

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135796.7 - phase: 0
(277 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
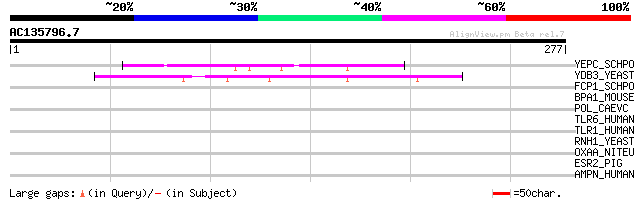
Score E
Sequences producing significant alignments: (bits) Value
YEPC_SCHPO (O13942) Hypothetical protein C23H3.12c in chromosome I 54 5e-07
YDB3_YEAST (P48569) Hypothetical 37.0 kDa protein in RPL41A-INH1... 49 1e-05
FCP1_SCHPO (Q9P376) RNA polymerase II subunit A C-terminal domai... 31 3.8
BPA1_MOUSE (Q91ZU6) Bullous pemphigoid antigen 1, isoforms 1/2/3... 31 3.8
POL_CAEVC (P33459) Pol polyprotein [Contains: Protease (Retropep... 30 4.9
TLR6_HUMAN (Q9Y2C9) Toll-like receptor 6 precursor 30 8.4
TLR1_HUMAN (Q15399) Toll-like receptor 1 precursor (Toll/interle... 30 8.4
RNH1_YEAST (Q04740) Ribonuclease H (EC 3.1.26.4) (RNase H) 30 8.4
OXAA_NITEU (P59810) Inner membrane protein oxaA 30 8.4
ESR2_PIG (Q9XSW2) Estrogen receptor beta (ER-beta) 30 8.4
AMPN_HUMAN (P15144) Aminopeptidase N (EC 3.4.11.2) (hAPN) (Alany... 30 8.4
>YEPC_SCHPO (O13942) Hypothetical protein C23H3.12c in chromosome I
Length = 226
Score = 53.5 bits (127), Expect = 5e-07
Identities = 41/151 (27%), Positives = 80/151 (52%), Gaps = 13/151 (8%)
Query: 57 AKAELFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSIS--KDV 114
AK D + N++ W + +A K+K+ LG +L E FL++I+ K +
Sbjct: 24 AKKVTIHDKIINRIYKYWDSW-SASKSYTKQKVVSLGNRILHATPYEENFLRAIAPVKKL 82
Query: 115 TGVE------VVYPSSMNARLVRRRL-RHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPL 167
E + +P ++ + + L R ++ T H + G++ +PL+ F ++PL
Sbjct: 83 NDTELHQTLYIEHPPNLESSTILAELNRSKQLQKT--HTNYLIGNIIGLPLTIPFILIPL 140
Query: 168 -PNVPFFWILFRSYSHWRALQGSEKLFQLVS 197
PN+P F++ +R+Y ++RA+QGS +L +++S
Sbjct: 141 IPNIPGFYLCYRAYCNFRAIQGSIQLARVMS 171
>YDB3_YEAST (P48569) Hypothetical 37.0 kDa protein in RPL41A-INH1
intergenic region
Length = 320
Score = 49.3 bits (116), Expect = 1e-05
Identities = 47/203 (23%), Positives = 87/203 (42%), Gaps = 25/203 (12%)
Query: 43 QLWKSINTGDKPFNAKAELFTDYVANKMNNGWVTLENAPDGSF-KKKIHGLGLWLLSRVK 101
+LW+ + K +N K + N +L P S+ K+I G + K
Sbjct: 84 KLWRKLKKSPKSYNKKIVSMVQSLLNSTPWSENSLLTIPSESYILKRIKG------EKDK 137
Query: 102 PSEIFL-------KSISKDVTGVEVVYPSSMNAR-LVRRRLRHIAMRGTIIHRKFFYGSV 153
EI L K+ D + V YP +++ R+++ + G I H+K+ +
Sbjct: 138 TQEIRLTLKDYTVKAEQVDTQPLHVYYPPGISSPDECLRQMKKLYQEGLIYHKKWTLYCL 197
Query: 154 SLIPLSSAFSILPL-PNVPFFWILFRSYSHWRALQGSEKLFQLVSDGSQS---------S 203
+PL+ ++PL PNVP F++ +R+Y + +A G++ L L+ Q+ +
Sbjct: 198 LGLPLTIPLILIPLIPNVPGFYLSYRAYVNIKAYLGAKHLKSLLESSKQNLEFRELLGYT 257
Query: 204 NTYSGKKETEHEDSENESLGLDE 226
Y + + ++ ES G E
Sbjct: 258 EVYKRGTSSRTQGNQKESKGAPE 280
>FCP1_SCHPO (Q9P376) RNA polymerase II subunit A C-terminal domain
phosphatase (EC 3.1.3.16) (CTD phosphatase fcp1)
Length = 723
Score = 30.8 bits (68), Expect = 3.8
Identities = 22/83 (26%), Positives = 37/83 (44%), Gaps = 4/83 (4%)
Query: 176 LFRSYSHWRALQGSEKLFQL---VSDGSQSSNTYSGKKETEHEDSENESLGLDEPHWVLT 232
L S S W+ L S+ L + D + S ++YS + E SE LDE W
Sbjct: 562 LTESLSQWKRLPESDYLLYPSYDLPDRNLSEHSYSSSSDDEQRISELNDRELDEIDW-QA 620
Query: 233 PSKELENIVRQEDGNDGLSRGTI 255
+++EN ++ ++ G+I
Sbjct: 621 ADQDVENALKDLSDDNDFDTGSI 643
>BPA1_MOUSE (Q91ZU6) Bullous pemphigoid antigen 1, isoforms 1/2/3/4
(BPA) (Hemidesmosomal plaque protein) (Dystonia
musculorum protein) (Dystonin)
Length = 7389
Score = 30.8 bits (68), Expect = 3.8
Identities = 16/39 (41%), Positives = 21/39 (53%)
Query: 182 HWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENE 220
H AL G EKL +L S+ + S + SG KE + S E
Sbjct: 3268 HIAALPGGEKLGELCSEPPEHSESTSGSKERSSDGSSKE 3306
>POL_CAEVC (P33459) Pol polyprotein [Contains: Protease
(Retropepsin) (EC 3.4.23.-); Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 1109
Score = 30.4 bits (67), Expect = 4.9
Identities = 20/82 (24%), Positives = 36/82 (43%), Gaps = 10/82 (12%)
Query: 170 VPFFWILFRSYSHWRALQGSEKLFQ---LVSDGSQSSNTYS-------GKKETEHEDSEN 219
+P FW +R ++ WR E++ + +DG + + S G+K +HE+ N
Sbjct: 553 MPKFWSCYRGHTRWRKRNIIEEVVEGPTYYTDGGKKNKVGSLGFIVSTGEKFRKHEEGTN 612
Query: 220 ESLGLDEPHWVLTPSKELENIV 241
+ L L L + N+V
Sbjct: 613 QQLELRAIEEALKQGPQTMNLV 634
>TLR6_HUMAN (Q9Y2C9) Toll-like receptor 6 precursor
Length = 796
Score = 29.6 bits (65), Expect = 8.4
Identities = 22/59 (37%), Positives = 28/59 (47%), Gaps = 10/59 (16%)
Query: 21 IDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAEL-----FTDYVANKMNNGW 74
IDH S +ADF QS Q +SI GD PF EL D V++++ GW
Sbjct: 502 IDHN-SVSHPSADFFQSC----QKMRSIKAGDNPFQCTCELREFVKNIDQVSSEVLEGW 555
>TLR1_HUMAN (Q15399) Toll-like receptor 1 precursor
(Toll/interleukin-1 receptor-like) (TIL)
Length = 786
Score = 29.6 bits (65), Expect = 8.4
Identities = 22/59 (37%), Positives = 28/59 (47%), Gaps = 10/59 (16%)
Query: 21 IDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAEL-----FTDYVANKMNNGW 74
IDH S +ADF QS Q +SI GD PF EL D V++++ GW
Sbjct: 497 IDHN-SVSHPSADFFQSC----QKMRSIKAGDNPFQCTCELGEFVKNIDQVSSEVLEGW 550
>RNH1_YEAST (Q04740) Ribonuclease H (EC 3.1.26.4) (RNase H)
Length = 348
Score = 29.6 bits (65), Expect = 8.4
Identities = 14/65 (21%), Positives = 33/65 (50%), Gaps = 7/65 (10%)
Query: 28 SSSTADFSQSPSTLKQLWKSINTGDKPFNAKAELFTDYVANKMNNGWVTLENAPDGSFKK 87
+++ A+ LK++W+ + + N + + ++YV +N+ ++T +N K
Sbjct: 230 TNNRAEIEAVSEALKKIWEKLTNEKEKVNYQIKTDSEYVTKLLNDRYMTYDN-------K 282
Query: 88 KIHGL 92
K+ GL
Sbjct: 283 KLEGL 287
>OXAA_NITEU (P59810) Inner membrane protein oxaA
Length = 614
Score = 29.6 bits (65), Expect = 8.4
Identities = 21/74 (28%), Positives = 33/74 (44%), Gaps = 10/74 (13%)
Query: 54 PFNAKAELFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKD 113
PF+ + DY AN NNGW+ + + + L WL + P E F K S +
Sbjct: 259 PFSDLDKNKADYPANA-NNGWIAM---------LEHYFLTAWLPQQQTPREFFAKRQSDN 308
Query: 114 VTGVEVVYPSSMNA 127
+ V+ P+ + A
Sbjct: 309 LYSAGVIVPAGVIA 322
>ESR2_PIG (Q9XSW2) Estrogen receptor beta (ER-beta)
Length = 526
Score = 29.6 bits (65), Expect = 8.4
Identities = 12/26 (46%), Positives = 16/26 (61%)
Query: 15 WCFTRSIDHTLPASSSTADFSQSPST 40
WC TRS++HTLP + T S S+
Sbjct: 104 WCDTRSLEHTLPVNRETLKRKASGSS 129
>AMPN_HUMAN (P15144) Aminopeptidase N (EC 3.4.11.2) (hAPN) (Alanyl
aminopeptidase) (Microsomal aminopeptidase)
(Aminopeptidase M) (gp150) (Myeloid plasma membrane
glycoprotein CD13)
Length = 966
Score = 29.6 bits (65), Expect = 8.4
Identities = 30/119 (25%), Positives = 49/119 (40%), Gaps = 18/119 (15%)
Query: 95 WLLSRVKPSEIFLKS------ISKDVTGVEVVYPSSMNARLVRRRLRH------IAMRGT 142
WL+ +++F S ++ +VTG V N R ++ +L+ + R
Sbjct: 600 WLIDVRAQNDLFSTSGNEWVLLNLNVTGYYRVNYDEENWRKIQTQLQRDHSAIPVINRAQ 659
Query: 143 IIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILFRSYSHWRALQGSEKLFQLVSDGSQ 201
II+ F S +P++ A N F I R Y W A S F+L+ D S+
Sbjct: 660 IINDAFNLASAHKVPVTLAL------NNTLFLIEERQYMPWEAALSSLSYFKLMFDRSE 712
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,898,622
Number of Sequences: 164201
Number of extensions: 1376411
Number of successful extensions: 3833
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 3823
Number of HSP's gapped (non-prelim): 14
length of query: 277
length of database: 59,974,054
effective HSP length: 109
effective length of query: 168
effective length of database: 42,076,145
effective search space: 7068792360
effective search space used: 7068792360
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC135796.7


