

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135233.1 + phase: 0
(87 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
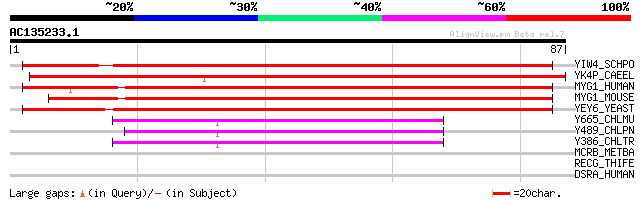
Score E
Sequences producing significant alignments: (bits) Value
YIW4_SCHPO (Q9P7T6) Hypothetical UPF0160 protein C694.04c in chr... 106 9e-24
YK4P_CAEEL (O17606) Hypothetical UPF0160 protein C27H6.8 in chro... 104 5e-23
MYG1_HUMAN (Q9HB07) MYG1 protein 99 2e-21
MYG1_MOUSE (Q9JK81) MYG1 protein (Gamm1 protein) 96 1e-20
YEY6_YEAST (P40093) Hypothetical UPF0160 protein YER156c 79 2e-15
Y665_CHLMU (Q9PK08) Hypothetical UPF0160 protein TC0665 41 5e-04
Y489_CHLPN (Q9Z862) UPF0160 protein CPn0489/CP0265/CPj0489/CpB0509 41 5e-04
Y386_CHLTR (O84391) Hypothetical UPF0160 protein CT386 40 8e-04
MCRB_METBA (P07955) Methyl-coenzyme M reductase beta subunit (EC... 28 4.1
RECG_THIFE (O50224) ATP-dependent DNA helicase recG (EC 3.6.1.-) 28 5.4
DSRA_HUMAN (P55265) Double-stranded RNA-specific adenosine deami... 27 7.0
>YIW4_SCHPO (Q9P7T6) Hypothetical UPF0160 protein C694.04c in
chromosome I
Length = 324
Score = 106 bits (265), Expect = 9e-24
Identities = 51/83 (61%), Positives = 60/83 (71%), Gaps = 2/83 (2%)
Query: 3 IDPPIKYVLYEDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCVFV 62
I+ KY +Y D K WRVQAVS+ P F R LP WRG+RD+ LS+ +GIPGC+FV
Sbjct: 239 IENQFKYAIYSD--GKAWRVQAVSIDPTSFTCRLPLPEPWRGIRDEKLSELTGIPGCIFV 296
Query: 63 HMSGFIGGNQTFEGALAMAKAAL 85
H SGFIGGNQTFEGAL MA+ AL
Sbjct: 297 HASGFIGGNQTFEGALEMARKAL 319
>YK4P_CAEEL (O17606) Hypothetical UPF0160 protein C27H6.8 in
chromosome V
Length = 340
Score = 104 bits (259), Expect = 5e-23
Identities = 52/85 (61%), Positives = 63/85 (73%), Gaps = 1/85 (1%)
Query: 4 DPPIKYVLYEDERSKQWRVQAVSVSP-DRFESRKALPSQWRGLRDDILSKESGIPGCVFV 62
D I Y+L+ D + WRVQA+ V FE+R LP+ WRGLRDD LSKESGIPG VFV
Sbjct: 243 DDNITYILFSDSTNASWRVQAIPVDKMSSFENRMPLPAAWRGLRDDDLSKESGIPGGVFV 302
Query: 63 HMSGFIGGNQTFEGALAMAKAALKM 87
H+SGFIGGN T EGA+AMA+ AL++
Sbjct: 303 HISGFIGGNLTREGAIAMARKALEI 327
>MYG1_HUMAN (Q9HB07) MYG1 protein
Length = 376
Score = 98.6 bits (244), Expect = 2e-21
Identities = 47/85 (55%), Positives = 60/85 (70%), Gaps = 3/85 (3%)
Query: 3 IDPPIK--YVLYEDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCV 60
+ PP+ +V+Y D+ QWR+Q V P F+SR LP WRGLRD+ L + SGIPGC+
Sbjct: 283 LSPPVAIFFVIYTDQAG-QWRIQCVPKEPHSFQSRLPLPEPWRGLRDEALDQVSGIPGCI 341
Query: 61 FVHMSGFIGGNQTFEGALAMAKAAL 85
FVH SGFIGG+ T EGAL+MA+A L
Sbjct: 342 FVHASGFIGGHPTREGALSMARATL 366
>MYG1_MOUSE (Q9JK81) MYG1 protein (Gamm1 protein)
Length = 380
Score = 96.3 bits (238), Expect = 1e-20
Identities = 47/79 (59%), Positives = 55/79 (69%), Gaps = 1/79 (1%)
Query: 7 IKYVLYEDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCVFVHMSG 66
I +V+Y D+ QWRVQ V P F+SR LP WRGLRD L + SGIPGC+FVH SG
Sbjct: 288 ITFVIYTDQAG-QWRVQCVPKEPHSFQSRLPLPEPWRGLRDKALDQVSGIPGCIFVHASG 346
Query: 67 FIGGNQTFEGALAMAKAAL 85
FIGG+ T EGAL MA+A L
Sbjct: 347 FIGGHHTREGALNMARATL 365
>YEY6_YEAST (P40093) Hypothetical UPF0160 protein YER156c
Length = 338
Score = 79.0 bits (193), Expect = 2e-15
Identities = 37/83 (44%), Positives = 55/83 (65%), Gaps = 1/83 (1%)
Query: 3 IDPPIKYVLYEDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCVFV 62
I+ I++VL+ D S WRV V ++ F+ R+ LP RGLRD+ LS +SG+PGC+F+
Sbjct: 256 IEKQIEFVLFTDS-SGAWRVSTVPINSTSFQFRRGLPEPLRGLRDEELSTKSGVPGCIFI 314
Query: 63 HMSGFIGGNQTFEGALAMAKAAL 85
H +GFIGG ++ E +AK +L
Sbjct: 315 HAAGFIGGAKSKEAVYELAKMSL 337
>Y665_CHLMU (Q9PK08) Hypothetical UPF0160 protein TC0665
Length = 289
Score = 41.2 bits (95), Expect = 5e-04
Identities = 21/53 (39%), Positives = 28/53 (52%), Gaps = 1/53 (1%)
Query: 17 SKQWRVQAVSVSPDR-FESRKALPSQWRGLRDDILSKESGIPGCVFVHMSGFI 68
S QW ++ + + DR E R P W GL D L K +GIPG +F H F+
Sbjct: 214 SDQWILRGIPPTLDRRMEVRIPFPEDWAGLLGDQLVKATGIPGAIFCHKGLFL 266
>Y489_CHLPN (Q9Z862) UPF0160 protein CPn0489/CP0265/CPj0489/CpB0509
Length = 290
Score = 41.2 bits (95), Expect = 5e-04
Identities = 22/51 (43%), Positives = 27/51 (52%), Gaps = 1/51 (1%)
Query: 19 QWRVQAVSVSPDR-FESRKALPSQWRGLRDDILSKESGIPGCVFVHMSGFI 68
QW ++ + + DR E R P W GL LSK SGIPG VF H F+
Sbjct: 217 QWILRGIPPNLDRRMEVRVPFPENWAGLLGKELSKVSGIPGAVFCHKGLFL 267
>Y386_CHLTR (O84391) Hypothetical UPF0160 protein CT386
Length = 289
Score = 40.4 bits (93), Expect = 8e-04
Identities = 20/53 (37%), Positives = 29/53 (53%), Gaps = 1/53 (1%)
Query: 17 SKQWRVQAVSVSPDR-FESRKALPSQWRGLRDDILSKESGIPGCVFVHMSGFI 68
S QW ++ + + DR E R P +W GL D L + +GIPG +F H F+
Sbjct: 214 SDQWILRGIPPTLDRRMEVRIPFPEEWAGLLGDQLVQATGIPGAIFCHKGLFL 266
>MCRB_METBA (P07955) Methyl-coenzyme M reductase beta subunit (EC
2.8.4.1) (Coenzyme-B sulfoethylthiotransferase beta)
Length = 433
Score = 28.1 bits (61), Expect = 4.1
Identities = 12/32 (37%), Positives = 19/32 (58%)
Query: 47 DDILSKESGIPGCVFVHMSGFIGGNQTFEGAL 78
+DIL KE+G+PGC + + G G F ++
Sbjct: 332 NDILEKETGLPGCDYGKVEGTAVGFSFFSHSI 363
>RECG_THIFE (O50224) ATP-dependent DNA helicase recG (EC 3.6.1.-)
Length = 652
Score = 27.7 bits (60), Expect = 5.4
Identities = 14/35 (40%), Positives = 22/35 (62%), Gaps = 5/35 (14%)
Query: 18 KQWRVQAVSVS---PDRFESRKALPSQWRGLRDDI 49
++W QA+++ PD E+R LP QW GLR+ +
Sbjct: 188 RRWMAQALTLLDQLPDYLENR--LPPQWPGLREGL 220
>DSRA_HUMAN (P55265) Double-stranded RNA-specific adenosine
deaminase (EC 3.5.4.-) (DRADA) (136 kDa double-stranded
RNA binding protein) (P136) (K88DSRBP)
(Interferon-inducible protein 4) (IFI-4 protein)
Length = 1226
Score = 27.3 bits (59), Expect = 7.0
Identities = 16/63 (25%), Positives = 27/63 (42%), Gaps = 8/63 (12%)
Query: 2 KIDPPIKY-----VLYEDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGI 56
++ PP+ Y Y D + QW + PD S +A P ++R + + G+
Sbjct: 431 RLKPPVHYNGPSKAGYVDFENGQWATDDI---PDDLNSIRAAPGEFRAIMEMPSFYSHGL 487
Query: 57 PGC 59
P C
Sbjct: 488 PRC 490
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,123,823
Number of Sequences: 164201
Number of extensions: 342129
Number of successful extensions: 661
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 651
Number of HSP's gapped (non-prelim): 11
length of query: 87
length of database: 59,974,054
effective HSP length: 63
effective length of query: 24
effective length of database: 49,629,391
effective search space: 1191105384
effective search space used: 1191105384
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135233.1


