

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135320.7 + phase: 0
(190 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
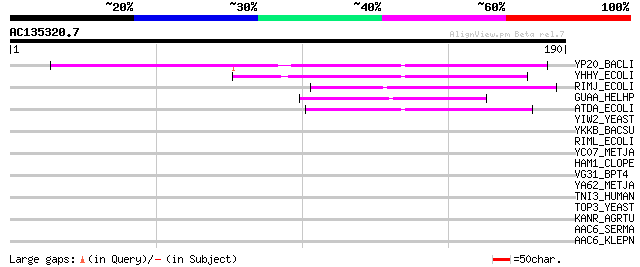
YP20_BACLI (P05332) Hypothetical P20 protein 79 8e-15
YHHY_ECOLI (P46854) Hypothetical acetyltransferase yhhY (EC 2.3.... 45 8e-05
RIMJ_ECOLI (P09454) Ribosomal-protein-alanine acetyltransferase ... 44 2e-04
GUAA_HELHP (Q7VG78) Probable GMP synthase [glutamine-hydrolyzing... 42 7e-04
ATDA_ECOLI (P37354) Spermidine N(1)-acetyltransferase (EC 2.3.1.... 42 7e-04
YIW2_YEAST (P40586) Hypothetical 27.4 kDa protein in HYR1 3'region 42 0.001
YKKB_BACSU (P49855) Hypothetical protein ykkB 41 0.002
RIML_ECOLI (P13857) Ribosomal-protein-serine acetyltransferase (... 38 0.016
YC07_METJA (Q58604) Hypothetical acetyltransferase MJ1207 (EC 2.... 32 0.68
HAM1_CLOPE (Q8XI68) HAM1 protein homolog 32 1.2
VG31_BPT4 (P17313) Capsid assembly protein Gp31 30 3.4
YA62_METJA (Q58462) Hypothetical protein MJ1062 30 4.4
TNI3_HUMAN (P21580) Tumor necrosis factor, alpha-induced protein... 29 5.7
TOP3_YEAST (P13099) DNA topoisomerase III (EC 5.99.1.2) 29 7.5
KANR_AGRTU (P14510) Kanamycin resistance protein (KM-R) (Fragment) 29 7.5
AAC6_SERMA (P20092) Aminoglycoside N(6')-acetyltransferase (EC 2... 29 7.5
AAC6_KLEPN (P19650) Aminoglycoside N(6')-acetyltransferase (EC 2... 29 7.5
>YP20_BACLI (P05332) Hypothetical P20 protein
Length = 178
Score = 78.6 bits (192), Expect = 8e-15
Identities = 56/176 (31%), Positives = 88/176 (49%), Gaps = 11/176 (6%)
Query: 15 ERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCK 74
E + ++TLR + L D D L + +D +V K+ + +T + I+ I L +
Sbjct: 2 ETLYTERLTLRKMELEDADVLCQYWSDPEVTKYMNITPFTDVSQARDMIQMINDLSLEGQ 61
Query: 75 A------ICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKV 128
A + D IG + N AE+GY LG +WGKG A+ V++++
Sbjct: 62 ANRFSIIVKETDEVIGTCGFNMID----QENGRAEIGYDLGRNHWGKGFASEAVQKLIDY 117
Query: 129 AFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVL 184
F L+ L R+EA V+ EN S ++L FQKEG+LR Y KG+ D+ + S+L
Sbjct: 118 GFTSLN-LNRIEAKVEPENTPSIKLLNSLSFQKEGLLRDYEKAKGRLIDVYMFSLL 172
>YHHY_ECOLI (P46854) Hypothetical acetyltransferase yhhY (EC
2.3.1.-)
Length = 162
Score = 45.4 bits (106), Expect = 8e-05
Identities = 27/101 (26%), Positives = 58/101 (56%), Gaps = 3/101 (2%)
Query: 77 CINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYL 136
CI+ +G +++ R+ A+ G + S++ +GVA+ ++++++++ L +
Sbjct: 57 CIDGDVVGHLTIDVQQR--PRRSHVADFGICVDSRWKNRGVASALMREMIEMCDNWLR-V 113
Query: 137 ERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRD 177
+R+E V V+NA + +V +K GF+ EG +KY + G+ D
Sbjct: 114 DRIELTVFVDNAPAIKVYKKYGFEIEGTGKKYALRNGEYVD 154
>RIMJ_ECOLI (P09454) Ribosomal-protein-alanine acetyltransferase (EC
2.3.1.128) (Acetylating enzyme for N-terminal of
ribosomal protein S5)
Length = 194
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/84 (28%), Positives = 45/84 (53%), Gaps = 1/84 (1%)
Query: 104 LGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKEG 163
LGY +G K+ GKG+ + ++ ++ R+ A N S +L + GF+KEG
Sbjct: 107 LGYSIGQKWQGKGLMFEALTAAIRY-MQRTQHIHRIMANYMPHNKRSGDLLARLGFEKEG 165
Query: 164 VLRKYLVMKGKSRDMIISSVLFTD 187
+ YL++ G+ RD +++++ D
Sbjct: 166 YAKDYLLIDGQWRDHVLTALTTPD 189
>GUAA_HELHP (Q7VG78) Probable GMP synthase [glutamine-hydrolyzing]
(EC 6.3.5.2) (Glutamine amidotransferase) (GMP
synthetase)
Length = 1375
Score = 42.4 bits (98), Expect = 7e-04
Identities = 23/64 (35%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Query: 100 KCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGF 159
K E+GY+L +WGKG + V + VK F E LE + L+ +N S +V ++
Sbjct: 401 KIVEIGYLLHRDFWGKGYGSEVARMCVKYGF-ETLGLEEVYCLIKEDNTASIKVAKRLEM 459
Query: 160 QKEG 163
QK G
Sbjct: 460 QKVG 463
>ATDA_ECOLI (P37354) Spermidine N(1)-acetyltransferase (EC 2.3.1.57)
(Diamine acetyltransferase) (SAT)
Length = 185
Score = 42.4 bits (98), Expect = 7e-04
Identities = 26/78 (33%), Positives = 39/78 (49%), Gaps = 1/78 (1%)
Query: 102 AELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQK 161
AE ++ +Y GKG+AT K + F L+ L +L +VD EN + + K GF
Sbjct: 83 AEFQIIISPEYQGKGLATRAAKLAMDYGFTVLN-LYKLYLIVDKENEKAIHIYRKLGFSV 141
Query: 162 EGVLRKYLVMKGKSRDMI 179
EG L + G+ R+ I
Sbjct: 142 EGELMHEFFINGQYRNAI 159
>YIW2_YEAST (P40586) Hypothetical 27.4 kDa protein in HYR1 3'region
Length = 236
Score = 41.6 bits (96), Expect = 0.001
Identities = 33/99 (33%), Positives = 50/99 (50%), Gaps = 6/99 (6%)
Query: 80 DRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGV-ATCVVKQVVKVAFCELSYLER 138
+RA+G + L + S E+GYV+ S K + AT ++K F +L Y R
Sbjct: 98 ERAVGTLCLIRIDEANGS----LEVGYVVFSPELQKTIIATEAQFLLMKYVFDDLQY-RR 152
Query: 139 LEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRD 177
E D N S+R + GF+ EG R+ +V KG++RD
Sbjct: 153 YEWKCDSLNGPSRRAAMRLGFKYEGTFRQVVVYKGRTRD 191
>YKKB_BACSU (P49855) Hypothetical protein ykkB
Length = 172
Score = 40.8 bits (94), Expect = 0.002
Identities = 21/66 (31%), Positives = 33/66 (49%), Gaps = 1/66 (1%)
Query: 103 ELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKE 162
E+GY+ ++WG G A + + F E + ++ AL+D N S RV EK G
Sbjct: 95 EIGYMFARRHWGNGYAQEAARACLDYGFNERQF-GKMAALIDPGNKASIRVAEKIGMHYS 153
Query: 163 GVLRKY 168
+RK+
Sbjct: 154 KTIRKW 159
>RIML_ECOLI (P13857) Ribosomal-protein-serine acetyltransferase (EC
2.3.1.-) (Acetylating enzyme for N-terminal of ribosomal
protein L7/L12)
Length = 179
Score = 37.7 bits (86), Expect = 0.016
Identities = 25/88 (28%), Positives = 43/88 (48%), Gaps = 5/88 (5%)
Query: 80 DRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERL 139
D IG +S + P NK AE+GY L + G+G+ + ++ ++ + + L R
Sbjct: 76 DELIGVISFNRIEP----LNKTAEIGYWLDESHQGQGIISQALQALIH-HYAQSGELRRF 130
Query: 140 EALVDVENAGSQRVLEKAGFQKEGVLRK 167
V+N S +V + GF EG L++
Sbjct: 131 VIKCRVDNPQSNQVALRNGFILEGCLKQ 158
>YC07_METJA (Q58604) Hypothetical acetyltransferase MJ1207 (EC
2.3.1.-)
Length = 226
Score = 32.3 bits (72), Expect = 0.68
Identities = 20/95 (21%), Positives = 50/95 (52%), Gaps = 3/95 (3%)
Query: 78 INDRAIGCVSLSSSSPGDKSRNKCAELGYV-LGSKYWGKGVATCVVKQVVKVAFCELSYL 136
+N + +G V+ + + + + AE+ + + + G+G+ T ++ + ++ A +
Sbjct: 130 VNGKPVGFVACDCNWISNIEKREVAEIHEIFVDPDFRGRGIGTALINKAIEYA--KKRGR 187
Query: 137 ERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVM 171
+E V VEN G+ ++ GF+++ V++ +L M
Sbjct: 188 RIVELWVGVENKGAIEFYKRLGFEEKEVVKGWLRM 222
>HAM1_CLOPE (Q8XI68) HAM1 protein homolog
Length = 204
Score = 31.6 bits (70), Expect = 1.2
Identities = 19/59 (32%), Positives = 34/59 (57%), Gaps = 3/59 (5%)
Query: 122 VKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMII 180
+K+++K L+ L EA +D++ + E+ F+K +RKYL+ KG+S D I+
Sbjct: 17 IKEILKDF--NLNILSLNEAGIDIDVEEDGKTFEENSFKKANEIRKYLLSKGES-DFIV 72
>VG31_BPT4 (P17313) Capsid assembly protein Gp31
Length = 111
Score = 30.0 bits (66), Expect = 3.4
Identities = 19/62 (30%), Positives = 27/62 (42%), Gaps = 3/62 (4%)
Query: 81 RAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQV---VKVAFCELSYLE 137
RA+G + S P + E G ++G + G+ CVV V V FCE+ L
Sbjct: 10 RAVGEYVILVSEPAQAGDEEVTESGLIIGKRVQGEVPELCVVHSVGPDVPEGFCEVGDLT 69
Query: 138 RL 139
L
Sbjct: 70 SL 71
>YA62_METJA (Q58462) Hypothetical protein MJ1062
Length = 484
Score = 29.6 bits (65), Expect = 4.4
Identities = 32/146 (21%), Positives = 69/146 (46%), Gaps = 12/146 (8%)
Query: 20 TQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSK-DDGINFIENIATKFLWCKAICI 78
++I +R + +DL+ LM W ++ + KF + K ++ ++ + + W +
Sbjct: 332 SKIIIRQITDNDLELLMAWRSNPLIYKFFYIQKEPLKWEEHYSWWMSRENRVDWIILLRE 391
Query: 79 ND--RAIGCVSLSSSSPGDKSRNKCAELGYVLGSKY-WGKGVATCVVKQVVKVAFCELSY 135
N+ R +G V++S + + E+G ++G + WGK + V V+K + +
Sbjct: 392 NNTIRKVGSVNVSQLNTDN------PEIGILIGEFFLWGKHIGRHSVSLVLK--WLKNIG 443
Query: 136 LERLEALVDVENAGSQRVLEKAGFQK 161
++ A + N S ++ E GF+K
Sbjct: 444 YKKAHARILENNIRSIKLFESLGFKK 469
>TNI3_HUMAN (P21580) Tumor necrosis factor, alpha-induced protein 3
(EC 3.-.-.-) (Putative DNA binding protein A20) (Zinc
finger protein A20)
Length = 790
Score = 29.3 bits (64), Expect = 5.7
Identities = 15/42 (35%), Positives = 23/42 (54%), Gaps = 1/42 (2%)
Query: 79 NDRAIGCVSLSSSSPGDKS-RNKCAELGYVLGSKYWGKGVAT 119
ND GC+S ++ +PGD++ +KC + G V KG T
Sbjct: 584 NDAPAGCLSQAARTPGDRTGTSKCRKAGCVYFGTPENKGFCT 625
>TOP3_YEAST (P13099) DNA topoisomerase III (EC 5.99.1.2)
Length = 656
Score = 28.9 bits (63), Expect = 7.5
Identities = 11/33 (33%), Positives = 18/33 (54%)
Query: 33 DDLMIWTTDEKVAKFCSWELYTSKDDGINFIEN 65
D LMIWT ++ ++ WE++ G I+N
Sbjct: 113 DYLMIWTDCDREGEYIGWEIWQEAKRGNRLIQN 145
>KANR_AGRTU (P14510) Kanamycin resistance protein (KM-R) (Fragment)
Length = 108
Score = 28.9 bits (63), Expect = 7.5
Identities = 15/52 (28%), Positives = 31/52 (58%), Gaps = 1/52 (1%)
Query: 114 GKGVATCVVKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVL 165
GKG+ T +V+ +V++ F + + +++ N + R EKAGF+++G +
Sbjct: 34 GKGLGTKLVRALVELLFNDPE-VTKIQTDPSPSNLRAIRCYEKAGFERQGTV 84
>AAC6_SERMA (P20092) Aminoglycoside N(6')-acetyltransferase (EC
2.3.1.82) (AAC(6'))
Length = 201
Score = 28.9 bits (63), Expect = 7.5
Identities = 15/52 (28%), Positives = 31/52 (58%), Gaps = 1/52 (1%)
Query: 114 GKGVATCVVKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVL 165
GKG+ T +V+ +V++ F + + +++ N + R EKAGF+++G +
Sbjct: 127 GKGLGTKLVRALVELLFNDPE-VTKIQTDPSPSNLRAIRCYEKAGFERQGTV 177
>AAC6_KLEPN (P19650) Aminoglycoside N(6')-acetyltransferase (EC
2.3.1.82) (Amikacin resistance protein) (AAC(6'))
Length = 201
Score = 28.9 bits (63), Expect = 7.5
Identities = 15/52 (28%), Positives = 31/52 (58%), Gaps = 1/52 (1%)
Query: 114 GKGVATCVVKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVL 165
GKG+ T +V+ +V++ F + + +++ N + R EKAGF+++G +
Sbjct: 127 GKGLGTKLVRALVELLFNDPE-VTKIQTDPSPSNLRAIRCYEKAGFERQGTV 177
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,509,311
Number of Sequences: 164201
Number of extensions: 824090
Number of successful extensions: 1713
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1701
Number of HSP's gapped (non-prelim): 17
length of query: 190
length of database: 59,974,054
effective HSP length: 104
effective length of query: 86
effective length of database: 42,897,150
effective search space: 3689154900
effective search space used: 3689154900
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135320.7


