

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135317.6 + phase: 0
(432 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
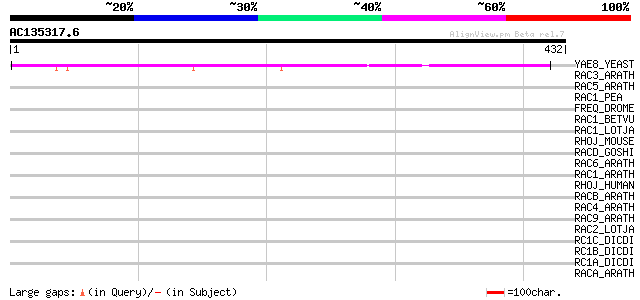
Score E
Sequences producing significant alignments: (bits) Value
YAE8_YEAST (P39722) Hypothetical 75.2 kDa protein in ACS1-GCV3 i... 194 5e-49
RAC3_ARATH (Q38912) RAC-like GTP binding protein ARAC3 (GTPase p... 40 0.012
RAC5_ARATH (Q38937) RAC-like GTP binding protein ARAC5 (GTPase p... 38 0.044
RAC1_PEA (Q35638) RAC-like GTP binding protein RHO1 (GTPase prot... 37 0.075
FREQ_DROME (P37236) Frequenin (d-FRQ) 37 0.075
RAC1_BETVU (Q39435) RAC-like GTP binding protein RHO1 (RHO1Bv) 37 0.098
RAC1_LOTJA (O04369) RAC-like GTP binding protein RAC1 37 0.13
RHOJ_MOUSE (Q9ER71) Rho-related GTP-binding protein RhoJ (Tc10-l... 36 0.17
RACD_GOSHI (Q41253) RAC-like GTP binding protein RAC13 36 0.17
RAC6_ARATH (Q9SBJ6) RAC-like GTP binding protein ARAC6 (GTPase p... 36 0.22
RAC1_ARATH (Q38902) RAC-like GTP binding protein ARAC1 (GTPase p... 36 0.22
RHOJ_HUMAN (Q9H4E5) Rho-related GTP-binding protein RhoJ (Tc10-l... 35 0.29
RACB_ARATH (P92978) RAC-like GTP binding protein ARAC11 (GTPase ... 35 0.29
RAC4_ARATH (Q38919) RAC-like GTP binding protein ARAC4 (GTPase p... 35 0.37
RAC9_ARATH (Q9XGU0) RAC-like GTP binding protein ARAC9 (GTPase p... 35 0.49
RAC2_LOTJA (Q40220) RAC-like GTP binding protein RAC2 35 0.49
RC1C_DICDI (P34146) RAS-related protein rac1C 34 0.64
RC1B_DICDI (P34145) RAS-related protein rac1B 34 0.64
RC1A_DICDI (P34144) RAS-related protein rac1A 34 0.64
RACA_ARATH (O82481) RAC-like GTP binding protein ARAC10 (GTPase ... 34 0.64
>YAE8_YEAST (P39722) Hypothetical 75.2 kDa protein in ACS1-GCV3
intergenic region
Length = 662
Score = 194 bits (492), Expect = 5e-49
Identities = 138/439 (31%), Positives = 216/439 (48%), Gaps = 25/439 (5%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVL---RNVPEGVN------SLGLTFPGF 52
D L D+E+ Q + F S+ ++ +K ++L ++ E +N G+T GF
Sbjct: 218 DSYLDDNEILGLQKKCFNKSIDVNELNFIKDLLLDISKHDQEYINRKLYVPGKGITKDGF 277
Query: 53 IEIHNMFLKKGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKG 112
+ ++ ++ ++GR ET WA+LR F Y + L + D L SVELS FL
Sbjct: 278 LVLNKIYAERGRHETTWAILRTFHYTDSLCINDKILHPRLVVPDTSSVELSPKGYRFLVD 337
Query: 113 VFRLVDTDKDQLLRPAEVDKLFDAAPESP--WKDAPYMDAAETTDMGYISLKGFLSRWAL 170
+F D D D L E+ +LF P P W + + + G I+L+G+L++W++
Sbjct: 338 IFLKFDIDNDGGLNNQELHRLFKCTPGLPKLWTSTNFPFSTVVNNKGCITLQGWLAQWSM 397
Query: 171 MTLLDPRYSLANLICIGYRDNPSAALRLTSRRSEDRKNQK------TERNVFQCYVFGSK 224
T L+ + A L+ G++++ AL++T R R++ K +R VF C+V G
Sbjct: 398 TTFLNYSTTTAYLVYFGFQEDARLALQVTKPRKMRRRSGKLYRSNINDRKVFNCFVIGKP 457
Query: 225 TAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNK 284
GKS+LL A LGR FS +Y+PT R A N +EL G + LIL+E+ E E + L NK
Sbjct: 458 CCGKSSLLEAFLGRSFSEEYSPTIKPRIAVNSLELKGGKQYYLILQELGEQEYA-ILENK 516
Query: 285 DCLAACDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDL-TPFPRA 343
D L CDV +DSSD S+ + LL+K + +L P + +A+K DL R
Sbjct: 517 DKLKECDVICLTYDSSDPESFSYLVSLLDKFTHLQDL-----PLVFVASKADLDKQQQRC 571
Query: 344 VLDSVKVAQELKIDAPIRLSIKSDDS-NNVYSKIINAAEHPHLSIPETEFVRKRKQHQQL 402
+ ++A EL ++ P+ +S + S N ++ KI AA P + P K
Sbjct: 572 QIQPDELADELFVNHPLHISSRWLSSLNELFIKITEAALDPGKNTPGLPEETAAKDVDYR 631
Query: 403 LHTFIFALAGAAMALAGFT 421
IF +AL FT
Sbjct: 632 QTALIFGSTVGFVALCSFT 650
>RAC3_ARATH (Q38912) RAC-like GTP binding protein ARAC3 (GTPase
protein ROP6)
Length = 198
Score = 40.0 bits (92), Expect = 0.012
Identities = 51/194 (26%), Positives = 76/194 (38%), Gaps = 18/194 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN+I + G L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVI--VDGNTINLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K V + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFLLAFSLVSKASYE-----NVSKKWVPELRHYAPGVPIILVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV S +ELK I AP + + NV + +AA L
Sbjct: 121 DLRDDKQFFAEHPGAVPISTAQGEELKKLIGAPAYIECSAKTQQNV-KAVFDAAIKVVLQ 179
Query: 387 IPETEFVRKRKQHQ 400
P+ + +KRK +
Sbjct: 180 PPKNKKKKKRKSQK 193
>RAC5_ARATH (Q38937) RAC-like GTP binding protein ARAC5 (GTPase
protein ROP4)
Length = 196
Score = 38.1 bits (87), Expect = 0.044
Identities = 47/191 (24%), Positives = 75/191 (38%), Gaps = 18/191 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ +L + F DY PT + ++AN++ + G L L + E
Sbjct: 8 KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV--VDGNTVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVAKKWIPELRHYAPGVPIILVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +ELK I +PI + S NV + +AA L
Sbjct: 121 DLRDDKQFFIDHPGAVPITTNQGEELKKLIGSPIYIECSSKTQQNV-KAVFDAAIKVVLQ 179
Query: 387 IPETEFVRKRK 397
P+ + +K K
Sbjct: 180 PPKQKKKKKNK 190
>RAC1_PEA (Q35638) RAC-like GTP binding protein RHO1 (GTPase protein
ROP1)
Length = 197
Score = 37.4 bits (85), Expect = 0.075
Identities = 46/192 (23%), Positives = 77/192 (39%), Gaps = 18/192 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGSTVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIILVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +EL+ I+AP + S NV + +AA L
Sbjct: 121 DLRDDKQFFVDHPGAVPITTAQGEELRKLINAPAYIECSSKSQQNV-KAVFDAAIRVVLQ 179
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 180 PPKQKKKKSKAQ 191
>FREQ_DROME (P37236) Frequenin (d-FRQ)
Length = 186
Score = 37.4 bits (85), Expect = 0.075
Identities = 34/115 (29%), Positives = 51/115 (43%), Gaps = 13/115 (11%)
Query: 24 QPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLKKGRTETFWA-VLRKFGYGNDLK 82
+ +I Q L++ P G+ LT GFI+I+ F +G F + V R F ND
Sbjct: 23 EKEIRQWHKGFLKDCPNGL----LTEQGFIKIYKQFFPQGDPSKFASLVFRVFDENNDGS 78
Query: 83 LR-DDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDKDQLLRPAEVDKLFDA 136
+ ++F+ SV G E L+ FRL D D D + E+ + DA
Sbjct: 79 IEFEEFIRA-------LSVTSKGNLDEKLQWAFRLYDVDNDGYITREEMYNIVDA 126
>RAC1_BETVU (Q39435) RAC-like GTP binding protein RHO1 (RHO1Bv)
Length = 197
Score = 37.0 bits (84), Expect = 0.098
Identities = 47/192 (24%), Positives = 76/192 (39%), Gaps = 18/192 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGATVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIVLVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +EL+ I AP + S NV + +AA L
Sbjct: 121 DLRDDKQFFIDHPGAVPITTAQGEELRKLIGAPAYIECSSKTQQNV-KAVFDAAIKVVLQ 179
Query: 387 IPETEFVRKRKQ 398
P+T+ + + Q
Sbjct: 180 PPKTKKKKSKAQ 191
>RAC1_LOTJA (O04369) RAC-like GTP binding protein RAC1
Length = 197
Score = 36.6 bits (83), Expect = 0.13
Identities = 48/195 (24%), Positives = 77/195 (38%), Gaps = 21/195 (10%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VDGSTVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NIAKKWIPELRHYAPGVPIILVGTKL 120
Query: 336 D-------LTPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
D L P AV + +EL+ I AP + S NV + +AA L
Sbjct: 121 DLRDDKHFLADHPGAVPITTAQGEELRKLIGAPAYIECSSKTQQNV-KAVFDAAIKVVLQ 179
Query: 387 IPETEFVRKRKQHQQ 401
P+ +K+K+ Q
Sbjct: 180 PPKQ---KKKKREAQ 191
>RHOJ_MOUSE (Q9ER71) Rho-related GTP-binding protein RhoJ (Tc10-like
GTP-binding protein TCL)
Length = 214
Score = 36.2 bits (82), Expect = 0.17
Identities = 15/49 (30%), Positives = 22/49 (44%)
Query: 208 NQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANI 256
N E + +C V G GK+ LL + F +Y PT + YA +
Sbjct: 14 NGHEENRILKCVVVGDGAVGKTCLLMSYANDAFPEEYVPTVFDHYAVTV 62
>RACD_GOSHI (Q41253) RAC-like GTP binding protein RAC13
Length = 196
Score = 36.2 bits (82), Expect = 0.17
Identities = 40/166 (24%), Positives = 65/166 (39%), Gaps = 17/166 (10%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ +L + F DY PT + ++AN++ + G+ L L + E
Sbjct: 8 KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV--VDGSTVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + H P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFLLAFSLISKASYE-----NIYKKWIPELRHYAHNVPVVLVGTKL 120
Query: 336 DLTPFPRAVLD-------SVKVAQELK--IDAPIRLSIKSDDSNNV 372
DL + ++D S +ELK I A + S NV
Sbjct: 121 DLRDDKQFLIDHPGATPISTSQGEELKKMIGAVTYIECSSKTQQNV 166
>RAC6_ARATH (Q9SBJ6) RAC-like GTP binding protein ARAC6 (GTPase
protein ROP5)
Length = 197
Score = 35.8 bits (81), Expect = 0.22
Identities = 47/192 (24%), Positives = 75/192 (38%), Gaps = 18/192 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGATVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIVLVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +ELK I AP + S NV + +AA L
Sbjct: 121 DLRDDKQFFIDHPGAVPITTVQGEELKKLIGAPAYIECSSKSQENV-KGVFDAAIRVVLQ 179
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 180 PPKQKKKKNKAQ 191
>RAC1_ARATH (Q38902) RAC-like GTP binding protein ARAC1 (GTPase
protein ROP3)
Length = 197
Score = 35.8 bits (81), Expect = 0.22
Identities = 47/192 (24%), Positives = 75/192 (38%), Gaps = 18/192 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGATVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIVLVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +ELK I AP + S NV + +AA L
Sbjct: 121 DLRDDKQFFIDHPGAVPITTAQGEELKKLIGAPAYIECSSKTQENV-KGVFDAAIRVVLQ 179
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 180 PPKQKKKKSKAQ 191
>RHOJ_HUMAN (Q9H4E5) Rho-related GTP-binding protein RhoJ (Tc10-like
GTP-binding protein TCL)
Length = 214
Score = 35.4 bits (80), Expect = 0.29
Identities = 14/45 (31%), Positives = 22/45 (48%)
Query: 212 ERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANI 256
E+ + +C V G GK+ LL + F +Y PT + YA +
Sbjct: 18 EKKMLKCVVVGDGAVGKTCLLMSYANDAFPEEYVPTVFDHYAVTV 62
>RACB_ARATH (P92978) RAC-like GTP binding protein ARAC11 (GTPase
protein ROP1)
Length = 197
Score = 35.4 bits (80), Expect = 0.29
Identities = 46/192 (23%), Positives = 76/192 (38%), Gaps = 18/192 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGSTVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIVLVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +EL+ I AP + S NV + +AA L
Sbjct: 121 DLRDDKQFFIDHPGAVPITTAQGEELRKQIGAPTYIECSSKTQENV-KAVFDAAIRVVLQ 179
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 180 PPKQKKKKSKAQ 191
>RAC4_ARATH (Q38919) RAC-like GTP binding protein ARAC4 (GTPase
protein ROP2)
Length = 195
Score = 35.0 bits (79), Expect = 0.37
Identities = 45/191 (23%), Positives = 74/191 (38%), Gaps = 18/191 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ +L + F DY PT + ++AN++ + G L L + E
Sbjct: 7 KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV--VDGNTVNLGLWDTAGQE 64
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + P +L+ K
Sbjct: 65 DYNRLRPLSYRGADVFILAFSLISKASYE-----NIAKKWIPELRHYAPGVPIILVGTKL 119
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +ELK I + + + S NV + +AA L
Sbjct: 120 DLRDDKQFFIDHPGAVPITTNQGEELKKLIGSAVYIECSSKTQQNV-KAVFDAAIKVVLQ 178
Query: 387 IPETEFVRKRK 397
P+ + +K K
Sbjct: 179 PPKQKKKKKNK 189
>RAC9_ARATH (Q9XGU0) RAC-like GTP binding protein ARAC9 (GTPase
protein ROP8)
Length = 209
Score = 34.7 bits (78), Expect = 0.49
Identities = 19/65 (29%), Positives = 28/65 (42%), Gaps = 6/65 (9%)
Query: 193 SAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERY 252
SA++ TS S T +C G GK+ LL + F DY PT + +
Sbjct: 2 SASMAATSTSSA------TATTFIKCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNF 55
Query: 253 AANII 257
AN++
Sbjct: 56 NANVL 60
>RAC2_LOTJA (Q40220) RAC-like GTP binding protein RAC2
Length = 196
Score = 34.7 bits (78), Expect = 0.49
Identities = 44/189 (23%), Positives = 76/189 (39%), Gaps = 16/189 (8%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ +L + F DY PT + ++AN++ + G+ L L + E
Sbjct: 8 KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV--VDGSTVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAACDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDD 336
N L A DV S++ ++ +K + + P +L+ K D
Sbjct: 66 DYNRLRPLSYRGA-DVFLLAFSLLSRASYE---NISKKWIPELRHYAPTVPIVLVGTKLD 121
Query: 337 LTPFPRAVLD-------SVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLSI 387
L + ++D + +ELK I A + L S NV + +AA L
Sbjct: 122 LREDRQYLIDHPGATPITTAQGEELKKAIGAAVYLECSSKTQQNV-KAVFDAAIKVVLQP 180
Query: 388 PETEFVRKR 396
P+ + RK+
Sbjct: 181 PKPKKKRKK 189
>RC1C_DICDI (P34146) RAS-related protein rac1C
Length = 193
Score = 34.3 bits (77), Expect = 0.64
Identities = 14/41 (34%), Positives = 22/41 (53%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C V G GK+ LL + F +Y PT + Y+AN++
Sbjct: 5 KCVVVGDGAVGKTCLLISYTTNAFPGEYIPTVFDNYSANVM 45
>RC1B_DICDI (P34145) RAS-related protein rac1B
Length = 194
Score = 34.3 bits (77), Expect = 0.64
Identities = 14/41 (34%), Positives = 22/41 (53%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C V G GK+ LL + F +Y PT + Y+AN++
Sbjct: 5 KCVVVGDGAVGKTCLLISYTTNAFPGEYIPTVFDNYSANVM 45
>RC1A_DICDI (P34144) RAS-related protein rac1A
Length = 194
Score = 34.3 bits (77), Expect = 0.64
Identities = 14/41 (34%), Positives = 22/41 (53%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C V G GK+ LL + F +Y PT + Y+AN++
Sbjct: 5 KCVVVGDGAVGKTCLLISYTTNAFPGEYIPTVFDNYSANVM 45
>RACA_ARATH (O82481) RAC-like GTP binding protein ARAC10 (GTPase
protein ROP11)
Length = 215
Score = 34.3 bits (77), Expect = 0.64
Identities = 19/65 (29%), Positives = 29/65 (44%), Gaps = 2/65 (3%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ +L F DY PT + ++AN++ + GT L L + E
Sbjct: 10 KCVTVGDGAVGKTCMLICYTSNKFPTDYIPTVFDNFSANVV--VEGTTVNLGLWDTAGQE 67
Query: 277 VSNFL 281
N L
Sbjct: 68 DYNRL 72
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,697,229
Number of Sequences: 164201
Number of extensions: 2020775
Number of successful extensions: 5264
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 62
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 5198
Number of HSP's gapped (non-prelim): 89
length of query: 432
length of database: 59,974,054
effective HSP length: 113
effective length of query: 319
effective length of database: 41,419,341
effective search space: 13212769779
effective search space used: 13212769779
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC135317.6


