

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135565.5 + phase: 0 /pseudo
(93 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
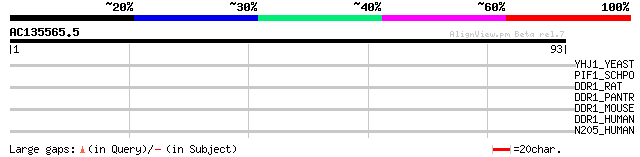
Score E
Sequences producing significant alignments: (bits) Value
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 32 0.37
PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1, m... 30 1.4
DDR1_RAT (Q63474) Epithelial discoidin domain receptor 1 precurs... 28 3.1
DDR1_PANTR (Q7YR43) Epithelial discoidin domain receptor 1 precu... 28 3.1
DDR1_MOUSE (Q03146) Epithelial discoidin domain receptor 1 precu... 28 3.1
DDR1_HUMAN (Q08345) Epithelial discoidin domain receptor 1 precu... 28 3.1
N205_HUMAN (Q92621) Nuclear pore complex protein Nup205 (Nucleop... 28 5.3
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 31.6 bits (70), Expect = 0.37
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Query: 29 PFGGKVFVLVGDFRQVLPVVRVATRQQIVSSAVNASKVWRRC 70
PFGG VL GDF Q+ PV + + V S++W+RC
Sbjct: 363 PFGGIQLVLTGDFFQLPPVAK--KDEHNVVKFCFESEMWKRC 402
>PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1,
mitochondrial precursor
Length = 805
Score = 29.6 bits (65), Expect = 1.4
Identities = 21/67 (31%), Positives = 29/67 (42%), Gaps = 8/67 (11%)
Query: 27 TKPFGGKVFVLVGDFRQVLPVVRVATRQQIVSSAVNASKVWRRCEVLRLTMNMGLSTANN 86
+KPFGG VL GDF Q+ PV + S+ W+ L +GL+
Sbjct: 441 SKPFGGIQLVLTGDFFQLPPVPENGKESKFCFE----SQTWKSA----LDFTIGLTHVFR 492
Query: 87 NADQEEV 93
D+E V
Sbjct: 493 QKDEEFV 499
>DDR1_RAT (Q63474) Epithelial discoidin domain receptor 1 precursor
(EC 2.7.1.112) (Tyrosine kinase DDR) (Discoidin receptor
tyrosine kinase) (Tyrosine-protein kinase CAK) (Cell
adhesion kinase) (Protein-tyrosine kinase PTK-3)
Length = 910
Score = 28.5 bits (62), Expect = 3.1
Identities = 21/66 (31%), Positives = 28/66 (41%), Gaps = 5/66 (7%)
Query: 7 FVALDKTLRNVMGSENKDNKTKPFGGKVFVLVGDF-----RQVLPVVRVATRQQIVSSAV 61
FV D RN + EN K FG + GD+ R VLP+ +A ++
Sbjct: 759 FVHRDLATRNCLVGENFTIKIADFGMSRNLYAGDYYRVQGRAVLPIRWMAWECILMGKFT 818
Query: 62 NASKVW 67
AS VW
Sbjct: 819 TASDVW 824
>DDR1_PANTR (Q7YR43) Epithelial discoidin domain receptor 1
precursor (EC 2.7.1.112) (Tyrosine kinase DDR)
(Discoidin receptor tyrosine kinase)
Length = 909
Score = 28.5 bits (62), Expect = 3.1
Identities = 21/66 (31%), Positives = 28/66 (41%), Gaps = 5/66 (7%)
Query: 7 FVALDKTLRNVMGSENKDNKTKPFGGKVFVLVGDF-----RQVLPVVRVATRQQIVSSAV 61
FV D RN + EN K FG + GD+ R VLP+ +A ++
Sbjct: 758 FVHRDLATRNCLVGENFTIKIADFGMSRNLYAGDYYRVQGRAVLPIRWMAWECILMGKFT 817
Query: 62 NASKVW 67
AS VW
Sbjct: 818 TASDVW 823
>DDR1_MOUSE (Q03146) Epithelial discoidin domain receptor 1
precursor (EC 2.7.1.112) (Tyrosine kinase DDR)
(Discoidin receptor tyrosine kinase) (Tyrosine-protein
kinase CAK) (Cell adhesion kinase) (Protein-tyrosine
kinase MPK-6)
Length = 911
Score = 28.5 bits (62), Expect = 3.1
Identities = 21/66 (31%), Positives = 28/66 (41%), Gaps = 5/66 (7%)
Query: 7 FVALDKTLRNVMGSENKDNKTKPFGGKVFVLVGDF-----RQVLPVVRVATRQQIVSSAV 61
FV D RN + EN K FG + GD+ R VLP+ +A ++
Sbjct: 760 FVHRDLATRNCLVGENFTIKIADFGMSRNLYAGDYYRVQGRAVLPIRWMAWECILMGKFT 819
Query: 62 NASKVW 67
AS VW
Sbjct: 820 TASDVW 825
>DDR1_HUMAN (Q08345) Epithelial discoidin domain receptor 1
precursor (EC 2.7.1.112) (Tyrosine kinase DDR)
(Discoidin receptor tyrosine kinase) (Tyrosine-protein
kinase CAK) (Cell adhesion kinase) (TRK E)
(Protein-tyrosine kinase RTK 6) (CD167a antigen) (
Length = 913
Score = 28.5 bits (62), Expect = 3.1
Identities = 21/66 (31%), Positives = 28/66 (41%), Gaps = 5/66 (7%)
Query: 7 FVALDKTLRNVMGSENKDNKTKPFGGKVFVLVGDF-----RQVLPVVRVATRQQIVSSAV 61
FV D RN + EN K FG + GD+ R VLP+ +A ++
Sbjct: 762 FVHRDLATRNCLVGENFTIKIADFGMSRNLYAGDYYRVQGRAVLPIRWMAWECILMGKFT 821
Query: 62 NASKVW 67
AS VW
Sbjct: 822 TASDVW 827
>N205_HUMAN (Q92621) Nuclear pore complex protein Nup205
(Nucleoporin Nup205) (205 kDa nucleoporin)
Length = 2012
Score = 27.7 bits (60), Expect = 5.3
Identities = 11/37 (29%), Positives = 22/37 (58%)
Query: 40 DFRQVLPVVRVATRQQIVSSAVNASKVWRRCEVLRLT 76
++ Q+L VR+ +++Q + V +++ RCE LT
Sbjct: 668 EYTQILQTVRIPSQRQAIGIEVELNEIESRCEEYPLT 704
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.133 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,742,103
Number of Sequences: 164201
Number of extensions: 296824
Number of successful extensions: 552
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 549
Number of HSP's gapped (non-prelim): 7
length of query: 93
length of database: 59,974,054
effective HSP length: 69
effective length of query: 24
effective length of database: 48,644,185
effective search space: 1167460440
effective search space used: 1167460440
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135565.5


