

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
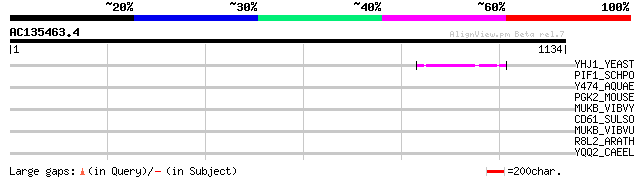
Query= AC135463.4 - phase: 2 /pseudo
(1134 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 57 4e-07
PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1, m... 42 0.009
Y474_AQUAE (O66772) Hypothetical UPF0004 protein AQ_474 35 1.5
PGK2_MOUSE (P09041) Phosphoglycerate kinase, testis specific (EC... 33 3.3
MUKB_VIBVY (Q7MJ64) Chromosome partition protein mukB (Structura... 33 4.4
CD61_SULSO (Q980N4) Cell division control protein 6 homolog 1 (C... 33 4.4
MUKB_VIBVU (Q8DAP8) Chromosome partition protein mukB (Structura... 33 5.7
R8L2_ARATH (Q9MAG6) Probable disease resistance RPP8-like protein 2 32 7.5
YQQ2_CAEEL (Q09305) Hypothetical protein F10B5.2 in chromosome II 32 9.7
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 56.6 bits (135), Expect = 4e-07
Identities = 51/189 (26%), Positives = 88/189 (45%), Gaps = 14/189 (7%)
Query: 831 EKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRS-EGKIVLNVASSSIASLLLPRG 889
E++VN++ + ++F+ G G GK+ + + S GK + + +S+ + + G
Sbjct: 237 ERVVNLIVKKRTNVFYT-GSAGTGKSVILQTIIRQLSSLYGKESIAITASTGLAAVTIGG 295
Query: 890 RTAH--SQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMR 947
T H S I +++ I+ + + ++I DE M+ + ++ R
Sbjct: 296 STLHKWSGIGIGNKTIDQLVKKIQSQKDLLAAWRYTKVLIIDEISMVDGNLLDKLEQIAR 355
Query: 948 DIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRC--RVL 1005
I N + PFGG +VL GDF Q LP V K+ ++V C S M W+RC + +
Sbjct: 356 RIRKN------DDPFGGIQLVLTGDFFQ-LPPVAKKDEHNVVKFCFESEM-WKRCIQKTI 407
Query: 1006 KLTKNIRLQ 1014
LTK R Q
Sbjct: 408 LLTKVFRQQ 416
>PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1,
mitochondrial precursor
Length = 805
Score = 42.0 bits (97), Expect = 0.009
Identities = 47/179 (26%), Positives = 73/179 (40%), Gaps = 19/179 (10%)
Query: 813 RTLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWN----ALSYHFRS 868
R + P L+ +Q + +V E H F G G GK+ L L +R
Sbjct: 301 RKKSVPSIFLSDEQKRILDMVV-----EQQHSIFFTGSAGTGKSVLLRKIIEVLKSKYRK 355
Query: 869 EGKIVLNVASSSIASLLLPRGRTAHSQFSIPLML--LEESGCTIEKEGNKAQLLTMASLI 926
+ V AS+ +A+ + G T HS + L ++ I+K ++
Sbjct: 356 QSDRVAVTASTGLAACNIG-GVTLHSFAGVGLARESVDLLVSKIKKNKKCVNRWLRTRVL 414
Query: 927 IWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGR 985
I DE M+ + + R I + + PFGG +VL GDF Q LP VP+ G+
Sbjct: 415 IIDEVSMVDAELMDKLEEVARVIRKD------SKPFGGIQLVLTGDFFQ-LPPVPENGK 466
>Y474_AQUAE (O66772) Hypothetical UPF0004 protein AQ_474
Length = 410
Score = 34.7 bits (78), Expect = 1.5
Identities = 32/127 (25%), Positives = 52/127 (40%), Gaps = 11/127 (8%)
Query: 174 RVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHS--KYIPLQYPVLFPL 231
R Y E EVV IV + G D++V ET+ K IP+ Y +FP
Sbjct: 270 RGYTREEYEEVVNFIVENRPISSIGTDVIVGFPTESEEDFQETYEFLKRIPISYMHIFPY 329
Query: 232 GEDRYEEKIKLNHRTTTGSVKKRVRV-------SLREFVAYRLQKRDNENSVIFQGMRLF 284
+ + + KL + K+RVR+ +EF Y K +++ + RL
Sbjct: 330 SDRPFTKASKLKPKLPERIKKERVRILKELDQKKRQEF--YEKNKGKELRALVIEENRLL 387
Query: 285 QQFIVDL 291
+ +D+
Sbjct: 388 TENYIDI 394
>PGK2_MOUSE (P09041) Phosphoglycerate kinase, testis specific (EC
2.7.2.3)
Length = 416
Score = 33.5 bits (75), Expect = 3.3
Identities = 40/160 (25%), Positives = 66/160 (41%), Gaps = 26/160 (16%)
Query: 834 VNVVDNELDHMFFVDGYGGIGKTFLWNALSYH-----FRSEG----KIVLNVASSSIASL 884
+ ++ N LD + F+ GG+ TFL + F EG K ++ A + +
Sbjct: 220 IQLIKNMLDKVNFMIIGGGMAYTFLKELKNMQIGASLFDEEGATIVKEIMEKAEKNGVKI 279
Query: 885 LLPRGRTAHSQFSIPLMLLE---ESG---------CTIEKEGNKAQLLTMASLIIWDEAP 932
+ P +F + + ESG C E AQ++ A LI+W+
Sbjct: 280 VFPVDFVTGDKFDENAKVGQATIESGIPSGWMGLDCGPESIKINAQIVAQAKLIVWNGP- 338
Query: 933 MISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGD 972
I ++AF + + +M VV SN G T++ GGD
Sbjct: 339 -IGVFEWDAFAKGTKALMDEVVKATSN---GCVTIIGGGD 374
>MUKB_VIBVY (Q7MJ64) Chromosome partition protein mukB (Structural
maintenance of chromosome related protein)
Length = 1487
Score = 33.1 bits (74), Expect = 4.4
Identities = 33/156 (21%), Positives = 74/156 (47%), Gaps = 21/156 (13%)
Query: 172 DSRVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGL-RRIHETHSKYIPLQYPVLFP 230
+ ++ + + E + +I GD D+F+D I E +G + R+++ +Y ++PV+
Sbjct: 722 EEKLVELDDCPEDLYIIEGDVDAFDDS-SIKAEELEGAVCVRLNDRQMRY--SRFPVIPL 778
Query: 231 LGEDRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVD 290
G E++++L V+K + + F A ++Q+ +FQ F QF+ +
Sbjct: 779 FGRAAREQRLELLRSEREEVVEKHAKAA---FDAQKMQR-------LFQA---FNQFVAE 825
Query: 291 ----LYSMVENQRLSYVRKNQSQIRSGLLWLDSRDK 322
++ Q L R ++Q+ L L+++++
Sbjct: 826 HIQVAFAADPEQALVIARDKRNQLTRTLAELEAKEQ 861
>CD61_SULSO (Q980N4) Cell division control protein 6 homolog 1 (CDC6
homolog 1)
Length = 397
Score = 33.1 bits (74), Expect = 4.4
Identities = 18/55 (32%), Positives = 32/55 (57%), Gaps = 1/55 (1%)
Query: 718 RLLLNRQIGCTSFEDIRTVDGHVYNSYREACAALGLLA-NDRQFIDLISEIVVVG 771
+L+L + +S E++ + G VY +Y C LG+ A R+ D+I+E+ +VG
Sbjct: 300 KLVLMAVVSISSEENVVSTTGAVYETYLNICKKLGVEAVTQRRVSDIINELDMVG 354
>MUKB_VIBVU (Q8DAP8) Chromosome partition protein mukB (Structural
maintenance of chromosome related protein)
Length = 1487
Score = 32.7 bits (73), Expect = 5.7
Identities = 33/155 (21%), Positives = 73/155 (46%), Gaps = 21/155 (13%)
Query: 172 DSRVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGL-RRIHETHSKYIPLQYPVLFP 230
+ ++ + + E + +I GD D+F+D I E +G + R+++ +Y ++PV+
Sbjct: 722 EEKLVELDDCPEDLYIIEGDVDAFDDS-SIKAEELEGAVCVRLNDRQMRY--SRFPVIPL 778
Query: 231 LGEDRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVD 290
G E++++L V+K + + F A ++Q+ +FQ F QF+ +
Sbjct: 779 FGRAAREQRLELLRSEREEVVEKHAKAA---FDAQKMQR-------LFQA---FNQFVAE 825
Query: 291 ----LYSMVENQRLSYVRKNQSQIRSGLLWLDSRD 321
++ Q L R ++Q+ L L++++
Sbjct: 826 HIQVAFAADPEQALVIARDKRNQLTRTLAELEAKE 860
>R8L2_ARATH (Q9MAG6) Probable disease resistance RPP8-like protein 2
Length = 1271
Score = 32.3 bits (72), Expect = 7.5
Identities = 23/64 (35%), Positives = 34/64 (52%), Gaps = 7/64 (10%)
Query: 808 LYERRRTL------NSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNA 861
L ER+R + NS L+ DQ S E + ++V+N+ + V G GGIGKT L
Sbjct: 144 LQERQREIRQTFSRNSESDLVGLDQ-SVEELVDHLVENDSVQVVSVSGMGGIGKTTLARQ 202
Query: 862 LSYH 865
+ +H
Sbjct: 203 VFHH 206
>YQQ2_CAEEL (Q09305) Hypothetical protein F10B5.2 in chromosome II
Length = 357
Score = 32.0 bits (71), Expect = 9.7
Identities = 13/43 (30%), Positives = 24/43 (55%), Gaps = 3/43 (6%)
Query: 674 LTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYC 716
L ++ FP+ + ++ +SW K+G+ L +PPG +YC
Sbjct: 20 LLFLGFPQASEFGIDYKSW---KTGEKFMGLKMIPPGVHFVYC 59
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.337 0.148 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 126,961,792
Number of Sequences: 164201
Number of extensions: 5302095
Number of successful extensions: 18779
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 18778
Number of HSP's gapped (non-prelim): 9
length of query: 1134
length of database: 59,974,054
effective HSP length: 121
effective length of query: 1013
effective length of database: 40,105,733
effective search space: 40627107529
effective search space used: 40627107529
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC135463.4


