

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136142.6 + phase: 0
(147 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
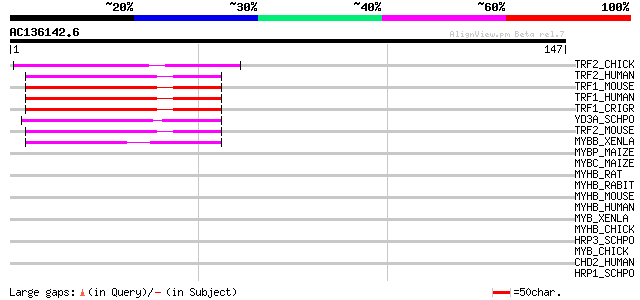
Score E
Sequences producing significant alignments: (bits) Value
TRF2_CHICK (Q9PU53) Telomeric repeat binding factor 2 (TTAGGG re... 48 9e-06
TRF2_HUMAN (Q15554) Telomeric repeat binding factor 2 (TTAGGG re... 46 3e-05
TRF1_MOUSE (P70371) Telomeric repeat binding factor 1 (TTAGGG re... 45 4e-05
TRF1_HUMAN (P54274) Telomeric repeat binding factor 1 (TTAGGG re... 45 6e-05
TRF1_CRIGR (O55036) Telomeric repeat binding factor 1 (TTAGGG re... 45 6e-05
YD3A_SCHPO (Q10274) Hypothetical protein C13G7.10 in chromosome I 45 7e-05
TRF2_MOUSE (O35144) Telomeric repeat binding factor 2 (TTAGGG re... 43 3e-04
MYBB_XENLA (P52551) Myb-related protein B (B-Myb) (Myb-related p... 42 4e-04
MYBP_MAIZE (P27898) Myb-related protein P 36 0.026
MYBC_MAIZE (P10290) Anthocyanin regulatory C1 protein 35 0.058
MYHB_RAT (Q63862) Myosin heavy chain, smooth muscle isoform (SMM... 35 0.076
MYHB_RABIT (P35748) Myosin heavy chain, smooth muscle isoform (S... 35 0.076
MYHB_MOUSE (O08638) Myosin heavy chain, smooth muscle isoform (S... 35 0.076
MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform (S... 35 0.076
MYB_XENLA (Q08759) Myb protein 35 0.076
MYHB_CHICK (P10587) Myosin heavy chain, gizzard smooth muscle 34 0.099
HRP3_SCHPO (O14139) Chromodomain helicase hrp3 34 0.099
MYB_CHICK (P01103) Myb proto-oncogene protein (C-myb) 34 0.13
CHD2_HUMAN (O14647) Chromodomain-helicase-DNA-binding protein 2 ... 34 0.13
HRP1_SCHPO (Q9US25) Chromodomain helicase hrp1 33 0.17
>TRF2_CHICK (Q9PU53) Telomeric repeat binding factor 2 (TTAGGG
repeat binding factor 2) (Telomeric DNA binding protein)
Length = 718
Score = 47.8 bits (112), Expect = 9e-06
Identities = 23/60 (38%), Positives = 36/60 (59%), Gaps = 4/60 (6%)
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTAI 61
G+ KQKWT +E +K GV K+G G+W+TI F + R++V +KD++R + I
Sbjct: 662 GSKKQKWTVQESEWIKDGVRKYGEGRWKTISEKYPF----QNRTSVQIKDRYRTMKKLGI 717
>TRF2_HUMAN (Q15554) Telomeric repeat binding factor 2 (TTAGGG
repeat binding factor 2) (Telomeric DNA binding protein)
Length = 500
Score = 46.2 bits (108), Expect = 3e-05
Identities = 23/52 (44%), Positives = 31/52 (59%), Gaps = 4/52 (7%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
KQKWT EE +KAGV K+G G W I + F + R+ V +KD+WR +
Sbjct: 447 KQKWTVEESEWVKAGVQKYGEGNWAAISKNYPFVN----RTAVMIKDRWRTM 494
>TRF1_MOUSE (P70371) Telomeric repeat binding factor 1 (TTAGGG
repeat binding factor 1)
Length = 421
Score = 45.4 bits (106), Expect = 4e-05
Identities = 22/52 (42%), Positives = 32/52 (61%), Gaps = 4/52 (7%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
+Q W EE+ LK GV K+G G W IL +F++ R++V LKD+WR +
Sbjct: 367 RQTWLWEEDRILKCGVKKYGEGNWAKILSHYKFNN----RTSVMLKDRWRTM 414
>TRF1_HUMAN (P54274) Telomeric repeat binding factor 1 (TTAGGG
repeat binding factor 1) (NIMA-interacting protein 2)
(Telomeric protein Pin2/TRF1)
Length = 439
Score = 45.1 bits (105), Expect = 6e-05
Identities = 21/52 (40%), Positives = 34/52 (65%), Gaps = 4/52 (7%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
+Q W EE+ L++GV K+G G W IL+ +F++ R++V LKD+WR +
Sbjct: 380 RQAWLWEEDKNLRSGVRKYGEGNWSKILLHYKFNN----RTSVMLKDRWRTM 427
>TRF1_CRIGR (O55036) Telomeric repeat binding factor 1 (TTAGGG
repeat binding factor 1) (Fragment)
Length = 438
Score = 45.1 bits (105), Expect = 6e-05
Identities = 21/52 (40%), Positives = 34/52 (65%), Gaps = 4/52 (7%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
+Q W EE+ L++GV K+G G W IL+ +F++ R++V LKD+WR +
Sbjct: 380 RQAWLWEEDKNLRSGVRKYGEGNWSKILLHYKFNN----RTSVMLKDRWRTM 427
>YD3A_SCHPO (Q10274) Hypothetical protein C13G7.10 in chromosome I
Length = 390
Score = 44.7 bits (104), Expect = 7e-05
Identities = 23/53 (43%), Positives = 29/53 (54%), Gaps = 2/53 (3%)
Query: 4 PKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
P+ KWT +E L G HG G W+ IL+D F RS DLKD++R I
Sbjct: 54 PRVKWTEKETNDLLRGCQIHGVGNWKKILLDERFH--FTNRSPNDLKDRFRTI 104
Score = 30.0 bits (66), Expect = 1.9
Identities = 23/72 (31%), Positives = 34/72 (46%), Gaps = 7/72 (9%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTAIWGS 64
++++T EE+ L G HG W I D L+ R + DL+D++RN A
Sbjct: 140 RKQFTPEEDERLLEGFFLHGPC-WTRISKDANLG--LQNRRSTDLRDRFRN----AFPER 192
Query: 65 RQKAKLALKNSP 76
A LKN+P
Sbjct: 193 YAAAGFKLKNNP 204
>TRF2_MOUSE (O35144) Telomeric repeat binding factor 2 (TTAGGG
repeat binding factor 2) (Telomeric DNA binding protein)
Length = 495
Score = 42.7 bits (99), Expect = 3e-04
Identities = 22/52 (42%), Positives = 29/52 (55%), Gaps = 4/52 (7%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
KQKWT EE +K GV K+G G W I F + R+ V +KD+WR +
Sbjct: 442 KQKWTIEESEWVKDGVRKYGEGNWAAISKSYPFVN----RTAVMIKDRWRTM 489
>MYBB_XENLA (P52551) Myb-related protein B (B-Myb) (Myb-related
protein 1) (XMYB1)
Length = 743
Score = 42.4 bits (98), Expect = 4e-04
Identities = 21/52 (40%), Positives = 29/52 (55%), Gaps = 6/52 (11%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
K KWT EE+ LKA V KHG G+W+TI +S L R+ + +W +
Sbjct: 31 KVKWTPEEDETLKALVKKHGQGEWKTI------ASNLNNRTEQQCQHRWLRV 76
>MYBP_MAIZE (P27898) Myb-related protein P
Length = 399
Score = 36.2 bits (82), Expect = 0.026
Identities = 18/55 (32%), Positives = 31/55 (55%), Gaps = 5/55 (9%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
+G + +WTAEE+ L + +HG G WR++ P+ + +LR + L +W N
Sbjct: 10 VGLKRGRWTAEEDQLLANYIAEHGEGSWRSL---PKNAGLLRCGKSCRL--RWIN 59
>MYBC_MAIZE (P10290) Anthocyanin regulatory C1 protein
Length = 273
Score = 35.0 bits (79), Expect = 0.058
Identities = 19/54 (35%), Positives = 29/54 (53%), Gaps = 5/54 (9%)
Query: 2 GAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRN 55
G + WT++E+ AL A V HG GKWR + P+ + + R + L +W N
Sbjct: 11 GVKRGAWTSKEDDALAAYVKAHGEGKWREV---PQKAGLRRCGKSCRL--RWLN 59
>MYHB_RAT (Q63862) Myosin heavy chain, smooth muscle isoform (SMMHC)
(Fragments)
Length = 1327
Score = 34.7 bits (78), Expect = 0.076
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 5/67 (7%)
Query: 18 AGVVKHGAGKWRTILMDP---EFSSILRTRSNVDLKDKWRNINVTAIWGSRQKAKLALKN 74
AG V + A W T MDP +S+L S+ + D W++++ I G Q AK+ +
Sbjct: 444 AGKVDYNASAWLTKNMDPLNDNVTSLLNASSDKFVADLWKDVD--RIVGLDQMAKMTESS 501
Query: 75 SPPAPKT 81
P A KT
Sbjct: 502 LPSASKT 508
>MYHB_RABIT (P35748) Myosin heavy chain, smooth muscle isoform
(SMMHC)
Length = 1972
Score = 34.7 bits (78), Expect = 0.076
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 5/67 (7%)
Query: 18 AGVVKHGAGKWRTILMDP---EFSSILRTRSNVDLKDKWRNINVTAIWGSRQKAKLALKN 74
AG V + A W T MDP +S+L S+ + D W++++ I G Q AK+ +
Sbjct: 581 AGKVDYNASAWLTKNMDPLNDNVTSLLNASSDKFVADLWKDVD--RIVGLDQMAKMTESS 638
Query: 75 SPPAPKT 81
P A KT
Sbjct: 639 LPSASKT 645
>MYHB_MOUSE (O08638) Myosin heavy chain, smooth muscle isoform
(SMMHC)
Length = 1972
Score = 34.7 bits (78), Expect = 0.076
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 5/67 (7%)
Query: 18 AGVVKHGAGKWRTILMDP---EFSSILRTRSNVDLKDKWRNINVTAIWGSRQKAKLALKN 74
AG V + A W T MDP +S+L S+ + D W++++ I G Q AK+ +
Sbjct: 581 AGKVDYNASAWLTKNMDPLNDNVTSLLNASSDKFVADLWKDVD--RIVGLDQMAKMTESS 638
Query: 75 SPPAPKT 81
P A KT
Sbjct: 639 LPSASKT 645
>MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform
(SMMHC)
Length = 1972
Score = 34.7 bits (78), Expect = 0.076
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 5/67 (7%)
Query: 18 AGVVKHGAGKWRTILMDP---EFSSILRTRSNVDLKDKWRNINVTAIWGSRQKAKLALKN 74
AG V + A W T MDP +S+L S+ + D W++++ I G Q AK+ +
Sbjct: 581 AGKVDYNASAWLTKNMDPLNDNVTSLLNASSDKFVADLWKDVD--RIVGLDQMAKMTESS 638
Query: 75 SPPAPKT 81
P A KT
Sbjct: 639 LPSASKT 645
>MYB_XENLA (Q08759) Myb protein
Length = 624
Score = 34.7 bits (78), Expect = 0.076
Identities = 16/52 (30%), Positives = 29/52 (55%), Gaps = 6/52 (11%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
K +WT EE+ LK V ++G +W+ I +S L R++V + +W+ +
Sbjct: 37 KTRWTREEDEKLKKLVEQNGTEEWKVI------ASFLPNRTDVQCQHRWQKV 82
>MYHB_CHICK (P10587) Myosin heavy chain, gizzard smooth muscle
Length = 1978
Score = 34.3 bits (77), Expect = 0.099
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 5/67 (7%)
Query: 18 AGVVKHGAGKWRTILMDP---EFSSILRTRSNVDLKDKWRNINVTAIWGSRQKAKLALKN 74
AG V + A W T MDP +S+L S+ + D W++++ I G Q AK+ +
Sbjct: 586 AGKVTYNASAWLTKNMDPLNDNVTSLLNQSSDKFVADLWKDVD--RIVGLDQMAKMTESS 643
Query: 75 SPPAPKT 81
P A KT
Sbjct: 644 LPSASKT 650
>HRP3_SCHPO (O14139) Chromodomain helicase hrp3
Length = 1388
Score = 34.3 bits (77), Expect = 0.099
Identities = 16/45 (35%), Positives = 25/45 (55%), Gaps = 4/45 (8%)
Query: 7 KWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKD 51
+W E++ L +G+ KHG G W I DPE L+ + + L+D
Sbjct: 1122 QWGPREDSMLLSGICKHGFGAWLEIRDDPE----LKMKDKIFLED 1162
>MYB_CHICK (P01103) Myb proto-oncogene protein (C-myb)
Length = 641
Score = 33.9 bits (76), Expect = 0.13
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 6/52 (11%)
Query: 5 KQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNI 56
K +WT EE+ LK V ++G W+ I +S L R++V + +W+ +
Sbjct: 40 KTRWTREEDEKLKKLVEQNGTEDWKVI------ASFLPNRTDVQCQHRWQKV 85
>CHD2_HUMAN (O14647) Chromodomain-helicase-DNA-binding protein 2
(CHD-2)
Length = 1739
Score = 33.9 bits (76), Expect = 0.13
Identities = 12/30 (40%), Positives = 18/30 (60%)
Query: 7 KWTAEEEAALKAGVVKHGAGKWRTILMDPE 36
+W E+++ L G+ +HG G W I DPE
Sbjct: 1260 EWGVEDDSRLLLGIYEHGYGNWELIKTDPE 1289
>HRP1_SCHPO (Q9US25) Chromodomain helicase hrp1
Length = 1373
Score = 33.5 bits (75), Expect = 0.17
Identities = 13/29 (44%), Positives = 19/29 (64%)
Query: 8 WTAEEEAALKAGVVKHGAGKWRTILMDPE 36
W +E++ L AG+ KHG G W+ I DP+
Sbjct: 1132 WGIKEDSMLLAGINKHGFGCWQAIKNDPD 1160
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.130 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,692,893
Number of Sequences: 164201
Number of extensions: 677301
Number of successful extensions: 1241
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 1185
Number of HSP's gapped (non-prelim): 78
length of query: 147
length of database: 59,974,054
effective HSP length: 100
effective length of query: 47
effective length of database: 43,553,954
effective search space: 2047035838
effective search space used: 2047035838
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC136142.6


