

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135801.4 + phase: 0
(61 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
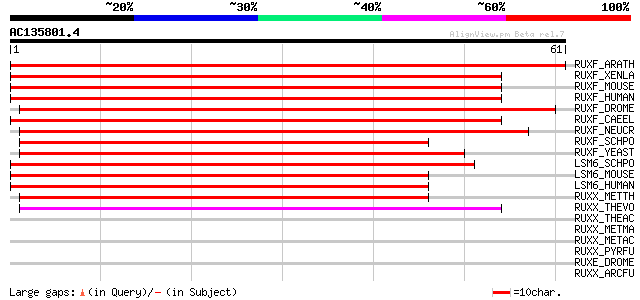
Score E
Sequences producing significant alignments: (bits) Value
RUXF_ARATH (Q9SUM2) Probable small nuclear ribonucleoprotein F (... 110 9e-25
RUXF_XENLA (P62321) Small nuclear ribonucleoprotein F (snRNP-F) ... 95 3e-20
RUXF_MOUSE (P62307) Small nuclear ribonucleoprotein F (snRNP-F) ... 95 3e-20
RUXF_HUMAN (P62306) Small nuclear ribonucleoprotein F (snRNP-F) ... 95 3e-20
RUXF_DROME (Q24297) Small nuclear ribonucleoprotein F (snRNP-F) ... 91 4e-19
RUXF_CAEEL (P34659) Probable small nuclear ribonucleoprotein F (... 86 2e-17
RUXF_NEUCR (Q9P5Z8) Probable small nuclear ribonucleoprotein F (... 73 1e-13
RUXF_SCHPO (O59734) Probable small nuclear ribonucleoprotein F (... 69 2e-12
RUXF_YEAST (P54999) Small nuclear ribonucleoprotein F (snRNP-F) ... 65 3e-11
LSM6_SCHPO (Q9UUI1) U6 snRNA-associated Sm-like protein LSm6 52 3e-07
LSM6_MOUSE (P62313) U6 snRNA-associated Sm-like protein LSm6 50 8e-07
LSM6_HUMAN (P62312) U6 snRNA-associated Sm-like protein LSm6 (Sm... 50 8e-07
RUXX_METTH (O26745) Putative snRNP Sm-like protein 43 2e-04
RUXX_THEVO (Q97BU5) Putative snRNP Sm-like protein 40 9e-04
RUXX_THEAC (P57670) Putative snRNP Sm-like protein 39 0.003
RUXX_METMA (Q8PZZ9) Putative snRNP Sm-like protein 39 0.003
RUXX_METAC (Q8TL47) Putative snRNP Sm-like protein 37 0.007
RUXX_PYRFU (Q8U0P4) Putative snRNP Sm-like protein 36 0.016
RUXE_DROME (Q9VLV5) Probable small nuclear ribonucleoprotein E (... 36 0.016
RUXX_ARCFU (O29386) Putative snRNP Sm-like protein 35 0.037
>RUXF_ARATH (Q9SUM2) Probable small nuclear ribonucleoprotein F
(snRNP-F) (Sm protein F) (Sm-F) (SmF)
Length = 88
Score = 110 bits (274), Expect = 9e-25
Identities = 51/61 (83%), Positives = 55/61 (89%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEEIEDAAD 60
MEYKG+L SVDSYMNLQL NTEEYI+G TGNLGEILIRCNNVLY+RGVPEDEE+EDA
Sbjct: 28 MEYKGFLASVDSYMNLQLGNTEEYIDGQLTGNLGEILIRCNNVLYVRGVPEDEELEDADQ 87
Query: 61 D 61
D
Sbjct: 88 D 88
>RUXF_XENLA (P62321) Small nuclear ribonucleoprotein F (snRNP-F)
(Sm protein F) (Sm-F) (SmF)
Length = 86
Score = 95.1 bits (235), Expect = 3e-20
Identities = 43/54 (79%), Positives = 50/54 (91%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEE 54
MEYKGYLVSVD YMN+QLANTEEYI+G +G+LGE+LIRCNNVLY+RGV E+EE
Sbjct: 27 MEYKGYLVSVDGYMNMQLANTEEYIDGALSGHLGEVLIRCNNVLYIRGVEEEEE 80
>RUXF_MOUSE (P62307) Small nuclear ribonucleoprotein F (snRNP-F)
(Sm protein F) (Sm-F) (SmF)
Length = 86
Score = 95.1 bits (235), Expect = 3e-20
Identities = 43/54 (79%), Positives = 50/54 (91%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEE 54
MEYKGYLVSVD YMN+QLANTEEYI+G +G+LGE+LIRCNNVLY+RGV E+EE
Sbjct: 27 MEYKGYLVSVDGYMNMQLANTEEYIDGALSGHLGEVLIRCNNVLYIRGVEEEEE 80
>RUXF_HUMAN (P62306) Small nuclear ribonucleoprotein F (snRNP-F)
(Sm protein F) (Sm-F) (SmF)
Length = 86
Score = 95.1 bits (235), Expect = 3e-20
Identities = 43/54 (79%), Positives = 50/54 (91%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEE 54
MEYKGYLVSVD YMN+QLANTEEYI+G +G+LGE+LIRCNNVLY+RGV E+EE
Sbjct: 27 MEYKGYLVSVDGYMNMQLANTEEYIDGALSGHLGEVLIRCNNVLYIRGVEEEEE 80
>RUXF_DROME (Q24297) Small nuclear ribonucleoprotein F (snRNP-F)
(Sm protein F) (Sm-F) (SmF) (Membrane-associated
protein Deb-B)
Length = 88
Score = 91.3 bits (225), Expect = 4e-19
Identities = 41/59 (69%), Positives = 51/59 (85%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEEIEDAAD 60
EYKG+LVSVD YMN+QLANTEE IEG+ TGNLGE+LIRCNNVLY++G+ +D+E + D
Sbjct: 30 EYKGFLVSVDGYMNMQLANTEEVIEGSVTGNLGEVLIRCNNVLYIKGMEDDDEEGEMRD 88
>RUXF_CAEEL (P34659) Probable small nuclear ribonucleoprotein F
(snRNP-F) (Sm protein F) (Sm-F) (SmF)
Length = 85
Score = 85.9 bits (211), Expect = 2e-17
Identities = 41/54 (75%), Positives = 46/54 (84%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEE 54
MEYKG LV+VDSYMNLQLA+ EEYI+GN GNLGEILIRCNNVLY+ GV + E
Sbjct: 29 MEYKGVLVAVDSYMNLQLAHAEEYIDGNSQGNLGEILIRCNNVLYVGGVDGENE 82
>RUXF_NEUCR (Q9P5Z8) Probable small nuclear ribonucleoprotein F
(snRNP-F) (Sm protein F) (Sm-F) (SmF)
Length = 90
Score = 73.2 bits (178), Expect = 1e-13
Identities = 34/56 (60%), Positives = 43/56 (76%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEEIED 57
EY G LVS+DSYMN+QL++T+EYI FTG LG++LIRCNNVLY++ E E D
Sbjct: 30 EYVGRLVSIDSYMNIQLSDTKEYINRKFTGALGQVLIRCNNVLYIKKADEAETSGD 85
>RUXF_SCHPO (O59734) Probable small nuclear ribonucleoprotein F
(snRNP-F) (Sm protein F) (Sm-F) (SmF)
Length = 78
Score = 68.9 bits (167), Expect = 2e-12
Identities = 33/45 (73%), Positives = 38/45 (84%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
EYKG L SVDSYMNLQL N EE ++G TG+LGEILIRCNNVL++
Sbjct: 29 EYKGTLQSVDSYMNLQLLNAEELVDGVKTGDLGEILIRCNNVLWV 73
>RUXF_YEAST (P54999) Small nuclear ribonucleoprotein F (snRNP-F)
(Sm protein F) (Sm-F) (SmF)
Length = 86
Score = 65.1 bits (157), Expect = 3e-11
Identities = 30/49 (61%), Positives = 36/49 (73%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVP 50
EY+G LVS D+Y NLQL EE++ G G LGEI IRCNNVLY+R +P
Sbjct: 37 EYRGTLVSTDNYFNLQLNEAEEFVAGVSHGTLGEIFIRCNNVLYIRELP 85
>LSM6_SCHPO (Q9UUI1) U6 snRNA-associated Sm-like protein LSm6
Length = 75
Score = 52.0 bits (123), Expect = 3e-07
Identities = 24/51 (47%), Positives = 32/51 (62%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPE 51
++YKG L +D YMNL L TEEY+ G T G+ IR NNVLY+ + +
Sbjct: 25 VDYKGILSCLDGYMNLALERTEEYVNGKKTNVYGDAFIRGNNVLYVSALDD 75
>LSM6_MOUSE (P62313) U6 snRNA-associated Sm-like protein LSm6
Length = 80
Score = 50.4 bits (119), Expect = 8e-07
Identities = 21/46 (45%), Positives = 29/46 (62%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
++Y+G L +D YMN+ L TEEY+ G G+ IR NNVLY+
Sbjct: 28 VDYRGVLACLDGYMNIALEQTEEYVNGQLKNKYGDAFIRGNNVLYI 73
>LSM6_HUMAN (P62312) U6 snRNA-associated Sm-like protein LSm6 (Sm
protein F)
Length = 80
Score = 50.4 bits (119), Expect = 8e-07
Identities = 21/46 (45%), Positives = 29/46 (62%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
++Y+G L +D YMN+ L TEEY+ G G+ IR NNVLY+
Sbjct: 28 VDYRGVLACLDGYMNIALEQTEEYVNGQLKNKYGDAFIRGNNVLYI 73
>RUXX_METTH (O26745) Putative snRNP Sm-like protein
Length = 81
Score = 42.7 bits (99), Expect = 2e-04
Identities = 20/45 (44%), Positives = 30/45 (66%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
E++G L S D +MNL L + EE +G T LG +LIR +N++Y+
Sbjct: 35 EFRGVLKSFDLHMNLVLNDAEELEDGEVTRRLGTVLIRGDNIVYI 79
>RUXX_THEVO (Q97BU5) Putative snRNP Sm-like protein
Length = 83
Score = 40.4 bits (93), Expect = 9e-04
Identities = 21/53 (39%), Positives = 27/53 (50%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEE 54
EY G L D YMN+ L N E I G G IL+R +NV+++ D E
Sbjct: 31 EYSGILEGYDVYMNVVLQNASEIINGENKGVFDRILVRGDNVIFVSPSKGDNE 83
>RUXX_THEAC (P57670) Putative snRNP Sm-like protein
Length = 83
Score = 38.9 bits (89), Expect = 0.003
Identities = 18/45 (40%), Positives = 25/45 (55%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
EY G L D YMN+ L N E I G G +L+R +NV+++
Sbjct: 31 EYSGILEGYDVYMNIVLQNASEIINGENKGVYDRVLVRGDNVIFV 75
>RUXX_METMA (Q8PZZ9) Putative snRNP Sm-like protein
Length = 72
Score = 38.5 bits (88), Expect = 0.003
Identities = 18/45 (40%), Positives = 26/45 (57%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
E++G L D +MNL L N EE EG ++IR +NV+Y+
Sbjct: 26 EFRGELKGYDIHMNLVLDNAEELREGEVVSKFSSVVIRGDNVVYV 70
>RUXX_METAC (Q8TL47) Putative snRNP Sm-like protein
Length = 72
Score = 37.4 bits (85), Expect = 0.007
Identities = 17/45 (37%), Positives = 26/45 (57%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
E++G L D +MNL L N EE +G ++IR +NV+Y+
Sbjct: 26 EFRGELKGYDIHMNLVLDNAEELRDGEVVSKFSSVVIRGDNVVYV 70
>RUXX_PYRFU (Q8U0P4) Putative snRNP Sm-like protein
Length = 76
Score = 36.2 bits (82), Expect = 0.016
Identities = 17/43 (39%), Positives = 27/43 (62%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVL 44
E++G L+ D ++N+ LAN E +G G+I+IR +NVL
Sbjct: 26 EFRGKLIGYDIHLNVVLANAELLQDGEVVKKYGKIVIRGDNVL 68
>RUXE_DROME (Q9VLV5) Probable small nuclear ribonucleoprotein E
(snRNP-E) (Sm protein E) (Sm-E) (SmE)
Length = 94
Score = 36.2 bits (82), Expect = 0.016
Identities = 17/50 (34%), Positives = 31/50 (62%), Gaps = 1/50 (2%)
Query: 1 MEYKGYLVSVDSYMNLQLANTEE-YIEGNFTGNLGEILIRCNNVLYMRGV 49
+ +G++V D YMNL L + EE Y++ NLG I+++ +N+ ++ V
Sbjct: 40 LRIEGHIVGFDEYMNLVLDDAEEVYVKTRQRRNLGRIMLKGDNITLIQNV 89
>RUXX_ARCFU (O29386) Putative snRNP Sm-like protein
Length = 77
Score = 35.0 bits (79), Expect = 0.037
Identities = 17/52 (32%), Positives = 28/52 (53%)
Query: 2 EYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDE 53
E++G L D +MNL L + EE G +G ++IR + V+++ P E
Sbjct: 26 EFRGTLDGYDIHMNLVLLDAEEIQNGEVVRKVGSVVIRGDTVVFVSPAPGGE 77
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.137 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,520,755
Number of Sequences: 164201
Number of extensions: 250005
Number of successful extensions: 435
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 406
Number of HSP's gapped (non-prelim): 45
length of query: 61
length of database: 59,974,054
effective HSP length: 37
effective length of query: 24
effective length of database: 53,898,617
effective search space: 1293566808
effective search space used: 1293566808
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135801.4


