

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
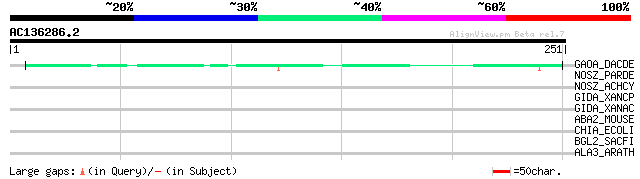
Query= AC136286.2 - phase: 0
(251 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
GAOA_DACDE (Q01745) Galactose oxidase precursor (EC 1.1.3.9) (GAO) 53 6e-07
NOSZ_PARDE (Q51705) Nitrous-oxide reductase precursor (EC 1.7.99... 35 0.22
NOSZ_ACHCY (P94127) Nitrous-oxide reductase precursor (EC 1.7.99... 33 0.85
GIDA_XANCP (Q8PDG1) Glucose inhibited division protein A 33 0.85
GIDA_XANAC (Q8PQE8) Glucose inhibited division protein A 31 2.5
ABA2_MOUSE (Q9D6J4) Amyloid beta A4 protein-binding family A mem... 31 2.5
CHIA_ECOLI (P13656) Probable bifunctional chitinase/lysozyme pre... 29 9.4
BGL2_SACFI (P22507) Beta-glucosidase 2 precursor (EC 3.2.1.21) (... 29 9.4
ALA3_ARATH (Q9XIE6) Potential phospholipid-transporting ATPase 3... 29 9.4
>GAOA_DACDE (Q01745) Galactose oxidase precursor (EC 1.1.3.9) (GAO)
Length = 680
Score = 53.1 bits (126), Expect = 6e-07
Identities = 62/246 (25%), Positives = 93/246 (37%), Gaps = 50/246 (20%)
Query: 8 RVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFELLAPASTP 67
R ++LP G+ I G G ++ PV P +Y P D F P S
Sbjct: 480 RTFHTSVVLPDGSTFITGGQRRGIPFEDST--PVFTPEIYVPEQDT----FYKQNPNSIV 533
Query: 68 RMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPELDTL--RPVIV 125
R+YHS ++LLPDGR+ GG D + + P+YL L RP I
Sbjct: 534 RVYHSISLLLPDGRVFNGGGG--LCGDCTTNH---FDAQIFTPNYLYNSNGNLATRPKIT 588
Query: 126 AVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLEVTALEE 185
+ + S D + + S++ TH+ +QR + L +T
Sbjct: 589 RTSTQSVKVGGRITIST--------DSSISKASLIRYGTATHTVNTDQRRIPLTLT---- 636
Query: 186 VVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIH-VGIPSV 244
N + + P VA PGY+MLFV++ G+PSV
Sbjct: 637 ------------------------NNGGNSYSFQVPSDSGVALPGYWMLFVMNSAGVPSV 672
Query: 245 ATWVHV 250
A+ + V
Sbjct: 673 ASTIRV 678
>NOSZ_PARDE (Q51705) Nitrous-oxide reductase precursor (EC 1.7.99.6)
(N(2)OR) (N2O reductase)
Length = 652
Score = 34.7 bits (78), Expect = 0.22
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 3/75 (4%)
Query: 149 VKDVNRIRVSM--VAPSFTTHSFAMNQ-RLLFLEVTALEEVVNSMQDQNFGEFGFGSSLG 205
++D N++RV M VAPSF+ SF + + + + VT L+E+ + G +G +G
Sbjct: 550 IRDGNKVRVYMSSVAPSFSIESFTVKEGDEVTVIVTNLDEIDDLTHGFTMGNYGVAMEIG 609
Query: 206 PGKIANSVYKATVRG 220
P ++ + A G
Sbjct: 610 PQMTSSVTFVAANPG 624
>NOSZ_ACHCY (P94127) Nitrous-oxide reductase precursor (EC 1.7.99.6)
(N(2)OR) (N2O reductase)
Length = 642
Score = 32.7 bits (73), Expect = 0.85
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 3/75 (4%)
Query: 149 VKDVNRIRVSM--VAPSFTTHSFAMNQ-RLLFLEVTALEEVVNSMQDQNFGEFGFGSSLG 205
++D N++RV M VAPSF+ SF + + + + VT L+E+ + G G +G
Sbjct: 540 IRDGNKVRVYMTSVAPSFSQPSFTVKEGDEVTVIVTNLDEIDDLTHGFTMGNHGVAMEVG 599
Query: 206 PGKIANSVYKATVRG 220
P + ++ + A G
Sbjct: 600 PQQTSSVTFVAANPG 614
>GIDA_XANCP (Q8PDG1) Glucose inhibited division protein A
Length = 634
Score = 32.7 bits (73), Expect = 0.85
Identities = 26/81 (32%), Positives = 39/81 (48%), Gaps = 7/81 (8%)
Query: 26 GAANGTAGWENAANPVLYPVLYKPGLDNPFMKFELLAPASTPRMYHSSAVLLPDGRILVG 85
G NGT G+E AA L L N + + L PA +PR + +L D I G
Sbjct: 375 GQINGTTGYEEAAAQGLLAGL------NAARQAQAL-PAWSPRRDEAYLGVLVDDLITHG 427
Query: 86 GSNPHRLYDFQAKYPTELSLD 106
+ P+R++ +A+Y +L D
Sbjct: 428 TTEPYRMFTSRAEYRLQLRED 448
>GIDA_XANAC (Q8PQE8) Glucose inhibited division protein A
Length = 634
Score = 31.2 bits (69), Expect = 2.5
Identities = 26/81 (32%), Positives = 38/81 (46%), Gaps = 7/81 (8%)
Query: 26 GAANGTAGWENAANPVLYPVLYKPGLDNPFMKFELLAPASTPRMYHSSAVLLPDGRILVG 85
G NGT G+E AA L L N + L PA +PR + +L D I G
Sbjct: 375 GQINGTTGYEEAAAQGLLAGL------NAARHVQGL-PAWSPRRDEAYLGVLVDDLITHG 427
Query: 86 GSNPHRLYDFQAKYPTELSLD 106
+ P+R++ +A+Y +L D
Sbjct: 428 TTEPYRMFTSRAEYRLQLRED 448
>ABA2_MOUSE (Q9D6J4) Amyloid beta A4 protein-binding family A member
2 binding protein (X11L-binding protein 51) (mXB51)
Length = 353
Score = 31.2 bits (69), Expect = 2.5
Identities = 26/80 (32%), Positives = 35/80 (43%), Gaps = 15/80 (18%)
Query: 111 DYLRPELDTLRPVIVAVEVVN---------STLSYESL-----FSVSFLLRE-VKDVNRI 155
DY L RPV+ A+E +N + L YE F FLLRE V + +
Sbjct: 88 DYFSTHLGVYRPVLAALESLNRAVLTAMDTTKLEYEQASKVDQFVTRFLLRETVNQLQAL 147
Query: 156 RVSMVAPSFTTHSFAMNQRL 175
+ S+ S T + A QRL
Sbjct: 148 QTSLEGASDTLEAQAHGQRL 167
>CHIA_ECOLI (P13656) Probable bifunctional chitinase/lysozyme
precursor [Includes: Chitinase (EC 3.2.1.14); Lysozyme
(EC 3.2.1.17)]
Length = 897
Score = 29.3 bits (64), Expect = 9.4
Identities = 30/121 (24%), Positives = 57/121 (46%), Gaps = 10/121 (8%)
Query: 47 YKPGLDNPFMKFELLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLD 106
+ PG DNP+ K+E PA + + +S V + R++V G L+ Q+ P ++
Sbjct: 445 FVPGGDNPWKKYE---PA---KAWSASTVYVKGDRVVVDGQAYEALFWTQSDNPALVANQ 498
Query: 107 AYYPDYLRPELDTLRPVIVAVEVVNSTLSY--ESLFSVSFLLR--EVKDVNRIRVSMVAP 162
RP + + E +N+ + E+L++ L+R V +++ +V V+P
Sbjct: 499 NATGSNSRPWKPLGKAQSYSNEELNNAPQFNPETLYASDTLIRFNGVNYISQSKVQKVSP 558
Query: 163 S 163
S
Sbjct: 559 S 559
>BGL2_SACFI (P22507) Beta-glucosidase 2 precursor (EC 3.2.1.21)
(Gentiobiase) (Cellobiase) (Beta-D-glucoside
glucohydrolase)
Length = 880
Score = 29.3 bits (64), Expect = 9.4
Identities = 31/128 (24%), Positives = 53/128 (41%), Gaps = 22/128 (17%)
Query: 101 TELSLDAYYPD------YLRPELDTLRP----VIVAVEVVNSTLSYESLFSVSFLLREVK 150
T+ S+ A PD YL P D++R V+ + VN+T S E+ + ++ LL+E
Sbjct: 231 TKESISANIPDRAMHELYLWPFADSIRAGVGSVMCSYNRVNNTYSCENSYMINHLLKE-- 288
Query: 151 DVNRIRVSMVAPSFTTHSFAMNQRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIA 210
+ F +A + ++ L+ SM + G + G S +
Sbjct: 289 -------ELGFQGFVVSDWAAQMSGAYSAISGLD---MSMPGELLGGWNTGKSYWGQNLT 338
Query: 211 NSVYKATV 218
+VY TV
Sbjct: 339 KAVYNETV 346
>ALA3_ARATH (Q9XIE6) Potential phospholipid-transporting ATPase 3
(EC 3.6.3.1) (Aminophospholipid flippase 3)
Length = 1213
Score = 29.3 bits (64), Expect = 9.4
Identities = 12/30 (40%), Positives = 19/30 (63%)
Query: 43 YPVLYKPGLDNPFMKFELLAPASTPRMYHS 72
YP LY+ G+ N F K+ ++A +T +Y S
Sbjct: 967 YPELYREGIRNSFFKWRVVAVWATSAVYQS 996
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,454,403
Number of Sequences: 164201
Number of extensions: 1353185
Number of successful extensions: 2877
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 2871
Number of HSP's gapped (non-prelim): 10
length of query: 251
length of database: 59,974,054
effective HSP length: 108
effective length of query: 143
effective length of database: 42,240,346
effective search space: 6040369478
effective search space used: 6040369478
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC136286.2


