

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
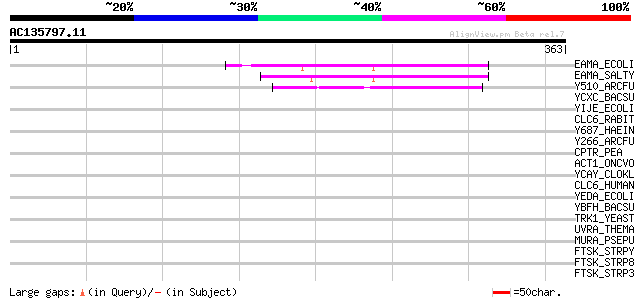
Query= AC135797.11 + phase: 0 /pseudo
(363 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
EAMA_ECOLI (P31125) Probable amino acid metabolite efflux pump 52 2e-06
EAMA_SALTY (Q56072) Probable amino acid metabolite efflux pump 48 3e-05
Y510_ARCFU (O29740) Hypothetical transport protein AF0510 45 4e-04
YCXC_BACSU (Q08794) Hypothetical transport protein ycxC 37 0.078
YIJE_ECOLI (P32667) Hypothetical transport protein yijE 35 0.30
CLC6_RABIT (Q9TT16) Chloride channel protein 6 (ClC-6) 35 0.30
Y687_HAEIN (P71356) Hypothetical transport protein HI0687 33 0.87
Y266_ARCFU (O29973) Hypothetical transport protein AF0266 32 3.3
CPTR_PEA (P21727) Triose phosphate/phosphate translocator, chlor... 32 3.3
ACT1_ONCVO (P30162) Actin 1 32 3.3
YCAY_CLOKL (P38943) Hypothetical transport protein in cat1 5'reg... 31 4.3
CLC6_HUMAN (P51797) Chloride channel protein 6 (ClC-6) 31 4.3
YEDA_ECOLI (P09185) Hypothetical transport protein yedA 30 7.3
YBFH_BACSU (O31448) Hypothetical transport protein ybhF 30 7.3
TRK1_YEAST (P12685) Potassium transport protein, high-affinity 30 7.3
UVRA_THEMA (Q9WYV0) UvrABC system protein A (UvrA protein) (Exci... 30 9.6
MURA_PSEPU (Q9Z3Z6) UDP-N-acetylglucosamine 1-carboxyvinyltransf... 30 9.6
FTSK_STRPY (Q9A155) DNA translocase ftsK 30 9.6
FTSK_STRP8 (Q8P276) DNA translocase ftsK 30 9.6
FTSK_STRP3 (Q8K8E8) DNA translocase ftsK 30 9.6
>EAMA_ECOLI (P31125) Probable amino acid metabolite efflux pump
Length = 299
Score = 52.0 bits (123), Expect = 2e-06
Identities = 46/177 (25%), Positives = 83/177 (45%), Gaps = 10/177 (5%)
Query: 142 IKGPILNLFGIHESSAQIQHNGGVNLQHAVKGSIMITIGC-FSCACFTIL--QAVTLETY 198
+ G L +FG+ + +N QH M+T+ FS AC I + ++ T
Sbjct: 117 LAGIALAIFGV-----LVLIEDSLNGQHVAMLGFMLTLAAAFSWACGNIFNKKIMSHSTR 171
Query: 199 PAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWD--TKLLAAVYSGIVCSGMAYYI 256
PA +SL W L+ + + +LI++ + SL T +L+ +Y V + + Y I
Sbjct: 172 PAVMSLVIWSALIPIIPFFVASLILDGSATMIHSLVTIDMTTILSLMYLAFVATIVGYGI 231
Query: 257 QGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVIILGLYLVVWG 313
G ++ V + L V+ + +L E++ + +GAV+I+ GLY+ V+G
Sbjct: 232 WGTLLGRYETWRVAPLSLLVPVVGLASAALLLDERLTGLQFLGAVLIMTGLYINVFG 288
>EAMA_SALTY (Q56072) Probable amino acid metabolite efflux pump
Length = 299
Score = 48.1 bits (113), Expect = 3e-05
Identities = 41/154 (26%), Positives = 72/154 (46%), Gaps = 5/154 (3%)
Query: 165 VNLQHAVKGSIMITIGC-FSCACFTILQAVTLE--TYPAELSLTAWICLLGTVEGGIVAL 221
+N QH M+T+ FS AC I ++ PA +SL W L+ + + +L
Sbjct: 135 LNGQHIAMSGFMLTLAAAFSWACGNIFNKKIMQHSPRPAVMSLVVWSALIPILPFLLSSL 194
Query: 222 IMERGEPSVWSLSWD--TKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVI 279
++E + SL T +L+ +Y V + + Y I G ++ V + L V+
Sbjct: 195 LLEGADHITQSLITIDMTTILSLLYLAFVATILGYGIWGALLGRYETWRVAPLSLLVPVV 254
Query: 280 VAIMSPFILAEKIYLGRVIGAVVIILGLYLVVWG 313
+ +L E + ++ GAV+I+ GLY+ V+G
Sbjct: 255 GLASAAVLLGETLTGMQLAGAVLIMAGLYINVFG 288
>Y510_ARCFU (O29740) Hypothetical transport protein AF0510
Length = 289
Score = 44.7 bits (104), Expect = 4e-04
Identities = 37/137 (27%), Positives = 62/137 (45%), Gaps = 4/137 (2%)
Query: 173 GSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWS 232
G +++ + F A +T+L + L Y E +LTA+ LGT+ AL+ P ++
Sbjct: 150 GDVLMIVDGFLWAVYTVLGSKMLLKYDHE-TLTAYAFALGTIFLIPFALMSGFANPVTFN 208
Query: 233 LSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKI 292
+ A +Y I+CS AY + + V + L + AI + + L EK
Sbjct: 209 ---PETVAALLYLSILCSVFAYVVWYYALTNADSTSVAVYVYLVPLFTAIFAFYALNEKP 265
Query: 293 YLGRVIGAVVIILGLYL 309
IG ++ I G+YL
Sbjct: 266 DFFTAIGGIITIAGVYL 282
>YCXC_BACSU (Q08794) Hypothetical transport protein ycxC
Length = 312
Score = 37.0 bits (84), Expect = 0.078
Identities = 39/184 (21%), Positives = 78/184 (42%), Gaps = 26/184 (14%)
Query: 129 TIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHA-VKGSIMITIGCFSCACF 187
T+ +VAG M + ++KG V+++ A +KGS++I + S A +
Sbjct: 130 TVLSVAGVMFIFVMKG--------------------VDVESASLKGSLLILLSALSSAMY 169
Query: 188 TILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTK----LLAAV 243
+ + LT + +G V +AL+ +V + + +LA V
Sbjct: 170 NTAARKMTQRFKLT-ELTYIMSAIGFVVFNAIALVRHGAAGTVGTYFLPFREPGFVLAIV 228
Query: 244 YSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVI 303
Y G++ S + ++ + ++ FN + ++ I IL E + + GAV I
Sbjct: 229 YLGVLSSLVTSFLSNYTLSRIEAFKMSAFNHVSTIVTMIAGFVILNESLAWYHLAGAVCI 288
Query: 304 ILGL 307
++G+
Sbjct: 289 MIGV 292
>YIJE_ECOLI (P32667) Hypothetical transport protein yijE
Length = 301
Score = 35.0 bits (79), Expect = 0.30
Identities = 30/111 (27%), Positives = 55/111 (49%), Gaps = 8/111 (7%)
Query: 202 LSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWD-TKLLAAVYSGIVCSGMAYYIQGVV 260
LSLT+W L + +VAL++ + E + W T A YS I+ + +A+ + V
Sbjct: 184 LSLTSWQMLYAALVMSVVALLVPQRE-----IDWQPTVFWALAYSAILATALAWSLWLFV 238
Query: 261 MRYRGPVFVTTFNPLCMVIVAIM-SPFILAEKIYLGRVIGAVVIILGLYLV 310
++ P + + + L + + ++ S ++L E G V+I+L L LV
Sbjct: 239 LK-NLPASIASLSTLAVPVCGVLFSWWLLGENPGAVEGSGIVLIVLALALV 288
>CLC6_RABIT (Q9TT16) Chloride channel protein 6 (ClC-6)
Length = 869
Score = 35.0 bits (79), Expect = 0.30
Identities = 29/100 (29%), Positives = 49/100 (49%), Gaps = 6/100 (6%)
Query: 108 LDSQMEKIKMRSVHSQAKIVGTIATVAGAMVMTL---IKGPILNLFGIHESSAQIQHNGG 164
L+ ++ K +MR+VH + K+V + ++ ++V TL + +L SS+QI NG
Sbjct: 354 LNKRLAKYRMRNVHPKPKLVRVLESLLVSLVTTLVVFVASMVLGECRQMSSSSQIS-NGS 412
Query: 165 VNLQHAVKGSIMITIGCFSCACFTILQAVTLETYPAELSL 204
+ LQ V + +I F C T TL P E ++
Sbjct: 413 LKLQ--VTSDVNSSIKAFFCPNDTYNDMATLFFNPQESAI 450
>Y687_HAEIN (P71356) Hypothetical transport protein HI0687
Length = 304
Score = 33.5 bits (75), Expect = 0.87
Identities = 22/77 (28%), Positives = 42/77 (53%), Gaps = 5/77 (6%)
Query: 246 GIVCSGMAYYIQGVVMRYR--GPVFVTTFNPLCMVI---VAIMSPFILAEKIYLGRVIGA 300
G+VC+G + G++M + +T FN L ++I AI+ L E+I + + I
Sbjct: 227 GLVCAGFYGMLTGMLMAFYIVQKQGITVFNILQLLIPLSTAIIGYLTLDERINIYQGISG 286
Query: 301 VVIILGLYLVVWGKSKD 317
+++I+G L + K+K+
Sbjct: 287 IIVIIGCVLALKRKNKE 303
>Y266_ARCFU (O29973) Hypothetical transport protein AF0266
Length = 276
Score = 31.6 bits (70), Expect = 3.3
Identities = 41/201 (20%), Positives = 81/201 (39%), Gaps = 32/201 (15%)
Query: 112 MEKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAV 171
+ I +R + +K+VG I G +V++ + N++GI
Sbjct: 104 LSAIFLRERITYSKVVGIIIAFLGVVVIS--EPSYANIYGI------------------- 142
Query: 172 KGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVW 231
++ + + A +T L Y ++LT+ +LG++ + S+
Sbjct: 143 ---ALVMVSTVAAAIYTTFGKSLLSKYNP-ITLTSNAMVLGSIP------LYPFLPDSIR 192
Query: 232 SLSWDTKLLAA-VYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAE 290
SL D L+ + V+ GI + Y + + F V+ + +LAE
Sbjct: 193 SLGGDLNLIGSIVFLGIFSTFFGYLGWYYFLEKEEASRASVFLLAIPVVSLLAGNILLAE 252
Query: 291 KIYLGRVIGAVVIILGLYLVV 311
+ L V G+ +++LG+Y+VV
Sbjct: 253 PLTLRTVAGSGLVLLGIYIVV 273
>CPTR_PEA (P21727) Triose phosphate/phosphate translocator,
chloroplast precursor (CTPT) (p36) (E30)
Length = 402
Score = 31.6 bits (70), Expect = 3.3
Identities = 20/65 (30%), Positives = 32/65 (48%), Gaps = 2/65 (3%)
Query: 1 MAFCNQKSFQNWKPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPF 60
MA + SF NW FI+ ++ + I SK A+ M + Y +A +V +P
Sbjct: 231 MASLTELSF-NWLGFISAMISNISFTYRSIYSKKAMT-DMDSTNIYAYISIIALIVCIPP 288
Query: 61 AVILE 65
A+I+E
Sbjct: 289 ALIIE 293
>ACT1_ONCVO (P30162) Actin 1
Length = 376
Score = 31.6 bits (70), Expect = 3.3
Identities = 25/90 (27%), Positives = 38/90 (41%), Gaps = 6/90 (6%)
Query: 236 DTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLG 295
D + +Y+ IV SG G+ R + V T P M I I P E+ Y
Sbjct: 287 DIDIRKDLYANIVLSGGTTMYPGIADRMQKEV--TALAPSTMKIKIIAPP----ERKYSV 340
Query: 296 RVIGAVVIILGLYLVVWGKSKDYDRPSPII 325
+ G+++ L + VW ++YD P I
Sbjct: 341 WIGGSILASLSTFQQVWISKQEYDESGPSI 370
>YCAY_CLOKL (P38943) Hypothetical transport protein in cat1 5'region
(ORFY)
Length = 311
Score = 31.2 bits (69), Expect = 4.3
Identities = 57/300 (19%), Positives = 112/300 (37%), Gaps = 22/300 (7%)
Query: 13 KPFIAVVLLQFGYAGMDILSKSALNKG-MSCYVLVVYRHAVAFVVIVPFAVILENGSQCA 71
K +I ++L Y+ +I K KG M + +++ + ++++P AV
Sbjct: 3 KGYIFILLTAIFYSTQEISGKMLAQKGAMDPFQVMMIVFLIGAIILLPMAV--------- 53
Query: 72 RACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKMRSVHSQAKIVGTIA 131
+ K GN++ Y C + S K +V V TI
Sbjct: 54 KDIKVKKLKLTGNDL--GYLALCGILAVSISMSMLQFAVTYTKASTAAVLFCTNAVFTIP 111
Query: 132 TVAGAMVMTLIKGP-----ILNLFGIHESSAQIQHNGGVNLQHAVKGSIMITIGCFSCAC 186
A ++ IKG I++L G+ + G+ + G + +
Sbjct: 112 -FAYFILKEKIKGITIVSIIVSLIGVVIIFNPAKVMEGIGGSRDLIGICFALVAAVVWSL 170
Query: 187 FTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTKLLAAVYSG 246
+T++ +E Y + G + ++ L++ G P ++ + +L +Y G
Sbjct: 171 YTVISKKRIEIYGGYV-FNCISFFFGVI--ALLILLVVTGRPIFSGITLNN-ILVLLYMG 226
Query: 247 IVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVIILG 306
I + Y ++ V +T + + +++ IL E I + VIG V II+G
Sbjct: 227 IFIKAVGYICYLGAIKETSAVTASTVFLIKPALATVLAILILGESIEVNVVIGIVFIIIG 286
>CLC6_HUMAN (P51797) Chloride channel protein 6 (ClC-6)
Length = 869
Score = 31.2 bits (69), Expect = 4.3
Identities = 22/97 (22%), Positives = 43/97 (43%)
Query: 108 LDSQMEKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNL 167
L+ ++ K +MR+VH + K+V + ++ ++V T++ + G + G +
Sbjct: 354 LNKRLAKYRMRNVHPKPKLVRVLESLLVSLVTTVVVFVASMVLGECRQMSSSSQIGNDSF 413
Query: 168 QHAVKGSIMITIGCFSCACFTILQAVTLETYPAELSL 204
Q V + +I F C T TL P E ++
Sbjct: 414 QLQVTEDVNSSIKTFFCPNDTYNDMATLFFNPQESAI 450
>YEDA_ECOLI (P09185) Hypothetical transport protein yedA
Length = 306
Score = 30.4 bits (67), Expect = 7.3
Identities = 43/176 (24%), Positives = 77/176 (43%), Gaps = 17/176 (9%)
Query: 159 IQHNGGVNLQHAVKGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVE--- 215
I N G NL G+I+I IG S A ++ Y + ++L + + G +E
Sbjct: 137 IMLNSGGNLSGNPWGAILILIGSISWAFGSV--------YGSRITLPVGM-MAGAIEMLA 187
Query: 216 GGIVALI--MERGEPSVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFN 273
G+V +I M GE + +L + LA Y + S +A ++R P T++
Sbjct: 188 AGVVLMIASMIAGE-KLTALPSLSGFLAVGYLALFGSIIAINAYMYLIRNVSPALATSYA 246
Query: 274 PLCMVIVAIMSPFILAEKIYLGRVIGAVVIILGLYLVVWGKSKDYDRP--SPIIKD 327
+ V+ ++ + E + + VI+ + LV GK +P +P+I+D
Sbjct: 247 YVNPVVAVLLGTGLGGETLSKIEWLALGVIVFAVVLVTLGKYLFPAKPVVAPVIQD 302
>YBFH_BACSU (O31448) Hypothetical transport protein ybhF
Length = 306
Score = 30.4 bits (67), Expect = 7.3
Identities = 24/97 (24%), Positives = 50/97 (50%), Gaps = 3/97 (3%)
Query: 217 GIVALIMERGEPSVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLC 276
G +ALI+ R + +W +LL A +G+ + + ++ + + Y V +
Sbjct: 47 GFIALILVRPNMIPFR-NWRQELLFAG-AGLFGVTLYFLLENIALTYTYASNVGMIVSII 104
Query: 277 MVIVAIMSPFIL-AEKIYLGRVIGAVVIILGLYLVVW 312
+I A+++ F+L EK+ L +IG + ++GL L+ +
Sbjct: 105 PMITAVLAHFLLEGEKLRLTFLIGFISALIGLLLITF 141
>TRK1_YEAST (P12685) Potassium transport protein, high-affinity
Length = 1235
Score = 30.4 bits (67), Expect = 7.3
Identities = 21/75 (28%), Positives = 37/75 (49%), Gaps = 12/75 (16%)
Query: 7 KSFQNWKPFIAVVLLQFGYAGMDILSKSALNK------GMSCYVLVVYR---HAVAFVVI 57
K++ +W+P I + G+ K L + C +LVVY H VAFV++
Sbjct: 738 KTYLSWQPTIG---RNSNFLGLTRAQKDELGGVEYRAIKLLCTILVVYYVGWHIVAFVML 794
Query: 58 VPFAVILENGSQCAR 72
VP+ ++ ++ S+ R
Sbjct: 795 VPWIILKKHYSEVVR 809
>UVRA_THEMA (Q9WYV0) UvrABC system protein A (UvrA protein)
(Excinuclease ABC subunit A)
Length = 916
Score = 30.0 bits (66), Expect = 9.6
Identities = 16/53 (30%), Positives = 27/53 (50%)
Query: 195 LETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTKLLAAVYSGI 247
L+ E+ L ++C+ G G +L+ME P++ +L TKL A + I
Sbjct: 600 LKNIDVEIPLGVFVCVTGVSGSGKSSLVMETLYPALMNLLHKTKLPAGEFDSI 652
>MURA_PSEPU (Q9Z3Z6) UDP-N-acetylglucosamine
1-carboxyvinyltransferase (EC 2.5.1.7) (Enoylpyruvate
transferase) (UDP-N-acetylglucosamine enolpyruvyl
transferase) (EPT)
Length = 419
Score = 30.0 bits (66), Expect = 9.6
Identities = 17/70 (24%), Positives = 32/70 (45%)
Query: 127 VGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVKGSIMITIGCFSCAC 186
+ IA GA++ T+ + ++++ +H AQIQ G + VK + G +
Sbjct: 316 LNAIAEGTGAVIETIFENRFMHVYEMHRMGAQIQVEGNTAIVTGVKALKVPGNGHLRASA 375
Query: 187 FTILQAVTLE 196
+L A+ E
Sbjct: 376 SLVLSALVAE 385
>FTSK_STRPY (Q9A155) DNA translocase ftsK
Length = 801
Score = 30.0 bits (66), Expect = 9.6
Identities = 19/54 (35%), Positives = 28/54 (51%), Gaps = 11/54 (20%)
Query: 269 VTTFNPLCMVIVAIMSPFILAEKIYL----------GRVIGAVVIILGLYLVVW 312
VTT+N + ++ ++ PF+ A IYL G + G V+ LGL LV W
Sbjct: 52 VTTYNMIRFLVGSLAYPFMFAWLIYLFCFKWLRQKDGMIAGVVIAFLGL-LVEW 104
>FTSK_STRP8 (Q8P276) DNA translocase ftsK
Length = 801
Score = 30.0 bits (66), Expect = 9.6
Identities = 19/54 (35%), Positives = 28/54 (51%), Gaps = 11/54 (20%)
Query: 269 VTTFNPLCMVIVAIMSPFILAEKIYL----------GRVIGAVVIILGLYLVVW 312
VTT+N + ++ ++ PF+ A IYL G + G V+ LGL LV W
Sbjct: 52 VTTYNMIRFLVGSLAYPFMFAWLIYLFCFKWLRQKDGMIAGVVIAFLGL-LVEW 104
>FTSK_STRP3 (Q8K8E8) DNA translocase ftsK
Length = 801
Score = 30.0 bits (66), Expect = 9.6
Identities = 19/54 (35%), Positives = 28/54 (51%), Gaps = 11/54 (20%)
Query: 269 VTTFNPLCMVIVAIMSPFILAEKIYL----------GRVIGAVVIILGLYLVVW 312
VTT+N + ++ ++ PF+ A IYL G + G V+ LGL LV W
Sbjct: 52 VTTYNMIRFLVGSLAYPFMFAWLIYLFCFKWLRQKDGMIAGVVIAFLGL-LVEW 104
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,215,436
Number of Sequences: 164201
Number of extensions: 1600248
Number of successful extensions: 4670
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 4664
Number of HSP's gapped (non-prelim): 27
length of query: 363
length of database: 59,974,054
effective HSP length: 111
effective length of query: 252
effective length of database: 41,747,743
effective search space: 10520431236
effective search space used: 10520431236
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC135797.11


