

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135799.3 + phase: 0
(242 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
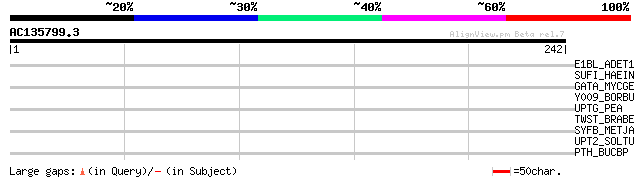
Score E
Sequences producing significant alignments: (bits) Value
E1BL_ADET1 (P04885) Early E1B 44 kDa protein 34 0.28
SUFI_HAEIN (P44847) Protein sufI homolog precursor 32 1.1
GATA_MYCGE (P47345) Glutamyl-tRNA(Gln) amidotransferase subunit ... 30 4.0
Y009_BORBU (O51042) Hypothetical protein BB0009 30 5.2
UPTG_PEA (O04300) Alpha-1,4-glucan-protein synthase [UDP-forming... 30 5.2
TWST_BRABE (O96642) Twist related protein (BBtwist) 30 6.8
SYFB_METJA (Q58508) Phenylalanyl-tRNA synthetase beta chain (EC ... 30 6.8
UPT2_SOLTU (Q8RU27) Alpha-1,4-glucan-protein synthase [UDP-formi... 29 8.9
PTH_BUCBP (P59490) Peptidyl-tRNA hydrolase (EC 3.1.1.29) (PTH) 29 8.9
>E1BL_ADET1 (P04885) Early E1B 44 kDa protein
Length = 391
Score = 34.3 bits (77), Expect = 0.28
Identities = 24/81 (29%), Positives = 37/81 (45%), Gaps = 10/81 (12%)
Query: 82 YSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVVQRR 141
Y+ +V+P T + F C D + + + ++ + NP L+Y + HNV+ L V RR
Sbjct: 238 YNSIVNPFTLQDSAEFSMVTCADAKVQLLHTIHIHSNPKLVYPQFMHNVLLRAKLFVGRR 297
Query: 142 ----------LKYLRLNLRLG 152
LKY L L G
Sbjct: 298 RGGFHPHFCSLKYSLLTLAKG 318
>SUFI_HAEIN (P44847) Protein sufI homolog precursor
Length = 311
Score = 32.3 bits (72), Expect = 1.1
Identities = 26/76 (34%), Positives = 36/76 (47%), Gaps = 10/76 (13%)
Query: 104 DVQFRN-GNPLYGYDNPHLLYKRSCHNVVKWVNLVVQR---RLKYLRLNLRLGYDLHLDV 159
D++F N G L+ + PH + R N ++ L V R RL+ L +L YDL LD
Sbjct: 188 DMEFNNDGLQLFKQNQPHFVGNRLLVNGIEAPYLDVARGWIRLRLLNASLARAYDLRLDN 247
Query: 160 DD------YDNSYLPK 169
D D +LPK
Sbjct: 248 DQEMLLIAQDLGFLPK 263
>GATA_MYCGE (P47345) Glutamyl-tRNA(Gln) amidotransferase subunit A
(EC 6.3.5.-) (Glu-ADT subunit A)
Length = 477
Score = 30.4 bits (67), Expect = 4.0
Identities = 14/39 (35%), Positives = 22/39 (55%)
Query: 47 LSKRWNNLWLSLPNIDFNNIKIDSIESNSKFIDSVYSVL 85
L K+WNNL +L ++ I +D K IDS+Y ++
Sbjct: 266 LQKKWNNLKKTLEQKNYQLIPLDFDVELLKVIDSIYKII 304
>Y009_BORBU (O51042) Hypothetical protein BB0009
Length = 334
Score = 30.0 bits (66), Expect = 5.2
Identities = 21/80 (26%), Positives = 38/80 (47%), Gaps = 5/80 (6%)
Query: 14 SLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKID-SIE 72
+L V+D + +P + L + V L+K+++N + PN+ FNN+K I+
Sbjct: 193 TLFVSDYLEVVPPNPL----VMFEKSHVVVNIYLNKKYSNTTIKSPNLIFNNLKNGLEIK 248
Query: 73 SNSKFIDSVYSVLVSPDTAI 92
K I+S + V T +
Sbjct: 249 DKEKIINSENKMFVKIKTRL 268
>UPTG_PEA (O04300) Alpha-1,4-glucan-protein synthase [UDP-forming]
(EC 2.4.1.112) (UDP-glucose:protein transglucosylase)
(UPTG) (Reversibly glycosylated polypeptide)
Length = 364
Score = 30.0 bits (66), Expect = 5.2
Identities = 14/48 (29%), Positives = 28/48 (58%), Gaps = 4/48 (8%)
Query: 37 STKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFIDSVYSV 84
S +E V T++ + LWL++P+ D + E N++F+D+V ++
Sbjct: 156 SLREGVPTAVS----HGLWLNIPDYDAPTQLVKPHERNTRFVDAVLTI 199
>TWST_BRABE (O96642) Twist related protein (BBtwist)
Length = 196
Score = 29.6 bits (65), Expect = 6.8
Identities = 16/57 (28%), Positives = 34/57 (59%), Gaps = 1/57 (1%)
Query: 35 LVSTKEAVATSILSKRWNNLWLSLPNIDFNNI-KIDSIESNSKFIDSVYSVLVSPDT 90
L + +E T L++ +++L +P + + + KI +++ +++ID +Y VL S DT
Sbjct: 99 LANVRERQRTQSLNEAFSSLRKIIPTLPSDKLSKIQTLKLAARYIDFLYQVLRSDDT 155
>SYFB_METJA (Q58508) Phenylalanyl-tRNA synthetase beta chain (EC
6.1.1.20) (Phenylalanine--tRNA ligase beta chain)
(PheRS)
Length = 548
Score = 29.6 bits (65), Expect = 6.8
Identities = 15/45 (33%), Positives = 27/45 (59%), Gaps = 3/45 (6%)
Query: 63 FNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQF 107
+ N+K D +E+ S++I+ V ++P T I +++ R LD QF
Sbjct: 272 YPNLKEDVLETTSEYINKVLGANLTPGTII---NYLRRCRLDAQF 313
>UPT2_SOLTU (Q8RU27) Alpha-1,4-glucan-protein synthase [UDP-forming]
2 (EC 2.4.1.112) (UDP-glucose:protein transglucosylase
2) (UPTG 2)
Length = 366
Score = 29.3 bits (64), Expect = 8.9
Identities = 13/48 (27%), Positives = 28/48 (58%), Gaps = 4/48 (8%)
Query: 37 STKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFIDSVYSV 84
S +E AT++ + LWL++P+ D + E N++++D+V ++
Sbjct: 157 SMREGAATAVS----HGLWLNIPDYDAPTQLVKPRERNTRYVDAVMTI 200
>PTH_BUCBP (P59490) Peptidyl-tRNA hydrolase (EC 3.1.1.29) (PTH)
Length = 207
Score = 29.3 bits (64), Expect = 8.9
Identities = 16/49 (32%), Positives = 25/49 (50%), Gaps = 2/49 (4%)
Query: 113 LYGYDNPHLLYKRSCHNVVKWV--NLVVQRRLKYLRLNLRLGYDLHLDV 159
+ G NP Y + HNV W+ +LV Q+ K + N LGY +++
Sbjct: 7 IVGLANPIKKYNDTRHNVGSWLVNSLVTQQNKKLKKNNKFLGYSTEINI 55
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,946,769
Number of Sequences: 164201
Number of extensions: 1097816
Number of successful extensions: 3246
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 3244
Number of HSP's gapped (non-prelim): 10
length of query: 242
length of database: 59,974,054
effective HSP length: 107
effective length of query: 135
effective length of database: 42,404,547
effective search space: 5724613845
effective search space used: 5724613845
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135799.3


