

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.8 + phase: 0
(189 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
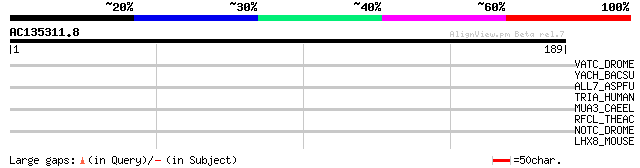
VATC_DROME (Q9V7N5) Vacuolar ATP synthase subunit C (EC 3.6.3.14... 31 1.9
YACH_BACSU (P37569) Hypothetical protein yacH 30 3.3
ALL7_ASPFU (O42799) Allergen Asp f 7 (Fragment) 30 4.3
TRIA_HUMAN (Q15643) Thyroid receptor interacting protein 11 (TRI... 29 7.4
MUA3_CAEEL (P34576) Transmembrane cell adhesion receptor mua-3 p... 29 7.4
RFCL_THEAC (Q9HIP7) Replication factor C large subunit (RFC larg... 28 9.7
NOTC_DROME (P07207) Neurogenic locus Notch protein precursor 28 9.7
LHX8_MOUSE (O35652) LIM/homeobox protein Lhx8 (LIM homeodomain L... 28 9.7
>VATC_DROME (Q9V7N5) Vacuolar ATP synthase subunit C (EC 3.6.3.14)
(V-ATPase C subunit) (Vacuolar proton pump C subunit)
Length = 714
Score = 30.8 bits (68), Expect = 1.9
Identities = 21/69 (30%), Positives = 28/69 (40%), Gaps = 6/69 (8%)
Query: 71 STHTKAYLTLNSFQKGGDGGGPSE----CDKQYHSDDTPVVALS--TGWFNHKSRCLNNI 124
ST A ++ GG GGGP+ C Y S P + S + S+ LNN
Sbjct: 265 STSAAAAAAASNSSGGGGGGGPASCSTLCSSAYFSTSAPTTSSSVHSSMSRSNSKRLNNN 324
Query: 125 TISANGRNV 133
T S N +
Sbjct: 325 TCSINNNKL 333
>YACH_BACSU (P37569) Hypothetical protein yacH
Length = 185
Score = 30.0 bits (66), Expect = 3.3
Identities = 23/78 (29%), Positives = 38/78 (48%), Gaps = 2/78 (2%)
Query: 67 SPSVSTHTKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVV-ALSTGWFNHKSRCLNNIT 125
S +S K +T F+K G G SEC K +HS+ TP++ + +G H + I
Sbjct: 75 SEQISACPKCGMTFQQFRKIGRFGC-SECYKTFHSNITPILRKVHSGNTVHAGKIPKRIG 133
Query: 126 ISANGRNVVAMVVDECDS 143
+ + R + M+ E +S
Sbjct: 134 GNLHVRRQIDMLKKELES 151
>ALL7_ASPFU (O42799) Allergen Asp f 7 (Fragment)
Length = 112
Score = 29.6 bits (65), Expect = 4.3
Identities = 21/69 (30%), Positives = 25/69 (35%), Gaps = 1/69 (1%)
Query: 80 LNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISANGRNVVAMVVD 139
L + PS C VVAL G C +TI+ NG A VVD
Sbjct: 18 LTYYDTATSASAPSSCGLTNDGFSENVVALPVGIMTDAD-CGKTVTITYNGITKTATVVD 76
Query: 140 ECDSRKGCD 148
+C K D
Sbjct: 77 KCMGCKPTD 85
>TRIA_HUMAN (Q15643) Thyroid receptor interacting protein 11 (TRIP-11)
(Golgi-associated microtubule-binding protein 210)
(GMAP-210) (Trip230) (Clonal evolution related gene on
chromosome 14)
Length = 1979
Score = 28.9 bits (63), Expect = 7.4
Identities = 17/63 (26%), Positives = 26/63 (40%)
Query: 94 ECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISANGRNVVAMVVDECDSRKGCDEQHDY 153
E ++ +H D V TGW S+ + N + N ++VV E + E H
Sbjct: 1816 EMEQLFHDDQGSVTRWMTGWLGGGSKSVPNTPLRPNQQSVVNSSFSELFVKFLETESHPS 1875
Query: 154 QPP 156
PP
Sbjct: 1876 IPP 1878
>MUA3_CAEEL (P34576) Transmembrane cell adhesion receptor mua-3
precursor
Length = 3767
Score = 28.9 bits (63), Expect = 7.4
Identities = 37/150 (24%), Positives = 56/150 (36%), Gaps = 29/150 (19%)
Query: 35 RIR-GKKAPPGKCNQENDSDCCV--------QGKMYTTYVCSPSVSTHTKAYLTLNSFQK 85
RIR G+ A PG CN + C V +G +YT C + H + L K
Sbjct: 1555 RIRNGQCAYPGSCNPNDPMSCDVRKRQQCLPRGNIYTCQ-CGRNEKRHPITDICL----K 1609
Query: 86 GGDGGGPSECDKQ---YHSDDTPVVALSTGWFNHKSRCLNNITISANGRNVVAMVVDECD 142
G +CD+ +D++ + A +G+ +H GR VA+ + D
Sbjct: 1610 NECLTGEHDCDRSARCIDTDESYICACQSGFIDHSPN-----PSERPGRVCVALQNECLD 1664
Query: 143 SRKGC-------DEQHDYQPPCPNNIVDAS 165
C D + Y C + VD S
Sbjct: 1665 GSNRCSPNALCTDTEEGYVCRCKSGFVDYS 1694
>RFCL_THEAC (Q9HIP7) Replication factor C large subunit (RFC large
subunit) (Clamp loader large subunit)
Length = 418
Score = 28.5 bits (62), Expect = 9.7
Identities = 15/49 (30%), Positives = 23/49 (46%)
Query: 81 NSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISAN 129
N F++GGD GG E K + P+V F + + + ISA+
Sbjct: 114 NMFERGGDTGGIYELSKIIKNSANPIVITMNDIFEFRKKNYASDVISAS 162
>NOTC_DROME (P07207) Neurogenic locus Notch protein precursor
Length = 2703
Score = 28.5 bits (62), Expect = 9.7
Identities = 28/135 (20%), Positives = 48/135 (34%), Gaps = 14/135 (10%)
Query: 44 GKCNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLTLNSFQKGGDGGGPSECD------- 96
GKC + +S C+ YT Y+C ++ + + G +C
Sbjct: 614 GKCIDDVNSFKCLCDPGYTGYICQKQINECESNPCQFDGHCQDRVGSYYCQCQAGTSGKN 673
Query: 97 -----KQYHSDDTPVVALSTGWFN-HKSRCLNNITISANGRNVVAMVVDECDSRKGC-DE 149
+ HS+ A N +K +C+ T +NV + C + C D+
Sbjct: 674 CEVNVNECHSNPCNNGATCIDGINSYKCQCVPGFTGQHCEKNVDECISSPCANNGVCIDQ 733
Query: 150 QHDYQPPCPNNIVDA 164
+ Y+ CP DA
Sbjct: 734 VNGYKCECPRGFYDA 748
>LHX8_MOUSE (O35652) LIM/homeobox protein Lhx8 (LIM homeodomain
Lhx7) (L3)
Length = 367
Score = 28.5 bits (62), Expect = 9.7
Identities = 23/91 (25%), Positives = 33/91 (35%), Gaps = 21/91 (23%)
Query: 32 PSGRIRGKKAPPGKCNQEND------------SDCCVQGKMYTTYVCSPSVSTHTKAYL- 78
P G PPGKC + +D C + + VC S+ HT Y+
Sbjct: 80 PQSMASGSVCPPGKCVCSSCGLEIVDKYLLKVNDLCWHVRCLSCSVCRTSLGRHTSCYIK 139
Query: 79 ------TLNSFQKGGDGGGPSECDKQYHSDD 103
L+ F++ G S C + HS D
Sbjct: 140 DKDIFCKLDYFRRYGT--RCSRCGRHIHSTD 168
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.134 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,187,160
Number of Sequences: 164201
Number of extensions: 1083785
Number of successful extensions: 1952
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1950
Number of HSP's gapped (non-prelim): 8
length of query: 189
length of database: 59,974,054
effective HSP length: 104
effective length of query: 85
effective length of database: 42,897,150
effective search space: 3646257750
effective search space used: 3646257750
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135311.8


