

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135464.5 - phase: 0 /pseudo
(1087 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
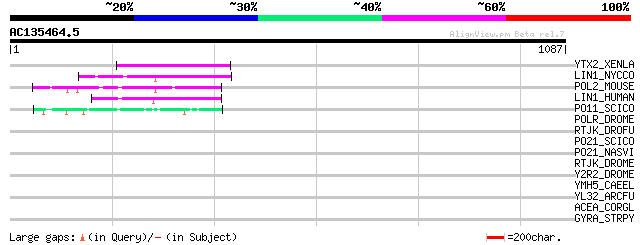
Score E
Sequences producing significant alignments: (bits) Value
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 97 2e-19
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 85 9e-16
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 82 6e-15
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 68 2e-10
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 47 3e-04
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 39 0.076
RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile elem... 39 0.099
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 39 0.099
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 37 0.22
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 37 0.29
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 34 2.4
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 33 3.2
YL32_ARCFU (O28148) Hypothetical protein AF2132 33 3.2
ACEA_CORGL (P42449) Isocitrate lyase (EC 4.1.3.1) (Isocitrase) (... 33 4.2
GYRA_STRPY (Q9L7Q5) DNA gyrase subunit A (EC 5.99.1.3) 33 5.4
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 97.1 bits (240), Expect = 2e-19
Identities = 64/225 (28%), Positives = 113/225 (49%), Gaps = 3/225 (1%)
Query: 210 QSRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLA-DSNLVEGVQPIRNAVLSHFRDHFA 268
+SR+ L D D S++F+++ + R I L A D +E + IR+ S +++ F+
Sbjct: 359 RSRMQLLCDMDRGSRFFYALEKKKGNRKQITCLFAEDGTPLEDPEAIRDRARSFYQNLFS 418
Query: 269 ARHTSRVGIDNL--PFKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIK 326
S + L +S L P + DE+ A+ KSPG DG+ F +
Sbjct: 419 PDPISPDACEELWDGLPVVSERRKERLETPITLDELSQALRLMPHNKSPGLDGLTIEFFQ 478
Query: 327 EFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKIL 386
FWD L D R +++ + G+L ++L+ K + I ++RP+SL+ + YKI+
Sbjct: 479 FFWDTLGPDFHRVLTEAFKKGELPLSCRRAVLSLLPKKGDLRLIKNWRPVSLLSTDYKIV 538
Query: 387 SKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVDEARK 431
+K ++ RLK VL ++I Q+ V R I D + + +++ AR+
Sbjct: 539 AKAISLRLKSVLAEVIHPDQSYTVPGRTIFDNVFLIRDLLHFARR 583
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 85.1 bits (209), Expect = 9e-16
Identities = 76/309 (24%), Positives = 138/309 (44%), Gaps = 22/309 (7%)
Query: 135 VLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGDVGDLSADEILELRGITH 194
VLR K ++ LK+ +N+ G + L+K ++ + EI ++R +
Sbjct: 293 VLRGKFIALQAFLKKTEREEVNNLMGHLKQLEK-----EEHSNPKPSRRKEITKIRAELN 347
Query: 195 DLHS---LSRVNSSILWQQSRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADSN--LV 249
++ + + ++N S W ++ + AN L+ ++R S+IS + + N +
Sbjct: 348 EIENKRIIQQINKSKSWFFEKINKIDKPLAN-------LTRKKRVKSLISSIRNGNDEIT 400
Query: 250 EGVQPIRNAVLSHFRDHFAARHTSRVGIDNLPFKC----LSHTDGSGLTKPFSADEVKSA 305
I+ + +++ ++ ++ + ID C LS + L +P S+ E+ S
Sbjct: 401 TDPSEIQKILNEYYKKLYSHKYENLKEIDQYLEACHLPRLSQKEVEMLNRPISSSEIAST 460
Query: 306 VWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKI- 364
+ + KSPGPDG F + F +EL ++ + + G L I LI K
Sbjct: 461 IQNLPKKKSPGPDGFTSEFYQTFKEELVPILLNLFQNIEKEGILPNTFYEANITLIPKPG 520
Query: 365 DSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANE 424
P R ++RPISL+ KIL+K+L NR++ + KII Q F+ Q I +
Sbjct: 521 KDPTRKENYRPISLMNIDAKILNKILTNRIQQHIKKIIHHDQVGFIPGSQGWFNIRKSIN 580
Query: 425 VVDEARKNK 433
V+ K K
Sbjct: 581 VIQHINKLK 589
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 82.4 bits (202), Expect = 6e-15
Identities = 95/398 (23%), Positives = 171/398 (42%), Gaps = 45/398 (11%)
Query: 45 FTWFKGDGNAMSRIDRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENWGPRPSR 104
+T+F S+ID + + ++ N V + LSDH L+L + +P+
Sbjct: 217 YTFFSAPHGTFSKIDHIIGHKTGLNRYKNIEIVPCI--LSDHHGLRLIFNNNINNGKPTF 274
Query: 105 MLKCWQDL-------PGYNQFVKDKLKSFQIEG------WGGF--VLREKLKFIKTALKE 149
K L G + +KD L+ + E W LR KL + + K+
Sbjct: 275 TWKLNNTLLNDTLVKEGIKKEIKDFLEFNENEATTYPNLWDTMKAFLRGKLIALSASKKK 334
Query: 150 WHIGHASNIPGKIDSLKKRQADFDDKGDVGDLSADEILELRGITHDLHS---LSRVNSSI 206
H S++ + +L+K++A+ + EI++LRG + + + + R+N +
Sbjct: 335 RETAHTSSLTTHLKALEKKEANSPKRS-----RRQEIIKLRGEINQVETRRTIQRINQTR 389
Query: 207 LWQQSRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADS-NLVEGVQPIRNAVLSHFRD 265
W ++ + K + G R + I + + ++ + I+N + S ++
Sbjct: 390 SWFFEKI------NKIDKPLARLTKGHRDKILINKIRNEKGDITTDPEEIQNTIRSFYKR 443
Query: 266 HFAARHTSRVGIDNLPFKC----LSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVN 321
++ + + +D + L+ L P S E+++ + + KSPGPDG
Sbjct: 444 LYSTKLENLDEMDKFLDRYQVPKLNQDQVDHLNSPISPKEIEAVINSLPTKKSPGPDG-- 501
Query: 322 FGFIKEFWDELKEDVMRFISD-FHR---NGKLTRGINTTFIALISKIDS-PQRINDFRPI 376
F EF+ KED++ + FH+ G L I LI K P +I +FRPI
Sbjct: 502 --FSAEFYQTFKEDLIPILHKLFHKIEVEGTLPNSFYEATITLIPKPQKDPTKIENFRPI 559
Query: 377 SLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQ 414
SL+ KIL+K+LANR++ + II Q F+ Q
Sbjct: 560 SLMNIDAKILNKILANRIQEHIKAIIHPDQVGFIPGMQ 597
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 67.8 bits (164), Expect = 2e-10
Identities = 57/266 (21%), Positives = 123/266 (45%), Gaps = 19/266 (7%)
Query: 161 KIDSLKKRQADFDDKGDVGDLSA--DEILELRGITHDLHS---LSRVNSSILWQQSRLLW 215
KID+L + + + + ++ EI+++R ++ + L ++N S W ++
Sbjct: 312 KIDTLISQLKELEKQEQTNSKASRRQEIIKIRAELKEIETQKTLQKINESRSWFFEKI-- 369
Query: 216 LKDGDANSKYFHSVLSGRRRRNSIISLLAD-SNLVEGVQPIRNAVLSHFRDHFAARHTSR 274
+ + ++ +R +N I ++ D ++ I+ + +++ +A + +
Sbjct: 370 ----NKIDRPLARLIKKKREKNQIDTIKNDRGDITTDPTEIQTTIREYYKHLYANKLENL 425
Query: 275 VGIDNL----PFKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWD 330
+D L+ + L +P ++ E+++ + + KSPGP+G F + + +
Sbjct: 426 EEMDKFLDTYTLPRLNQEEVESLNRPITSSEIEAIINSLPNKKSPGPEGFTAEFYQRYKE 485
Query: 331 ELKEDVMRFISDFHRNGKLTRGINTTFIALISK--IDSPQRINDFRPISLVGSLYKILSK 388
EL +++ + G L I LI K D+ ++ N FRPISL+ KIL+K
Sbjct: 486 ELVPFLLKLFQSIEKEGILPNSFYEASIILIPKPGRDTTKKEN-FRPISLMNIDAKILNK 544
Query: 389 VLANRLKLVLGKIISDSQTAFVKDRQ 414
+LAN+++ + K+I Q F+ Q
Sbjct: 545 ILANQIQQHIKKLIHHDQVGFIPAMQ 570
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 47.0 bits (110), Expect = 3e-04
Identities = 96/416 (23%), Positives = 155/416 (37%), Gaps = 76/416 (18%)
Query: 47 WFKGDG-NAMSRIDRFLLSE-------EWCLQ--WPNSFQVALLRGLSDH--CPLQLSVD 94
W+ G N S ID L++E EW +Q W G+SDH +++S+D
Sbjct: 144 WYTFSGPNGSSDIDVTLVNEAGGRFGYEWSVQPEW----------GVSDHNLIRIRVSLD 193
Query: 95 EENWGPRPSRMLKCWQ----DLPGYNQFVKDKLKSFQIEGWGGFVLREKLKF-------- 142
PS WQ D Y VK K F + + + EK+
Sbjct: 194 GLVADASPSPQSARWQTRDTDWGEYMGDVKAKADVFGLAQYENVSVDEKVDLLTEWIYGA 253
Query: 143 ----------IKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGDVGDLS-ADEILELRG 191
++T EW + + K L++R+ F + G S AD + R
Sbjct: 254 NDWNMRRHTAVRTFQNEWW---SVELAEKRSELRRRRHAFQRIRNAGAASLADRLQAFRD 310
Query: 192 ITHDLHSLSRVNSSILWQQSRLLWLKDGDANSKYFHSVLS---GRRRRNSIISLLADSNL 248
+ + WQ+ ++N + V GRR+ + S+ D
Sbjct: 311 CKIEYKRMLCEAKLRCWQE-----FVASESNENPWGRVFKLCRGRRKPVDVCSVKVDGVY 365
Query: 249 VEGVQPIRNAVLSHFRDHFAARHTSRVGIDNLPFKCLSHTDGSGLTKPFSADEVKSAVWD 308
+ + NA+++ F F A ID L K ++ L DEV +V
Sbjct: 366 TDTWEGSVNAMMNVF---FPASIDDASEIDRL--KAIARP----LPPDLEMDEVSDSVRR 416
Query: 309 CDSYKSPGPDGVNFGFIKEFWDELKEDVMRFI------SDFHRNGKLTRGINTTFIALIS 362
C KSPGPDG+ ++ W + E + S F + K+ + + L+
Sbjct: 417 CKVRKSPGPDGIVGEMVRAVWGAIPEYMFCLYKQCLLESYFPQKWKIASLV--ILLKLLD 474
Query: 363 KIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKI-ISDSQTAFVKDRQILD 417
+I S +RPI L+ +L K+L ++ RL L + +S Q AF + D
Sbjct: 475 RIRSDP--GSYRPICLLDNLGKVLEGIMVKRLDQKLMDVEVSPYQFAFTYGKSTED 528
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 38.9 bits (89), Expect = 0.076
Identities = 41/157 (26%), Positives = 62/157 (39%), Gaps = 12/157 (7%)
Query: 286 SHTDGSGLTKPFSADEVKSAVWDCDSY-------KSPGPDGVNFGFIKEFWDELKEDVMR 338
S G + S + V SA+ + D SPGPDG+ +E + +M
Sbjct: 324 SSCSGEVIQMDHSLERVWSAITEQDLRASRVSLSSSPGPDGITPKSAREVPSGIMLRIMN 383
Query: 339 FISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVL 398
I G L I I K + +R DFRPIS+ L + L+ +LA RL +
Sbjct: 384 LILWC---GNLPHSIRLARTVFIPKTVTAKRPQDFRPISVPSVLVRQLNAILATRLNSSI 440
Query: 399 GKIISDSQTAFVKDRQILDGILIANEVVDEARKNKRN 435
Q F+ D I + V+ + K+ R+
Sbjct: 441 N--WDPRQRGFLPTDGCADNATIVDLVLRHSHKHFRS 475
>RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 38.5 bits (88), Expect = 0.099
Identities = 28/100 (28%), Positives = 49/100 (49%), Gaps = 1/100 (1%)
Query: 296 PFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINT 355
P + +EVK V K+PG D ++ I+ D+ ++ + R G + T
Sbjct: 437 PVTLEEVKELVSKLKPKKAPGEDLLDNRTIRLLPDQALLYLVLIFNSILRVGYFPKARPT 496
Query: 356 TFIALISKIDS-PQRINDFRPISLVGSLYKILSKVLANRL 394
I +I K P ++ +RP SL+ SL K+L +++ NR+
Sbjct: 497 ASIIMILKPGKQPLDVDSYRPTSLLPSLGKMLERLILNRI 536
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 38.5 bits (88), Expect = 0.099
Identities = 35/143 (24%), Positives = 63/143 (43%), Gaps = 9/143 (6%)
Query: 293 LTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISD-FHRNGKLTR 351
L P S E+KSA + K GPDGV W+ L + R + + F G++
Sbjct: 155 LWDPVSLIEIKSA--RASNEKGAGPDGVT----PRSWNALDDRYKRLLYNIFVFYGRVPS 208
Query: 352 GINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVK 411
I + KI+ FRP+S+ + + +K+LA R V + QTA++
Sbjct: 209 PIKGSRTVFTPKIEGGPDPGVFRPLSICSVILREFNKILARR--FVSCYTYDERQTAYLP 266
Query: 412 DRQILDGILIANEVVDEARKNKR 434
+ + + ++ EA++ ++
Sbjct: 267 IDGVCINVSMLTAIIAEAKRLRK 289
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 37.4 bits (85), Expect = 0.22
Identities = 32/140 (22%), Positives = 60/140 (42%), Gaps = 12/140 (8%)
Query: 293 LTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRG 352
L +P S DE+K + + GPDG+ W+ + E + + +G+ R
Sbjct: 315 LWRPISNDEIKEV--EACKRTAAGPDGMT----TTAWNSIDECIKSLFNMIMYHGQCPRR 368
Query: 353 INTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGK--IISDSQTAFV 410
+ LI K FRP+S+ + ++LANR +G+ ++ Q AF+
Sbjct: 369 YLDSRTVLIPKEPGTMDPACFRPLSIASVALRHFHRILANR----IGEHGLLDTRQRAFI 424
Query: 411 KDRQILDGILIANEVVDEAR 430
+ + + + ++ EAR
Sbjct: 425 VADGVAENTSLLSAMIKEAR 444
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 37.0 bits (84), Expect = 0.29
Identities = 28/106 (26%), Positives = 53/106 (49%), Gaps = 4/106 (3%)
Query: 293 LTKPFSA---DEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKL 349
++ P SA +EVK+ + K+PG D ++ I+ D+ + + + G
Sbjct: 431 MSLPLSAVTLEEVKNLIAKLPLKKAPGEDLLDNRTIRLLPDQALQFLALIFNSVLDVGYF 490
Query: 350 TRGINTTFIALISKID-SPQRINDFRPISLVGSLYKILSKVLANRL 394
+ + I +I K +P ++ +RP SL+ SL KI+ +++ NRL
Sbjct: 491 PKAWKSASIIMIHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRL 536
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 33.9 bits (76), Expect = 2.4
Identities = 26/100 (26%), Positives = 42/100 (42%), Gaps = 2/100 (2%)
Query: 301 EVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIAL 360
EV + V S +SPG DG+N K W + E + S R G +
Sbjct: 438 EVDTCVARLKSRRSPGLDGINGTICKAVWRAIPEHLASLFSRCIRLGYFPAEWKCPRVVS 497
Query: 361 ISKIDSPQRI--NDFRPISLVGSLYKILSKVLANRLKLVL 398
+ K + + +R I L+ K+L ++ NR++ VL
Sbjct: 498 LLKGPDKDKCEPSSYRGICLLPVFGKVLEAIMVNRVREVL 537
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 33.5 bits (75), Expect = 3.2
Identities = 16/75 (21%), Positives = 38/75 (50%)
Query: 361 ISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGIL 420
I K +P +++RPISL +I+ +++ +R++ ++S Q F+ R ++
Sbjct: 673 IPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHLLSPHQHGFLNFRSCPSSLV 732
Query: 421 IANEVVDEARKNKRN 435
+ + KN+++
Sbjct: 733 RSISLYHSILKNEKS 747
>YL32_ARCFU (O28148) Hypothetical protein AF2132
Length = 244
Score = 33.5 bits (75), Expect = 3.2
Identities = 45/179 (25%), Positives = 77/179 (42%), Gaps = 27/179 (15%)
Query: 873 MVSFCSIISLTGGNGTRILLAVIVWEAPIFILLLHWSLQTFLWETLYGISRCRLRFPFLL 932
++ F ++++ N T + +++ +PIF +LL ++L FL + Y + + + +
Sbjct: 18 VIVFFWLLAVPPENFTADRIVMVLLISPIFAILLAFNLSRFLKQHGYSLRQVHVALEQNI 77
Query: 933 GDYCVIGCRLRIIFLGVA*STVRFVCVSQGAVWKRLHLIFLF------IAPLLVLCGILS 986
D+ RL I L + F+ GA LIF + IA L L ++
Sbjct: 78 ADFIYEFQRLVFIHL----MPIAFL----GA------LIFYYGEFTPAIAEFLSLFALIF 123
Query: 987 GLGSECQVLTLLLLRPISFSLFTLQVTQRLVSHFFNLFGYFVFWWFGTNVIIGFLIISK 1045
L LL ISF + LQV +R+ +F FW + +I +IISK
Sbjct: 124 ALSP-------LLFLGISFFVAMLQVYERVRKKSLLRANFFNFWIWIHLFVILLIIISK 175
>ACEA_CORGL (P42449) Isocitrate lyase (EC 4.1.3.1) (Isocitrase)
(Isocitratase) (ICL)
Length = 431
Score = 33.1 bits (74), Expect = 4.2
Identities = 20/57 (35%), Positives = 33/57 (57%), Gaps = 5/57 (8%)
Query: 156 SNIPGKIDSLKKRQADFDDK----GDVGDLSADEILELRGITHDLHSLSRVNSSILW 208
SN+ GK + ++ Q D+D G D +AD++ +L+G + H+L+R S ILW
Sbjct: 1 SNV-GKPRTAQEIQQDWDTNPRWNGITRDYTADQVADLQGSVIEEHTLARRGSEILW 56
>GYRA_STRPY (Q9L7Q5) DNA gyrase subunit A (EC 5.99.1.3)
Length = 828
Score = 32.7 bits (73), Expect = 5.4
Identities = 33/117 (28%), Positives = 52/117 (44%), Gaps = 22/117 (18%)
Query: 330 DELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKV 389
DE + +RF+ + R T +N F K+ S Q F +++ + KILS
Sbjct: 291 DESSREGVRFVIEIRREASATVILNNLF-----KLTSLQTNFSFNMLAIENGVPKILS-- 343
Query: 390 LANRLKLVLGKIISDSQ------TAFVKDR-----QILDGILIANEVVDEARKNKRN 435
L+ ++ IS + T F KD+ IL+G+LIA + +DE RN
Sbjct: 344 ----LRQIIDNYISHQKEVIIRRTRFDKDKAEARAHILEGLLIALDHLDEVIAIIRN 396
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.344 0.155 0.538
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 120,970,025
Number of Sequences: 164201
Number of extensions: 5049018
Number of successful extensions: 18132
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 18113
Number of HSP's gapped (non-prelim): 17
length of query: 1087
length of database: 59,974,054
effective HSP length: 121
effective length of query: 966
effective length of database: 40,105,733
effective search space: 38742138078
effective search space used: 38742138078
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 71 (32.0 bits)
Medicago: description of AC135464.5


