

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135317.3 + phase: 0
(701 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
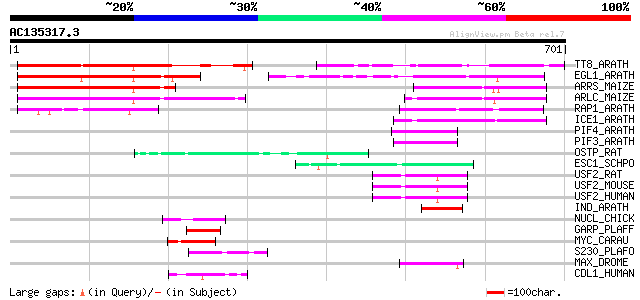
Score E
Sequences producing significant alignments: (bits) Value
TT8_ARATH (Q9FT81) TRANSPARENT TESTA 8 protein (Basic helix-loop... 294 5e-79
EGL1_ARATH (Q9CAD0) Transcription factor EGL1 (ENHANCER OF GLABR... 199 2e-50
ARRS_MAIZE (P13027) Anthocyanin regulatory R-S protein 177 1e-43
ARLC_MAIZE (P13526) Anthocyanin regulatory Lc protein 174 9e-43
RAP1_ARATH (Q39204) Transcription factor AtMYC2 (R-homologous Ar... 98 6e-20
ICE1_ARATH (Q9LSE2) Transcription factor ICE1 (Inducer of CBF ex... 83 2e-15
PIF4_ARATH (Q8W2F3) Phytochrome-interacting factor 4 (Basic heli... 62 7e-09
PIF3_ARATH (O80536) Phytochrome-interacting factor 3 (Phytochrom... 59 3e-08
OSTP_RAT (P08721) Osteopontin precursor (Bone sialoprotein 1) (S... 51 9e-06
ESC1_SCHPO (Q04635) Protein esc1 49 3e-05
USF2_RAT (Q63665) Upstream stimulatory factor 2 (Upstream transc... 47 1e-04
USF2_MOUSE (Q64705) Upstream stimulatory factor 2 (Upstream tran... 47 1e-04
USF2_HUMAN (Q15853) Upstream stimulatory factor 2 (Upstream tran... 47 1e-04
IND_ARATH (O81313) INDEHISCENT protein 47 1e-04
NUCL_CHICK (P15771) Nucleolin (Protein C23) 47 2e-04
GARP_PLAFF (P13816) Glutamic acid-rich protein precursor 47 2e-04
MYC_CARAU (P49709) Myc protein (c-myc) 46 3e-04
S230_PLAFO (Q08372) Transmission-blocking target antigen S230 pr... 46 4e-04
MAX_DROME (P91664) Max protein (dMax) 46 4e-04
CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2... 46 4e-04
>TT8_ARATH (Q9FT81) TRANSPARENT TESTA 8 protein (Basic
helix-loop-helix protein 42) (bHLH42) (AtbHLH042)
Length = 518
Score = 294 bits (753), Expect = 5e-79
Identities = 159/305 (52%), Positives = 196/305 (64%), Gaps = 69/305 (22%)
Query: 10 KLQNMLQAAVQSVQWTYSLFWQLCPQQLILVWGDGYYNGSIKTRKTVQPMEVSAEEASLQ 69
+LQ +L+ AVQSV WTYS+FWQ CPQQ +LVWG+GYYNG+IKTRKT QP EV+AEEA+L+
Sbjct: 19 ELQGLLKTAVQSVDWTYSVFWQFCPQQRVLVWGNGYYNGAIKTRKTTQPAEVTAEEAALE 78
Query: 70 RSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGVGLPGRAYT 129
RSQQLRELYE+L AGE+ R C +LSPEDLTE+EWFYLMCVSFSFPP G+PG+AY
Sbjct: 79 RSQQLRELYETLLAGESTSEA-RACTALSPEDLTETEWFYLMCVSFSFPPPSGMPGKAYA 137
Query: 130 KRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDGVVEFGTTDKVQEDLNFIKHV 184
+R+H+WL+GANEVDSK FSRAILAK TVVCIP+LDGVVE GTT KV+ED+ F++
Sbjct: 138 RRKHVWLSGANEVDSKTFSRAILAKSAKIQTVVCIPMLDGVVELGTTKKVREDVEFVELT 197
Query: 185 KSFFLDGHSLPPKPALSEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDEEEEEE 244
KSFF D PKPALSEHST E E
Sbjct: 198 KSFFYDHCKTNPKPALSEHST----------------------------------YEVHE 223
Query: 245 EDEEDDEVESESEDETGGRIRRATSMTAIVEPSELMQIEMPDDIRIGSPND---GSNHLD 301
E E+++EVE E + M +++R+GSP+D + +L
Sbjct: 224 EAEDEEEVEEE--------------------------MTMSEEMRLGSPDDEDVSNQNLH 257
Query: 302 SDFHL 306
SD H+
Sbjct: 258 SDLHI 262
Score = 192 bits (488), Expect = 3e-48
Identities = 126/315 (40%), Positives = 176/315 (55%), Gaps = 62/315 (19%)
Query: 388 NIHTPHVL--HHQLEDLTQEDTHYSQTVSTILQNQSTQWTIDSSPSINYITSSNQSSFTN 445
+I + H L H + +L +E +YSQTV+T+L + T DS + +YI QSSF
Sbjct: 261 HIESTHTLDTHMDMMNLMEEGGNYSQTVTTLLMSHPTSLLSDSVSTSSYI----QSSFAT 316
Query: 446 WTNHHNFHPLPPPETTTTTSQCLLKYILFTVPYLHTKNHDETSPQTHDAGVDPSSKLRGK 505
W + T +SQ +LK ++F VP+LH D K
Sbjct: 317 WRVENGKEH--QQVKTAPSSQWVLKQMIFRVPFLHDNTKD-------------------K 355
Query: 506 GTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQ 565
P+++LS HV+AERRRREKLNE+FI LRS+VPFVTKMDK SILGDTI Y+ LR+++
Sbjct: 356 RLPREDLS--HVVAERRRREKLNEKFITLRSMVPFVTKMDKVSILGDTIAYVNHLRKRVH 413
Query: 566 DLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSVQVSIIES 625
+LE + EQQ + R + + + V+VSIIE+
Sbjct: 414 ELENTHH----EQQHK------------------------RTRTCKRKTSEEVEVSIIEN 445
Query: 626 DALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKVKENGGNGKKVS 685
D LLE+ C +R+GLLLD++ +L EL IE V +S+N+ F AE+RAKV+ GKK S
Sbjct: 446 DVLLEMRCEYRDGLLLDILQVLHELGIETTAVHTSVNDHDFEAEIRAKVR-----GKKAS 500
Query: 686 IVEVKRALNQIIPHN 700
I EVKRA++Q+I H+
Sbjct: 501 IAEVKRAIHQVIIHD 515
>EGL1_ARATH (Q9CAD0) Transcription factor EGL1 (ENHANCER OF GLABRA3)
(Basic helix-loop-helix protein 2) (bHLH2) (AtbHLH002)
(AtMyc-146)
Length = 596
Score = 199 bits (506), Expect = 2e-50
Identities = 109/246 (44%), Positives = 154/246 (62%), Gaps = 20/246 (8%)
Query: 11 LQNMLQAAVQSVQWTYSLFWQLCPQQL-ILVWGDGYYNGSIKTRKTVQPMEVSAEEASLQ 69
L+ L +V+++QW+Y +FW + Q +L WGDGYYNG IKTRKT+Q EV ++ L+
Sbjct: 13 LKKQLAVSVRNIQWSYGIFWSVSASQPGVLEWGDGYYNGDIKTRKTIQAAEVKIDQLGLE 72
Query: 70 RSQQLRELYESLSAGETNPP-----TRRP-CASLSPEDLTESEWFYLMCVSFSFPPGVGL 123
RS+QLRELYESLS E++ TRR A+LSPEDLT++EW+YL+C+SF F G G+
Sbjct: 73 RSEQLRELYESLSLAESSASGSSQVTRRASAAALSPEDLTDTEWYYLVCMSFVFNIGEGI 132
Query: 124 PGRAYTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDGVVEFGTTDKVQEDL 178
PG A + + IWL A DSK+F+R++LAK TVVC P L GV+E GTT+ ++ED+
Sbjct: 133 PGGALSNGEPIWLCNAETADSKVFTRSLLAKSASLQTVVCFPFLGGVLEIGTTEHIKEDM 192
Query: 179 NFIKHVKSFFLDGHSLPPKPALSEHS----TSNPTSSTDHIPTIMYTMADPPSTNPNQDD 234
N I+ VK+ FL+ PP +S S +P S + P + T A P ++ +
Sbjct: 193 NVIQSVKTLFLEA---PPYTTISTRSDYQEIFDPLSDDKYTP-VFITEAFPTTSTSGFEQ 248
Query: 235 MDEDEE 240
ED +
Sbjct: 249 EPEDHD 254
Score = 131 bits (330), Expect = 5e-30
Identities = 102/361 (28%), Positives = 169/361 (46%), Gaps = 64/361 (17%)
Query: 328 QRWGPIEEPVGNLLQVQLPSSDPYIYMWSILINSIVYPIFAGSTSMQNRTQFNAQPPKSK 387
Q W + E + N + L SSD V F G+T + P KS+
Sbjct: 266 QSWQFVGEEISNCIHQSLNSSD------------CVSQTFVGTTG-----RLACDPRKSR 308
Query: 388 NIHTPHVLHHQLEDLTQEDTHYSQTVSTILQNQSTQWTIDSSPSINYITSSNQSSFTNWT 447
+ +D HY +STI + +T I N+ +SSFT W
Sbjct: 309 IQRLGQIQEQSNHVNMDDDVHYQGVISTIFK--TTHQLILGPQFQNF---DKRSSFTRWK 363
Query: 448 NHHNFHPLPPPETTTTTSQCLLKYILFTVPYLHTKNHDETSPQTHDAGVDPSSKLRGKGT 507
+ +T SQ ++K ILF VP ++ K +E P T
Sbjct: 364 RSSSV------KTLGEKSQKMIKKILFEVPLMNKK--EELLPDT---------------- 399
Query: 508 PQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDL 567
E + NH L+E++RREKLNERF+ LRS++P ++K+DK SIL DTIEYL+ L++++Q+L
Sbjct: 400 --PEETGNHALSEKKRREKLNERFMTLRSIIPSISKIDKVSILDDTIEYLQDLQKRVQEL 457
Query: 568 ETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEV------------- 614
E+ +TE +R + P D+++ R C ++ K + V
Sbjct: 458 ESCRESADTE--TRITMMKRKKPDDEEE-RASANCMNSKRKGSDVNVGEDEPADIGYAGL 514
Query: 615 VSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKV 674
++++S + ++ ++E+ C REG+LL++M ++ +L ++ VQSS +G+ + K
Sbjct: 515 TDNLRISSLGNEVVIELRCAWREGILLEIMDVISDLNLDSHSVQSSTGDGLLCLTVNCKH 574
Query: 675 K 675
K
Sbjct: 575 K 575
>ARRS_MAIZE (P13027) Anthocyanin regulatory R-S protein
Length = 612
Score = 177 bits (448), Expect = 1e-43
Identities = 87/207 (42%), Positives = 135/207 (65%), Gaps = 12/207 (5%)
Query: 11 LQNMLQAAVQSVQWTYSLFWQLCPQQL-ILVWGDGYYNGSIKTRKTVQPMEVSAEEASLQ 69
+++ L AA +S+ W+Y+LFW + Q +L W DG+YNG +KTRK +E+++++ +Q
Sbjct: 24 MRSQLAAAARSINWSYALFWSISDTQPGVLTWTDGFYNGEVKTRKISNSVELTSDQLVMQ 83
Query: 70 RSQQLRELYESLSAGETN--PPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGVGLPGRA 127
RS QLRELYE+L +GE + RP SLSPEDL ++EW+Y++ ++++F PG GLPGR+
Sbjct: 84 RSDQLRELYEALLSGEGDRRAAPARPAGSLSPEDLGDTEWYYVVSMTYAFRPGQGLPGRS 143
Query: 128 YTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDGVVEFGTTDKVQEDLNFIK 182
+ +H+WL A+ SK F RA+LAK +++CIPV+ GV+E GTTD V E + +
Sbjct: 144 FASDEHVWLCNAHLAGSKAFPRALLAKSASIQSILCIPVMGGVLELGTTDTVPEAPDLVS 203
Query: 183 HVKSFFLDGHSLPPKPALSEHSTSNPT 209
+ F + P P SE +S+P+
Sbjct: 204 RATAAFWE----PQCPTYSEEPSSSPS 226
Score = 104 bits (260), Expect = 7e-22
Identities = 63/184 (34%), Positives = 109/184 (59%), Gaps = 17/184 (9%)
Query: 511 ELSA--NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLE 568
E+SA NHV++ER+RREKLNE F++L+SL+P + +++KASIL +TI YLK+L+R++Q+LE
Sbjct: 412 EMSATKNHVMSERKRREKLNEMFLVLKSLLPSIHRVNKASILAETIAYLKELQRRVQELE 471
Query: 569 TRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKA-------VETEVV------ 615
+ ++ + + + + VR E C ++ K+ VE V
Sbjct: 472 SSREPASRPSETTTRLITRPSRGNNESVR-KEVCAGSKRKSPELGRDDVERPPVLTMDAG 530
Query: 616 -SSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKV 674
S+V V++ + D LLE++C E L+ V ++ L ++V+ VQ+S +G ++RA+
Sbjct: 531 SSNVTVTVSDKDVLLEVQCRWEELLMTRVFDAIKSLHLDVLSVQASAPDGFMGLKIRAQF 590
Query: 675 KENG 678
+G
Sbjct: 591 AGSG 594
>ARLC_MAIZE (P13526) Anthocyanin regulatory Lc protein
Length = 610
Score = 174 bits (440), Expect = 9e-43
Identities = 102/298 (34%), Positives = 167/298 (55%), Gaps = 16/298 (5%)
Query: 11 LQNMLQAAVQSVQWTYSLFWQLCPQQL-ILVWGDGYYNGSIKTRKTVQPMEVSAEEASLQ 69
+++ L AA +S+ W+Y+LFW + Q +L W DG+YNG +KTRK +E+++++ +Q
Sbjct: 24 MRSQLAAAARSINWSYALFWSISDTQPGVLTWTDGFYNGEVKTRKISNSVELTSDQLVMQ 83
Query: 70 RSQQLRELYESLSAGETN--PPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGVGLPGRA 127
RS QLRELYE+L +GE + RP SLSPEDL ++EW+Y++ ++++F PG GLPGR+
Sbjct: 84 RSDQLRELYEALLSGEGDRRAAPARPAGSLSPEDLGDTEWYYVVSMTYAFRPGQGLPGRS 143
Query: 128 YTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDGVVEFGTTDKVQEDLNFIK 182
+ +H+WL A+ SK F RA+LAK +++CIPV+ GV+E GTTD V E + +
Sbjct: 144 FASDEHVWLCNAHLAGSKAFPRALLAKSASIQSILCIPVMGGVLELGTTDTVPEAPDLVS 203
Query: 183 HVKSFFLDGHSLPPKPALSEHSTSNPT-SSTDHIPTIMYTMADPPSTNPNQDDMDEDEEE 241
+ F + P P+ S +N T + T + D + + + M
Sbjct: 204 RATAAFWE----PQCPSSSPSGRANETGEAAADDGTFAFEELDHNNGMDDIEAMTAAGGH 259
Query: 242 EEEEDEEDDEVESESEDETGGRI-RRATSMTAIVEPSELMQIEMPDDIRIGSPNDGSN 298
+EE+ E E+ S+D + I + ++ + +L + +P + G D SN
Sbjct: 260 GQEEELRLREAEALSDDASLEHITKEIEEFYSLCDEMDLQALPLP--LEDGWTVDASN 315
Score = 104 bits (259), Expect = 9e-22
Identities = 61/194 (31%), Positives = 109/194 (55%), Gaps = 21/194 (10%)
Query: 499 SSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLK 558
+ ++ G GT NHV++ER+RREKLNE F++L+SL+P + +++KASIL +TI YLK
Sbjct: 406 AQEMSGTGTK------NHVMSERKRREKLNEMFLVLKSLLPSIHRVNKASILAETIAYLK 459
Query: 559 QLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVE------- 611
+L+R++Q+LE+ ++ + + + + VR E C ++ K+ E
Sbjct: 460 ELQRRVQELESSREPASRPSETTTRLITRPSRGNNESVR-KEVCAGSKRKSPELGRDDVE 518
Query: 612 -------TEVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNG 664
S+V V++ + D LLE++C E L+ V ++ L ++V+ VQ+S +G
Sbjct: 519 RPPVLTMDAGTSNVTVTVSDKDVLLEVQCRWEELLMTRVFDAIKSLHLDVLSVQASAPDG 578
Query: 665 VFVAELRAKVKENG 678
++RA+ +G
Sbjct: 579 FMGLKIRAQFAGSG 592
>RAP1_ARATH (Q39204) Transcription factor AtMYC2 (R-homologous
Arabidopsis protein-1) (RAP-1) (Basic helix-loop-helix
protein 6) (bHLH6) (AtbHLH006) (rd22BP1)
Length = 623
Score = 98.2 bits (243), Expect = 6e-20
Identities = 60/193 (31%), Positives = 98/193 (50%), Gaps = 23/193 (11%)
Query: 11 LQNMLQAAVQSVQ--WTYSLFWQLCPQ---QLILVWGDGYYNG-----SIKTRKTVQPME 60
LQ LQA ++ WTY++FWQ +L WGDGYY G + + R + P
Sbjct: 68 LQQRLQALIEGTHEGWTYAIFWQPSYDFSGASVLGWGDGYYKGEEDKANPRRRSSSPPFS 127
Query: 61 VSAEEASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPG 120
A++ R + LREL +S G P E++T++EWF+L+ ++ SF G
Sbjct: 128 TPADQE--YRKKVLRELNSLISGGVA------PSDDAVDEEVTDTEWFFLVSMTQSFACG 179
Query: 121 VGLPGRAYTKRQHIWLTGANEVDSKIFSRA-----ILAKTVVCIPVLDGVVEFGTTDKVQ 175
GL G+A+ +W++G++++ RA T+ CIP +GVVE G+T+ ++
Sbjct: 180 AGLAGKAFATGNAVWVSGSDQLSGSGCERAKQGGVFGMHTIACIPSANGVVEVGSTEPIR 239
Query: 176 EDLNFIKHVKSFF 188
+ + I V+ F
Sbjct: 240 QSSDLINKVRILF 252
Score = 86.3 bits (212), Expect = 3e-16
Identities = 58/184 (31%), Positives = 95/184 (51%), Gaps = 10/184 (5%)
Query: 493 DAGVDPSSKLRGKGTPQD-ELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILG 551
+ V+ K RG+ E NHV AER+RREKLN+RF LR++VP V+KMDKAS+LG
Sbjct: 429 EVAVEKRPKKRGRKPANGREEPLNHVEAERQRREKLNQRFYALRAVVPNVSKMDKASLLG 488
Query: 552 DTIEYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVG-PTDKKKVRIVEECGATRAKAV 610
D I Y+ +L+ K+ ++T + +++ + Q L G + C + + +
Sbjct: 489 DAIAYINELKSKV--VKTESEKLQIKNQLEEVKLELAGRKASASGGDMSSSCSSIKPVGM 546
Query: 611 ETEVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAEL 670
E ++V II DA++ +E R +M L +L +EV S+ N + + +
Sbjct: 547 E------IEVKIIGWDAMIRVESSKRNHPAARLMSALMDLELEVNHASMSVVNDLMIQQA 600
Query: 671 RAKV 674
K+
Sbjct: 601 TVKM 604
>ICE1_ARATH (Q9LSE2) Transcription factor ICE1 (Inducer of CBF
expression 1) (Basic helix-loop-helix protein 116)
(bHLH116) (AtbHLH116)
Length = 494
Score = 83.2 bits (204), Expect = 2e-15
Identities = 64/197 (32%), Positives = 101/197 (50%), Gaps = 9/197 (4%)
Query: 485 DETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKM 544
+E+ + K + KG P A +++AERRRR+KLN+R +LRS+VP ++KM
Sbjct: 282 NESGKAAESVQIGGGGKGKKKGMP-----AKNLMAERRRRKKLNDRLYMLRSVVPKISKM 336
Query: 545 DKASILGDTIEYLKQLRRKIQDL--ETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEEC 602
D+ASILGD I+YLK+L ++I DL E + + + S L R+ EE
Sbjct: 337 DRASILGDAIDYLKELLQRINDLHNELESTPPGSLPPTSSSFHPLTPTPQTLSCRVKEEL 396
Query: 603 GATRAKAVETEVVSSVQVSIIESDAL-LEIECLHREGLLLDVMVMLRELRIEVIGVQSSL 661
+ + + + + V+V + E A+ + + C R GLLL M L L ++V S
Sbjct: 397 CPSSLPSPKGQ-QARVEVRLREGRAVNIHMFCGRRPGLLLATMKALDNLGLDVQQAVISC 455
Query: 662 NNGVFVAELRAKVKENG 678
NG + RA+ + G
Sbjct: 456 FNGFALDVFRAEQCQEG 472
>PIF4_ARATH (Q8W2F3) Phytochrome-interacting factor 4 (Basic
helix-loop-helix protein 9) (bHLH9) (AtbHLH009) (Short
under red-light 2)
Length = 430
Score = 61.6 bits (148), Expect = 7e-09
Identities = 33/83 (39%), Positives = 49/83 (58%), Gaps = 1/83 (1%)
Query: 483 NHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVT 542
NH + S DA + S R + + H L+ERRRR+++NER L+ L+P +
Sbjct: 230 NHTDESVSLSDA-IGNKSNQRSGSNRRSRAAEVHNLSERRRRDRINERMKALQELIPHCS 288
Query: 543 KMDKASILGDTIEYLKQLRRKIQ 565
K DKASIL + I+YLK L+ ++Q
Sbjct: 289 KTDKASILDEAIDYLKSLQLQLQ 311
>PIF3_ARATH (O80536) Phytochrome-interacting factor 3
(Phytochrome-associated protein 3) (Basic
helix-loop-helix protein 8) (bHLH8) (AtbHLH008)
Length = 524
Score = 59.3 bits (142), Expect = 3e-08
Identities = 28/81 (34%), Positives = 49/81 (59%)
Query: 485 DETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKM 544
++ ++ D + G G+ + + H L+ERRRR+++NE+ L+ L+P K+
Sbjct: 317 EDVEEESGDGRKEAGPSRTGLGSKRSRSAEVHNLSERRRRDRINEKMRALQELIPNCNKV 376
Query: 545 DKASILGDTIEYLKQLRRKIQ 565
DKAS+L + IEYLK L+ ++Q
Sbjct: 377 DKASMLDEAIEYLKSLQLQVQ 397
>OSTP_RAT (P08721) Osteopontin precursor (Bone sialoprotein 1)
(Secreted phosphoprotein 1) (SPP-1)
Length = 317
Score = 51.2 bits (121), Expect = 9e-06
Identities = 71/303 (23%), Positives = 119/303 (38%), Gaps = 43/303 (14%)
Query: 158 CIPVLDGVVEFGTTDKVQEDLNFIKHVKSFFLDGHSLPPKPALSEH--STSNPTSSTDHI 215
C+PV V EFG+++ E ++ KH + L P P+ ++ + N SS +
Sbjct: 16 CLPVK--VAEFGSSE---EKAHYSKHSDAV---ATWLKPDPSQKQNLLAPQNSVSSEETD 67
Query: 216 PTIMYTMADPPSTNPNQDDMDEDEEEEEEED--EEDDEVESESEDETGGRIRRATSMTAI 273
T+ P ++N + D MD+D++++++ D E +D V S+ DE+ S TA
Sbjct: 68 DFKQETL--PSNSNESHDHMDDDDDDDDDGDHAESEDSVNSDESDESHHSDESDESFTAS 125
Query: 274 VEPSELMQIEMPDDIRIGSPNDGSNHLDSDFHLLVVSNQGNPLGHVDSYKTDLTQRWGPI 333
+ L I D+ G + + L S VS++ P D+ DLT R
Sbjct: 126 TQADVLTPIAPTVDVPDGRGDSLAYGLRSKSRSFPVSDEQYP----DATDEDLTSR---- 177
Query: 334 EEPVGNLLQVQLPSSDPYIYMWSILINSIVYPIFAGSTSMQNRTQFNAQPPKSKNIHTPH 393
++ SD I V P+ + ++ +S + P
Sbjct: 178 ---------MKSQESDEAIK---------VIPVAQRLSVPSDQDSNGKTSHESSQLDEPS 219
Query: 394 VLHHQLE---DLTQEDTHYSQTVSTILQNQSTQWTIDSSPSINYITSSNQSSFTNWTNHH 450
V H LE + Q +H S S + + IDS+ + I S S + H
Sbjct: 220 VETHSLEQSKEYKQRASHESTEQSDAIDSAEKPDAIDSAERSDAIDSQASSKASLEHQSH 279
Query: 451 NFH 453
FH
Sbjct: 280 EFH 282
>ESC1_SCHPO (Q04635) Protein esc1
Length = 413
Score = 49.3 bits (116), Expect = 3e-05
Identities = 58/234 (24%), Positives = 94/234 (39%), Gaps = 17/234 (7%)
Query: 361 SIVYPIFAGSTSMQNRTQFNAQPPKSKN------IHTPH-VLHHQLEDLTQEDTHYSQTV 413
S+ P + +T + N+ Q SKN +H+ H Q + Q+ T +S
Sbjct: 185 SVTSPASSSATPLPNQPS-QQQFLVSKNDAFTTFVHSVHNTPMQQSMYVPQQQTSHSSGA 243
Query: 414 STILQNQSTQWTIDSSPSINYITSSNQSSFTNWTNHHNFHPLPPPETTTTTSQCLLKYIL 473
S QN+S + S +Y S + + P+ P + + I
Sbjct: 244 S--YQNESANPPVQSPMQYSYSQGQPFSYPQHKNQSFSASPIDPSMSYVYRAPESFSSIN 301
Query: 474 FTVPYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFII 533
VPY + + V + G T EL +H LAER+RR+++ E F
Sbjct: 302 ANVPYGRNEYLRRVTSL-----VPNQPEYTGPYTRNPELRTSHKLAERKRRKEIKELFDD 356
Query: 534 LRSLVPF--VTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQSRSGVT 585
L+ +P TK K +L I+Y++QL+ + LE + +E QS VT
Sbjct: 357 LKDALPLDKSTKSSKWGLLTRAIQYIEQLKSEQVALEAYVKSLEENMQSNKEVT 410
>USF2_RAT (Q63665) Upstream stimulatory factor 2 (Upstream
transcription factor 2) (Major late transcription factor
2) (Fragment)
Length = 291
Score = 47.4 bits (111), Expect = 1e-04
Identities = 37/129 (28%), Positives = 60/129 (45%), Gaps = 13/129 (10%)
Query: 459 ETTTTTSQCLLKYILFTVPY--LHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELS-AN 515
+TT + Q ++ + P L T +P+TH S K+ G TP+DE A
Sbjct: 129 QTTDQSLQAGGQFYVMMTPQDVLQTGTQRTIAPRTHPY----SPKIDGTRTPRDERRRAQ 184
Query: 516 HVLAERRRREKLNERFIILRSLVP------FVTKMDKASILGDTIEYLKQLRRKIQDLET 569
H ERRRR+K+N + L ++P T K IL +Y+++LR+ Q ++
Sbjct: 185 HNEVERRRRDKINNWIVQLSKIIPDCHADNSKTGASKGGILSKACDYIRELRQTNQRMQE 244
Query: 570 RNRQIETEQ 578
++ E Q
Sbjct: 245 TFKEAERLQ 253
>USF2_MOUSE (Q64705) Upstream stimulatory factor 2 (Upstream
transcription factor 2) (Major late transcription factor
2)
Length = 346
Score = 47.4 bits (111), Expect = 1e-04
Identities = 37/129 (28%), Positives = 60/129 (45%), Gaps = 13/129 (10%)
Query: 459 ETTTTTSQCLLKYILFTVPY--LHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELS-AN 515
+TT + Q ++ + P L T +P+TH S K+ G TP+DE A
Sbjct: 184 QTTDQSLQAGGQFYVMMTPQDVLQTGTQRTIAPRTHPY----SPKIDGTRTPRDERRRAQ 239
Query: 516 HVLAERRRREKLNERFIILRSLVP------FVTKMDKASILGDTIEYLKQLRRKIQDLET 569
H ERRRR+K+N + L ++P T K IL +Y+++LR+ Q ++
Sbjct: 240 HNEVERRRRDKINNWIVQLSKIIPDCHADNSKTGASKGGILSKACDYIRELRQTNQRMQE 299
Query: 570 RNRQIETEQ 578
++ E Q
Sbjct: 300 TFKEAERLQ 308
>USF2_HUMAN (Q15853) Upstream stimulatory factor 2 (Upstream
transcription factor 2) (FOS-interacting protein) (FIP)
(Major late transcription factor 2)
Length = 346
Score = 47.4 bits (111), Expect = 1e-04
Identities = 37/129 (28%), Positives = 60/129 (45%), Gaps = 13/129 (10%)
Query: 459 ETTTTTSQCLLKYILFTVPY--LHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELS-AN 515
+TT + Q ++ + P L T +P+TH S K+ G TP+DE A
Sbjct: 184 QTTDQSLQAGGQFYVMMTPQDVLQTGTQRTIAPRTHPY----SPKIDGTRTPRDERRRAQ 239
Query: 516 HVLAERRRREKLNERFIILRSLVP------FVTKMDKASILGDTIEYLKQLRRKIQDLET 569
H ERRRR+K+N + L ++P T K IL +Y+++LR+ Q ++
Sbjct: 240 HNEVERRRRDKINNWIVQLSKIIPDCNADNSKTGASKGGILSKACDYIRELRQTNQRMQE 299
Query: 570 RNRQIETEQ 578
++ E Q
Sbjct: 300 TFKEAERLQ 308
>IND_ARATH (O81313) INDEHISCENT protein
Length = 169
Score = 47.4 bits (111), Expect = 1e-04
Identities = 22/52 (42%), Positives = 37/52 (70%)
Query: 521 RRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNR 572
RRRRE+++E+ IL+ +VP KMD AS+L + I Y K L+R+++ L+ ++
Sbjct: 99 RRRRERISEKIRILKRIVPGGAKMDTASMLDEAIRYTKFLKRQVRILQPHSQ 150
>NUCL_CHICK (P15771) Nucleolin (Protein C23)
Length = 694
Score = 46.6 bits (109), Expect = 2e-04
Identities = 25/80 (31%), Positives = 40/80 (49%), Gaps = 13/80 (16%)
Query: 193 SLPPKPALSEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDEEEEEEEDEEDDEV 252
++P KPA + + + P +P Q+ +E+EE++EEEDEEDDE
Sbjct: 143 AVPVKPAAKKSAAAVPAKKPAVVPA-------------KQESEEEEEEDDEEEDEEDDES 189
Query: 253 ESESEDETGGRIRRATSMTA 272
E E+ D T +++ T A
Sbjct: 190 EDEAMDTTPAPVKKPTPAKA 209
Score = 42.0 bits (97), Expect = 0.006
Identities = 17/26 (65%), Positives = 23/26 (88%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESE 257
+D+ DEDE+EE+E+DEE+DE ESE E
Sbjct: 222 EDEEDEDEDEEDEDDEEEDEEESEDE 247
Score = 40.0 bits (92), Expect = 0.021
Identities = 15/47 (31%), Positives = 32/47 (67%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDETGGRIRRATSMTAIVEPSE 278
++ +EDE++E++E++ED+E ES+ E+E ++ A +A P++
Sbjct: 114 EESEEEDEDDEDDEEDEDEEEESDEEEEPAVPVKPAAKKSAAAVPAK 160
Score = 40.0 bits (92), Expect = 0.021
Identities = 16/45 (35%), Positives = 27/45 (59%)
Query: 223 ADPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDETGGRIRRA 267
A P +D +++E+E+E+E++EDDE E E E E ++ A
Sbjct: 209 ATPAKAKAESEDEEDEEDEDEDEEDEDDEEEDEEESEDEKPVKEA 253
Score = 38.5 bits (88), Expect = 0.061
Identities = 18/59 (30%), Positives = 30/59 (50%)
Query: 202 EHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDET 260
+ + + T P T A + DE++EE+E+EDEED++ E E E+E+
Sbjct: 186 DDESEDEAMDTTPAPVKKPTPAKATPAKAKAESEDEEDEEDEDEDEEDEDDEEEDEEES 244
Score = 37.4 bits (85), Expect = 0.14
Identities = 18/59 (30%), Positives = 31/59 (52%)
Query: 201 SEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDE 259
SE + T + PT ++ +EDE+E+EE++++++E E ESEDE
Sbjct: 189 SEDEAMDTTPAPVKKPTPAKATPAKAKAESEDEEDEEDEDEDEEDEDDEEEDEEESEDE 247
Score = 35.4 bits (80), Expect = 0.52
Identities = 15/40 (37%), Positives = 26/40 (64%)
Query: 233 DDMDEDEEEEEEEDEEDDEVESESEDETGGRIRRATSMTA 272
+D DE++E++EEEDEE+ E E ++ G R + + +A
Sbjct: 227 EDEDEEDEDDEEEDEEESEDEKPVKEAPGKRKKEMANKSA 266
Score = 34.3 bits (77), Expect = 1.2
Identities = 13/30 (43%), Positives = 22/30 (73%)
Query: 231 NQDDMDEDEEEEEEEDEEDDEVESESEDET 260
N + ++E EEE+ED+EDDE + + E+E+
Sbjct: 107 NGKNAKKEESEEEDEDDEDDEEDEDEEEES 136
Score = 33.5 bits (75), Expect = 2.0
Identities = 13/29 (44%), Positives = 20/29 (68%)
Query: 231 NQDDMDEDEEEEEEEDEEDDEVESESEDE 259
N + +EE+E++ED+E+DE E E DE
Sbjct: 110 NAKKEESEEEDEDDEDDEEDEDEEEESDE 138
>GARP_PLAFF (P13816) Glutamic acid-rich protein precursor
Length = 678
Score = 46.6 bits (109), Expect = 2e-04
Identities = 19/43 (44%), Positives = 31/43 (71%)
Query: 224 DPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDETGGRIRR 266
D + +DD +ED++E+E+EDEED+E E E E+E+ +I+R
Sbjct: 628 DAEEDDDEEDDDEEDDDEDEDEDEEDEEEEEEEEEESEKKIKR 670
Score = 42.0 bits (97), Expect = 0.006
Identities = 16/29 (55%), Positives = 26/29 (89%)
Query: 231 NQDDMDEDEEEEEEEDEEDDEVESESEDE 259
++++++EDEEEEEEE+EE++E E E E+E
Sbjct: 568 DEEEVEEDEEEEEEEEEEEEEEEEEEEEE 596
Score = 41.6 bits (96), Expect = 0.007
Identities = 17/43 (39%), Positives = 30/43 (69%)
Query: 224 DPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDETGGRIRR 266
D + +++D DEDE+E+EE++EE++E E ESE + +R+
Sbjct: 632 DDDEEDDDEEDDDEDEDEDEEDEEEEEEEEEESEKKIKRNLRK 674
Score = 40.4 bits (93), Expect = 0.016
Identities = 16/36 (44%), Positives = 26/36 (71%)
Query: 224 DPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDE 259
D ++++ +E+EEEEEEE+EE++E E E E+E
Sbjct: 568 DEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEE 603
Score = 40.4 bits (93), Expect = 0.016
Identities = 16/28 (57%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EE++E E E EDE
Sbjct: 580 EEEEEEEEEEEEEEEEEEEEEEEEEEDE 607
Score = 40.4 bits (93), Expect = 0.016
Identities = 16/28 (57%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EE++E E E EDE
Sbjct: 582 EEEEEEEEEEEEEEEEEEEEEEEEDEDE 609
Score = 40.0 bits (92), Expect = 0.021
Identities = 14/36 (38%), Positives = 26/36 (71%)
Query: 224 DPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDE 259
D ++DD +ED++EE++++E+DDE E E E++
Sbjct: 618 DEDDAEEDEDDAEEDDDEEDDDEEDDDEDEDEDEED 653
Score = 40.0 bits (92), Expect = 0.021
Identities = 17/37 (45%), Positives = 26/37 (69%), Gaps = 1/37 (2%)
Query: 224 DPPSTNPNQDDMDEDEEE-EEEEDEEDDEVESESEDE 259
D ++DD +EDE++ EE++DEEDD+ E + EDE
Sbjct: 611 DEDDAEEDEDDAEEDEDDAEEDDDEEDDDEEDDDEDE 647
Score = 38.9 bits (89), Expect = 0.047
Identities = 15/28 (53%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EE++E E E E+E
Sbjct: 578 EEEEEEEEEEEEEEEEEEEEEEEEEEEE 605
Score = 38.9 bits (89), Expect = 0.047
Identities = 15/28 (53%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EE++E E E E+E
Sbjct: 577 EEEEEEEEEEEEEEEEEEEEEEEEEEEE 604
Score = 38.5 bits (88), Expect = 0.061
Identities = 16/28 (57%), Positives = 23/28 (82%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
Q+D +E EE+EEEE+EE++E E E E+E
Sbjct: 566 QEDEEEVEEDEEEEEEEEEEEEEEEEEE 593
Score = 38.1 bits (87), Expect = 0.080
Identities = 14/28 (50%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EE++E + + EDE
Sbjct: 585 EEEEEEEEEEEEEEEEEEEEEDEDEEDE 612
Score = 37.7 bits (86), Expect = 0.10
Identities = 14/37 (37%), Positives = 26/37 (69%)
Query: 223 ADPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDE 259
A+ + +D+ D +E+++EE+D+E+D+ E E EDE
Sbjct: 615 AEEDEDDAEEDEDDAEEDDDEEDDDEEDDDEDEDEDE 651
Score = 37.7 bits (86), Expect = 0.10
Identities = 14/28 (50%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EE++E E E E++
Sbjct: 579 EEEEEEEEEEEEEEEEEEEEEEEEEEED 606
Score = 37.7 bits (86), Expect = 0.10
Identities = 14/28 (50%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EE++E E E E++
Sbjct: 584 EEEEEEEEEEEEEEEEEEEEEEDEDEED 611
Score = 37.4 bits (85), Expect = 0.14
Identities = 14/28 (50%), Positives = 23/28 (82%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEE+EDEED++ E ED+
Sbjct: 594 EEEEEEEEEEEEDEDEEDEDDAEEDEDD 621
Score = 37.4 bits (85), Expect = 0.14
Identities = 14/27 (51%), Positives = 23/27 (84%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESED 258
+++ +E+EEEEEEE+EE+DE E + +D
Sbjct: 588 EEEEEEEEEEEEEEEEEEDEDEEDEDD 614
Score = 37.0 bits (84), Expect = 0.18
Identities = 13/29 (44%), Positives = 24/29 (81%)
Query: 231 NQDDMDEDEEEEEEEDEEDDEVESESEDE 259
++D+ DED+ EE+E+D E+DE ++E +D+
Sbjct: 606 DEDEEDEDDAEEDEDDAEEDEDDAEEDDD 634
Score = 37.0 bits (84), Expect = 0.18
Identities = 13/28 (46%), Positives = 25/28 (88%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE++ED+E E ++E++
Sbjct: 591 EEEEEEEEEEEEEEEDEDEEDEDDAEED 618
Score = 37.0 bits (84), Expect = 0.18
Identities = 15/33 (45%), Positives = 24/33 (72%)
Query: 227 STNPNQDDMDEDEEEEEEEDEEDDEVESESEDE 259
S +D+ + +E+EEEEE+EE++E E E E+E
Sbjct: 562 SKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEE 594
Score = 37.0 bits (84), Expect = 0.18
Identities = 13/28 (46%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EE+++ + E ED+
Sbjct: 587 EEEEEEEEEEEEEEEEEEEDEDEEDEDD 614
Score = 36.6 bits (83), Expect = 0.23
Identities = 14/28 (50%), Positives = 23/28 (82%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EED++ E E + E
Sbjct: 589 EEEEEEEEEEEEEEEEEDEDEEDEDDAE 616
Score = 36.2 bits (82), Expect = 0.30
Identities = 13/28 (46%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+EE++E E E +++
Sbjct: 586 EEEEEEEEEEEEEEEEEEEEDEDEEDED 613
Score = 36.2 bits (82), Expect = 0.30
Identities = 13/28 (46%), Positives = 23/28 (81%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEEDE++++ + EDE
Sbjct: 592 EEEEEEEEEEEEEEDEDEEDEDDAEEDE 619
Score = 34.7 bits (78), Expect = 0.88
Identities = 11/28 (39%), Positives = 25/28 (89%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEE+++E+DE ++E +++
Sbjct: 593 EEEEEEEEEEEEEDEDEEDEDDAEEDED 620
Score = 34.7 bits (78), Expect = 0.88
Identities = 12/28 (42%), Positives = 23/28 (81%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE+E++DE + + +E
Sbjct: 590 EEEEEEEEEEEEEEEEDEDEEDEDDAEE 617
Score = 34.3 bits (77), Expect = 1.2
Identities = 11/28 (39%), Positives = 24/28 (85%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +EDE+EE+E+D E+DE ++E +++
Sbjct: 600 EEEEEEDEDEEDEDDAEEDEDDAEEDED 627
Score = 33.9 bits (76), Expect = 1.5
Identities = 12/36 (33%), Positives = 24/36 (66%)
Query: 224 DPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDE 259
D + ++D D+ EE+E++ +E+DDE + + ED+
Sbjct: 608 DEEDEDDAEEDEDDAEEDEDDAEEDDDEEDDDEEDD 643
Score = 33.5 bits (75), Expect = 2.0
Identities = 12/28 (42%), Positives = 22/28 (77%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEE+E+E++EDD E E + E
Sbjct: 596 EEEEEEEEEEDEDEEDEDDAEEDEDDAE 623
Score = 33.5 bits (75), Expect = 2.0
Identities = 10/27 (37%), Positives = 22/27 (81%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESED 258
+++ DEDEE+E++ +E++D+ E + +D
Sbjct: 602 EEEEDEDEEDEDDAEEDEDDAEEDEDD 628
Score = 32.3 bits (72), Expect = 4.4
Identities = 11/28 (39%), Positives = 22/28 (78%)
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EE+E+EEDE+D E + + +E
Sbjct: 597 EEEEEEEEEDEDEEDEDDAEEDEDDAEE 624
Score = 31.6 bits (70), Expect = 7.5
Identities = 15/30 (50%), Positives = 22/30 (73%), Gaps = 2/30 (6%)
Query: 232 QDDMDEDEEEEEE--EDEEDDEVESESEDE 259
Q++ E +E+EEE EDEE++E E E E+E
Sbjct: 559 QEESKEVQEDEEEVEEDEEEEEEEEEEEEE 588
>MYC_CARAU (P49709) Myc protein (c-myc)
Length = 399
Score = 46.2 bits (108), Expect = 3e-04
Identities = 24/60 (40%), Positives = 36/60 (60%), Gaps = 2/60 (3%)
Query: 200 LSEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDE 259
L+E S SN + + P ++ S++ D DE+EEEEEEE+EE++E E E E+E
Sbjct: 174 LTESSKSNKVAPSQ--PMLVLDTPPNSSSSSGSDSEDEEEEEEEEEEEEEEEEEEEEEEE 231
Score = 34.3 bits (77), Expect = 1.2
Identities = 33/114 (28%), Positives = 50/114 (42%), Gaps = 30/114 (26%)
Query: 480 HTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVP 539
H K TSP+T D+ + + K R H + ER+RR +L F LR +P
Sbjct: 297 HGKQRKCTSPRTSDS--EDNDKRR-----------THNVLERQRRNELKLSFFALRDEIP 343
Query: 540 FVTKMDKAS---ILGDTIEYL--------------KQLRRKIQDLETRNRQIET 576
V +KA+ IL E + +QLRRK + L+ R +Q+ +
Sbjct: 344 EVANNEKAAKVVILKKATECIHSMQLDEQRLLSIKEQLRRKSEQLKHRLQQLRS 397
Score = 32.0 bits (71), Expect = 5.7
Identities = 19/65 (29%), Positives = 33/65 (50%), Gaps = 7/65 (10%)
Query: 195 PPKPALSEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDEEEEEEEDEEDDEVES 254
P +P L + N +SS+ + ++ +++ +E+EEEEEEE+EE D V
Sbjct: 185 PSQPMLVLDTPPNSSSSSG-------SDSEDEEEEEEEEEEEEEEEEEEEEEEEIDVVTV 237
Query: 255 ESEDE 259
E +
Sbjct: 238 EKRQK 242
>S230_PLAFO (Q08372) Transmission-blocking target antigen S230
precursor
Length = 3135
Score = 45.8 bits (107), Expect = 4e-04
Identities = 32/100 (32%), Positives = 47/100 (47%), Gaps = 8/100 (8%)
Query: 226 PSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDETGGRIRRATSMTAIVEPSELMQIEMP 285
P N DD +E+EEEEEEE+EE++E E E E+E + + E E +Q E
Sbjct: 271 PRDNFVIDDEEEEEEEEEEEEEEEEEEEEEEEEEYDDYVYEESG----DETEEQLQEEHQ 326
Query: 286 DDIRIGSPNDGSNHLDSDFHLLVVSNQGNPLGHVDSYKTD 325
+++ S + N D D V + + VD Y D
Sbjct: 327 EEVGAESSEESFNDEDED----SVEARDGDMIRVDEYYED 362
Score = 42.7 bits (99), Expect = 0.003
Identities = 30/105 (28%), Positives = 47/105 (44%), Gaps = 8/105 (7%)
Query: 182 KHVKSFFLDGHS------LPPKPALSEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDM 235
KH +FF++ L P + + + D P + + D +++
Sbjct: 231 KHKSNFFINSSLKYIYMYLTPSDSFNLVRRNRNLDEEDMSPRDNFVIDDEEEEEEEEEEE 290
Query: 236 DEDEEEEEEEDEE--DDEVESESEDETGGRIRRATSMTAIVEPSE 278
+E+EEEEEEE+EE DD V ES DET +++ E SE
Sbjct: 291 EEEEEEEEEEEEEEYDDYVYEESGDETEEQLQEEHQEEVGAESSE 335
>MAX_DROME (P91664) Max protein (dMax)
Length = 161
Score = 45.8 bits (107), Expect = 4e-04
Identities = 32/87 (36%), Positives = 43/87 (48%), Gaps = 6/87 (6%)
Query: 493 DAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFV--TKMDKASIL 550
D G+ S Q E A+H ERRRR+ + E F LR VP + K +A IL
Sbjct: 21 DTGLGSSRHTNTANFTQAEKRAHHNALERRRRDHIKESFTNLREAVPTLKGEKASRAQIL 80
Query: 551 GDTIEYLKQLRRKI----QDLETRNRQ 573
T E ++ +RRKI +D+E RQ
Sbjct: 81 KKTTECIQTMRRKISENQKDIEEIKRQ 107
>CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1) (CLK-1)
(CDK11) (p58 CLK-1)
Length = 795
Score = 45.8 bits (107), Expect = 4e-04
Identities = 33/103 (32%), Positives = 51/103 (49%), Gaps = 21/103 (20%)
Query: 201 SEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDEEEE---EEEDEEDDEVESESE 257
SE TS+ SS+ A+ S + +++ +E+EEEE EE EE++E E E E
Sbjct: 285 SERKTSSAESSS----------AESGSGSEEEEEEEEEEEEEGSTSEESEEEEEEEEEEE 334
Query: 258 DETGGRIRRATSMTAIVEPSELMQIEMPDDIRIGSPNDGSNHL 300
+ETG A+ +A E+ + EM +D + NHL
Sbjct: 335 EETGSNSEEASEQSA----EEVSEEEMSED----EERENENHL 369
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.131 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 84,947,983
Number of Sequences: 164201
Number of extensions: 3806176
Number of successful extensions: 41503
Number of sequences better than 10.0: 855
Number of HSP's better than 10.0 without gapping: 579
Number of HSP's successfully gapped in prelim test: 290
Number of HSP's that attempted gapping in prelim test: 25076
Number of HSP's gapped (non-prelim): 5952
length of query: 701
length of database: 59,974,054
effective HSP length: 117
effective length of query: 584
effective length of database: 40,762,537
effective search space: 23805321608
effective search space used: 23805321608
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC135317.3


