC +N PCA KF H C+ WL C C+
Sbjct: 1143 PCVICHENLSPENLSVLPCA----HKF-HSQCIRPWLMQQGT--CPTCR 1184
>YQDA_CAEEL (Q09268) Hypothetical RING finger protein C32D5.10 in
chromosome II
Length = 610
Score = 36.6 bits (83), Expect = 0.34
Identities = 27/97 (27%), Positives = 45/97 (45%), Gaps = 12/97 (12%)
Query: 25 SPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPLRY 84
SPS+S S + + ++ + T P +K + D+ +EE C +C+N L
Sbjct: 3 SPSTSESPLKKNTIQDFGESNM----TESPQSKEEIDEPEEE--CSVCKNEIIDTTSLSD 56
Query: 85 PCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSF 121
C +F + DC++ WL + C +CK P SF
Sbjct: 57 CCH-----EFCY-DCIVGWLTKGSGPFCPMCKTPVSF 87
>NU1M_ALBCO (P48897) NADH-ubiquinone oxidoreductase chain 1 (EC
1.6.5.3)
Length = 299
Score = 36.6 bits (83), Expect = 0.34
Identities = 18/60 (30%), Positives = 30/60 (50%), Gaps = 5/60 (8%)
Query: 848 LWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLGCLSFSFVCFC 907
+W L ++ I+ L+ + + ++ ARGV+P Y L++N W L FS C C
Sbjct: 237 VWFLYSDMIFIMTLLILLIAMAFLFARGVYPRHRYDLLMN-----LCWKSFLPFSLCCIC 291
>TF1B_MOUSE (Q62318) Transcription intermediary factor 1-beta
(TIF1-beta) (Tripartite motif-containing protein 28)
(KRAB-A interacting protein) (KRIP-1)
Length = 834
Score = 36.2 bits (82), Expect = 0.44
Identities = 26/97 (26%), Positives = 38/97 (38%), Gaps = 14/97 (14%)
Query: 21 PSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADN 80
P + P +S S S G E+ A V +A + P +D +CR+C+ PGD
Sbjct: 585 PGAEGPRLASPSGSTSSGLEVVAPEVTSAPVSGPGIL-----DDSATICRVCQKPGDL-- 637
Query: 81 PLRYPCACSGSIKFVHQDCLLQWLNHSNARQ--CEVC 115
C+ H DC L L + C +C
Sbjct: 638 -----VMCNQCEFCFHLDCHLPALQDVPGEEWSCSLC 669
>DZP3_HUMAN (Q86Y13) Ubiquitin ligase protein DZIP3 (EC 6.3.2.-)
(DAZ-interacting protein 3) (RNA-binding ubiquitin ligase
of 138 kDa) (hRUL138)
Length = 1208
Score = 36.2 bits (82), Expect = 0.44
Identities = 26/101 (25%), Positives = 41/101 (39%), Gaps = 11/101 (10%)
Query: 16 IAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNP 75
+ +PS S ++ P + + ++ AT A D++E+EE+ C IC
Sbjct: 1099 VNCVSPSHSPSQPDAAQPPKPAWRPLTSQGPATWE----GASNPDEEEEEEEPCVICHEN 1154
Query: 76 GDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCK 116
+N PCA KF H C+ WL C C+
Sbjct: 1155 LSPENLSVLPCA----HKF-HAQCIRPWLMQQGT--CPTCR 1188
>AMR1_HUMAN (Q9Y4X0) AMME syndrome candidate gene 1 protein
Length = 333
Score = 36.2 bits (82), Expect = 0.44
Identities = 23/47 (48%), Positives = 28/47 (58%), Gaps = 5/47 (10%)
Query: 3 IANEPPPS-----LDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAE 44
IA PPPS L TP AAT+ S SS S++SSS GS+ + AE
Sbjct: 83 IALSPPPSCGVGTLLSTPAAATSSSPSSSSAASSSSPGSRKMVVSAE 129
>YDBA_ECOLI (P33666) Hypothetical protein ydbA
Length = 2003
Score = 35.4 bits (80), Expect = 0.75
Identities = 46/187 (24%), Positives = 77/187 (40%), Gaps = 21/187 (11%)
Query: 262 NVNGDANGEDAVAAQGVAGAGQVIRRNAENVAARWEMQAARLEAHVEQMFDGLDDADGAE 321
++NG+ N +AA+ + GQ A + + + +V + D AD A
Sbjct: 634 DLNGN-NNSVTLAAKDLKVVGQ----KATGINVSGDANTVNITGNV--LVDKDKTADNAA 686
Query: 322 DVPFDELVGMQVPPTDASLSLANITLKNALTAVQNLSTATQESGSIGQIAEMLKVNASEL 381
+ FD VG+ V +D N+TL LT V + +++S AE S L
Sbjct: 687 EYFFDPSVGINVYGSD-----NNVTLDGKLTVVSDSEVTSRQSNLFDGSAE----KTSGL 737
Query: 382 SEMSNNITASVSDDL-LKGGSIGTSRISDVTTLAXGYIFLSTLIFCYFGVVALIRYTKGE 440
+ + T +++ L L G + S VT+L GY + S ++ V Y G+
Sbjct: 738 VVIGDGNTVNMNGGLELIGEKNALADGSQVTSLRTGYSYTSVIVVSGESSV----YLNGD 793
Query: 441 PLTAGRF 447
+G F
Sbjct: 794 TTISGEF 800
>SYN1_CANFA (O62732) Synapsin-1 (Synapsin I) (Fragment)
Length = 415
Score = 35.0 bits (79), Expect = 0.98
Identities = 16/45 (35%), Positives = 24/45 (52%)
Query: 13 GTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAK 57
G P AA P+S SP + P+ ++ + +A AS APPS +
Sbjct: 269 GAPPAARPPASPSPQRQAGPPQATRQTSVSGQAPPKASGAPPSGQ 313
>RUS1_MOUSE (Q8BG26) RUN and SH3 domain containing protein 1
Length = 893
Score = 34.7 bits (78), Expect = 1.3
Identities = 22/76 (28%), Positives = 35/76 (45%), Gaps = 9/76 (11%)
Query: 2 EIANEPPPSLD-GTPIAATTPSSSSPSSSSSSPRGSKG--------KEIDAEAVATASTA 52
E AN P +L+ G+P+A PS+ SP S SP G + + A+ T
Sbjct: 129 EDANPQPSTLELGSPLAPAGPSTCSPDSFCCSPDSCSGISSPPGPDLDSNCNALTTCQDL 188
Query: 53 PPSAKYDDDDEDEEDV 68
P +++D E+D+
Sbjct: 189 PSPGLEEEEDSGEQDL 204
>TRL3_HUMAN (Q9HCF6) Long transient receptor potential channel 3
(LTrpC3) (Fragment)
Length = 1017
Score = 34.3 bits (77), Expect = 1.7
Identities = 29/111 (26%), Positives = 50/111 (44%), Gaps = 17/111 (15%)
Query: 14 TPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICR 73
TP+ +T PSSS+ ++ + + R + ID E + + T S+ Y E C+
Sbjct: 743 TPVPSTAPSSSAYATLAPTDR-PPSRSIDFEDITSMDTRSFSSDYTHLPE--------CQ 793
Query: 74 NPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCK-HPFSFSP 123
NP D++ P+ + S S +++ L + + K H F FSP
Sbjct: 794 NPWDSEPPMYHTIERSKSSRYLATTPFL-------LEEAPIVKSHSFMFSP 837
>TF1B_HUMAN (Q13263) Transcription intermediary factor 1-beta
(TIF1-beta) (Tripartite motif-containing protein 28)
(Nuclear corepressor KAP-1) (KRAB-associated protein 1)
(KAP-1) (KRAB-interacting protein 1) (KRIP-1)
Length = 835
Score = 34.3 bits (77), Expect = 1.7
Identities = 29/114 (25%), Positives = 41/114 (35%), Gaps = 14/114 (12%)
Query: 4 ANEPPPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDE 63
A E P + A P + P +S S S G E+ A +A P +
Sbjct: 568 ATEGPETKPVLMALAEGPGAEGPRLASPSGSTSSGLEVVAPEGTSAPGGGPGTL-----D 622
Query: 64 DEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQ--CEVC 115
D +CR+C+ PGD C+ H DC L L + C +C
Sbjct: 623 DSATICRVCQKPGDL-------VMCNQCEFCFHLDCHLPALQDVPGEEWSCSLC 669
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.141 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 108,200,855
Number of Sequences: 164201
Number of extensions: 4480329
Number of successful extensions: 22898
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 46
Number of HSP's that attempted gapping in prelim test: 22569
Number of HSP's gapped (non-prelim): 213
length of query: 982
length of database: 59,974,054
effective HSP length: 120
effective length of query: 862
effective length of database: 40,269,934
effective search space: 34712683108
effective search space used: 34712683108
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 71 (32.0 bits)
Medicago: description of AC136449.11


