

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135232.7 + phase: 0
(221 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
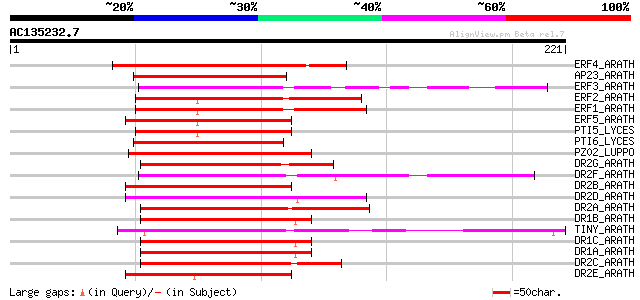
Score E
Sequences producing significant alignments: (bits) Value
ERF4_ARATH (O80340) Ethylene responsive element binding factor 4... 102 9e-22
AP23_ARATH (P42736) AP2 domain transcription factor RAP2.3 (Rela... 99 6e-21
ERF3_ARATH (O80339) Ethylene responsive element binding factor 3... 99 8e-21
ERF2_ARATH (O80338) Ethylene responsive element binding factor 2... 96 7e-20
ERF1_ARATH (O80337) Ethylene responsive element binding factor 1... 96 7e-20
ERF5_ARATH (O80341) Ethylene responsive element binding factor 5... 95 1e-19
PTI5_LYCES (O04681) Pathogenesis-related genes transcriptional a... 91 2e-18
PTI6_LYCES (O04682) Pathogenesis-related genes transcriptional a... 88 1e-17
PZ02_LUPPO (P16146) PPLZ02 protein 86 9e-17
DR2G_ARATH (P61827) Putative dehydration responsive element bind... 86 9e-17
DR2F_ARATH (Q9SVX5) Dehydration responsive element binding prote... 85 1e-16
DR2B_ARATH (O82133) Dehydration responsive element binding prote... 85 1e-16
DR2D_ARATH (Q9LQZ2) Putative dehydration responsive element bind... 83 4e-16
DR2A_ARATH (O82132) Dehydration responsive element binding prote... 83 4e-16
DR1B_ARATH (P93835) Dehydration responsive element binding prote... 80 3e-15
TINY_ARATH (Q39127) Transcriptional factor TINY 80 4e-15
DR1C_ARATH (Q9SYS6) Dehydration responsive element binding prote... 79 8e-15
DR1A_ARATH (Q9M0L0) Dehydration responsive element binding prote... 79 1e-14
DR2C_ARATH (Q8LFR2) Dehydration responsive element binding prote... 78 2e-14
DR2E_ARATH (O80917) Dehydration responsive element binding prote... 76 5e-14
>ERF4_ARATH (O80340) Ethylene responsive element binding factor 4
(AtERF4)
Length = 222
Score = 102 bits (253), Expect = 9e-22
Identities = 52/93 (55%), Positives = 67/93 (71%), Gaps = 1/93 (1%)
Query: 42 NVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRG 101
N + + + +YRGVR+RPWG++AAEIRDP K TRVWLGTF+TAE+AA+AYD A+ FRG
Sbjct: 14 NQTHNNAKEIRYRGVRKRPWGRYAAEIRDPGKKTRVWLGTFDTAEEAARAYDTAARDFRG 73
Query: 102 NKAKLNFPENVKLKQQQSTPTHLNISHSNSALL 134
KAK NFP ++L Q+ PT S S S+ L
Sbjct: 74 AKAKTNFPTFLELSDQK-VPTGFARSPSQSSTL 105
>AP23_ARATH (P42736) AP2 domain transcription factor RAP2.3 (Related
to AP2 protein 3) (Cadmium-induced protein AS30)
Length = 248
Score = 99.4 bits (246), Expect = 6e-21
Identities = 44/61 (72%), Positives = 51/61 (83%)
Query: 50 KRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
K YRG+R+RPWGKWAAEIRDP K RVWLGTF TAE+AA AYD A+ + RG+KAKLNFP
Sbjct: 76 KNVYRGIRKRPWGKWAAEIRDPRKGVRVWLGTFNTAEEAAMAYDVAAKQIRGDKAKLNFP 135
Query: 110 E 110
+
Sbjct: 136 D 136
>ERF3_ARATH (O80339) Ethylene responsive element binding factor 3
(AtERF3)
Length = 225
Score = 99.0 bits (245), Expect = 8e-21
Identities = 60/163 (36%), Positives = 89/163 (53%), Gaps = 30/163 (18%)
Query: 52 KYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPEN 111
++RGVR+RPWG++AAEIRDP+K RVWLGTF++AE+AA+AYD A+ RG KAK NFP
Sbjct: 27 RFRGVRKRPWGRFAAEIRDPWKKARVWLGTFDSAEEAARAYDSAARNLRGPKAKTNFP-- 84
Query: 112 VKLKQQQSTPTHLNISHSNSALLSHQPRTDPDPIVHNETLHTLQSSNKYYDYFNGQNFPM 171
+ P +L + + +Q + DP + H L + ++ Q FP+
Sbjct: 85 --IDSSSPPPPNLRFNQ-----IRNQNQNQVDPFMD----HRLFTDHQ-------QQFPI 126
Query: 172 ASLQTSVSPSASYSSSYTTTTFASSFSSPQTTSSIPSGLTAWP 214
+ TS S S++ SFS P+ T+ P+ +P
Sbjct: 127 VNRPTSSSMSST----------VESFSGPRPTTMKPATTKRYP 159
>ERF2_ARATH (O80338) Ethylene responsive element binding factor 2
(AtERF2)
Length = 243
Score = 95.9 bits (237), Expect = 7e-20
Identities = 50/91 (54%), Positives = 62/91 (67%), Gaps = 3/91 (3%)
Query: 51 RKYRGVRQRPWGKWAAEIRDPFK-ATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
+ YRGVRQRPWGK+AAEIRDP K RVWLGTFETAEDAA AYD A+ R RG++A LNFP
Sbjct: 115 KHYRGVRQRPWGKFAAEIRDPAKNGARVWLGTFETAEDAALAYDIAAFRMRGSRALLNFP 174
Query: 110 ENVKLKQQQSTPTHLNISHSNSALLSHQPRT 140
+++ + P + S+S+ S T
Sbjct: 175 --LRVNSGEPDPVRITSKRSSSSSSSSSSST 203
>ERF1_ARATH (O80337) Ethylene responsive element binding factor 1
(AtERF1) (EREBP-2 protein)
Length = 268
Score = 95.9 bits (237), Expect = 7e-20
Identities = 51/93 (54%), Positives = 60/93 (63%), Gaps = 5/93 (5%)
Query: 51 RKYRGVRQRPWGKWAAEIRDPFK-ATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
+ YRGVRQRPWGK+AAEIRDP K RVWLGTFETAEDAA AYD+A+ R RG++A LNFP
Sbjct: 146 KHYRGVRQRPWGKFAAEIRDPAKNGARVWLGTFETAEDAALAYDRAAFRMRGSRALLNFP 205
Query: 110 ENVKLKQQQSTPTHLNISHSNSALLSHQPRTDP 142
L+ P + I S+ S P
Sbjct: 206 ----LRVNSGEPDPVRIKSKRSSFSSSNENGAP 234
>ERF5_ARATH (O80341) Ethylene responsive element binding factor 5
(AtERF5)
Length = 300
Score = 94.7 bits (234), Expect = 1e-19
Identities = 45/67 (67%), Positives = 56/67 (83%), Gaps = 1/67 (1%)
Query: 47 DQPKRKYRGVRQRPWGKWAAEIRDPFK-ATRVWLGTFETAEDAAKAYDQASLRFRGNKAK 105
++ K+ YRGVRQRPWGK+AAEIRDP K +RVWLGTF+TA +AA+AYD+A+ R RG+KA
Sbjct: 150 EEEKKHYRGVRQRPWGKFAAEIRDPNKRGSRVWLGTFDTAIEAARAYDEAAFRLRGSKAI 209
Query: 106 LNFPENV 112
LNFP V
Sbjct: 210 LNFPLEV 216
>PTI5_LYCES (O04681) Pathogenesis-related genes transcriptional
activator PTI5
Length = 161
Score = 90.9 bits (224), Expect = 2e-18
Identities = 43/63 (68%), Positives = 51/63 (80%), Gaps = 1/63 (1%)
Query: 51 RKYRGVRQRPWGKWAAEIRDPFK-ATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
+KYRGVR+RPWGK+AAEIRD + RVWLGTFETAE+AA AYD+A+ R RG KA LNFP
Sbjct: 57 KKYRGVRRRPWGKYAAEIRDSARHGARVWLGTFETAEEAALAYDRAAFRMRGAKALLNFP 116
Query: 110 ENV 112
+
Sbjct: 117 SEI 119
>PTI6_LYCES (O04682) Pathogenesis-related genes transcriptional
activator PTI6
Length = 248
Score = 88.2 bits (217), Expect = 1e-17
Identities = 36/60 (60%), Positives = 49/60 (81%)
Query: 50 KRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
++K+RGVRQRPWG+WAAEIRDP + RVWLGT++T E+AA YD+A+++ +G A NFP
Sbjct: 95 RKKFRGVRQRPWGRWAAEIRDPTRGKRVWLGTYDTPEEAAVVYDKAAVKLKGPDAVTNFP 154
>PZ02_LUPPO (P16146) PPLZ02 protein
Length = 164
Score = 85.5 bits (210), Expect = 9e-17
Identities = 37/73 (50%), Positives = 50/73 (67%)
Query: 48 QPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLN 107
+P+++YRG RQR WG W +EIR TR+W GTFE+AEDAA+AYD+A+ G +A+ N
Sbjct: 3 RPQQRYRGFRQRHWGSWVSEIRHSILKTRIWQGTFESAEDAARAYDEAARLMCGTRARTN 62
Query: 108 FPENVKLKQQQST 120
FP N Q S+
Sbjct: 63 FPYNPNASQSSSS 75
>DR2G_ARATH (P61827) Putative dehydration responsive element binding
protein 2G (DREB2G protein)
Length = 307
Score = 85.5 bits (210), Expect = 9e-17
Identities = 41/77 (53%), Positives = 55/77 (71%), Gaps = 3/77 (3%)
Query: 53 YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPENV 112
+RGVRQR WGKW AEIR+P + TR+WLGTF T+ +AA AYD+A+ + G++AKLN V
Sbjct: 34 FRGVRQRTWGKWVAEIREPNRGTRLWLGTFNTSVEAAMAYDEAAKKLYGHEAKLNL---V 90
Query: 113 KLKQQQSTPTHLNISHS 129
+QQQ + N+S S
Sbjct: 91 HPQQQQQVVVNRNLSFS 107
>DR2F_ARATH (Q9SVX5) Dehydration responsive element binding protein
2F (DREB2F protein)
Length = 277
Score = 85.1 bits (209), Expect = 1e-16
Identities = 56/162 (34%), Positives = 82/162 (50%), Gaps = 15/162 (9%)
Query: 52 KYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPEN 111
+YRGVRQR WGKW AEIR+P K R+WLG+F TAE+AA AYD+A+L+ G+ A LN P
Sbjct: 27 QYRGVRQRTWGKWVAEIREPKKRARLWLGSFATAEEAAMAYDEAALKLYGHDAYLNLPH- 85
Query: 112 VKLKQQQSTPTHLNISH----SNSALLSHQPRTDPDPIVHNETLHTLQSSNKYYDYFNGQ 167
Q+ + P+ N + +S P + ++H +Q + +
Sbjct: 86 ---LQRNTRPSLSNSQRFKWVPSRKFISMFPSCGMLNVNAQPSVHIIQQRLE-------E 135
Query: 168 NFPMASLQTSVSPSASYSSSYTTTTFASSFSSPQTTSSIPSG 209
L S S S+S + S T T+F +S T ++ G
Sbjct: 136 LKKTGLLSQSYSSSSSSTESKTNTSFLDEKTSKGETDNMFEG 177
>DR2B_ARATH (O82133) Dehydration responsive element binding protein
2B (DREB2B protein)
Length = 330
Score = 85.1 bits (209), Expect = 1e-16
Identities = 39/66 (59%), Positives = 48/66 (72%)
Query: 47 DQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKL 106
D +RGVRQR WGKW AEIR+P TR+WLGTF TAE AA AYD+A+ G+ A+L
Sbjct: 72 DNSHCSFRGVRQRIWGKWVAEIREPKIGTRLWLGTFPTAEKAASAYDEAATAMYGSLARL 131
Query: 107 NFPENV 112
NFP++V
Sbjct: 132 NFPQSV 137
>DR2D_ARATH (Q9LQZ2) Putative dehydration responsive element binding
protein 2D (DREB2D protein)
Length = 206
Score = 83.2 bits (204), Expect = 4e-16
Identities = 41/97 (42%), Positives = 57/97 (58%), Gaps = 1/97 (1%)
Query: 47 DQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKL 106
D Y+GVRQR WGKW AEIR+P + R+WLGTF+T+ +AA AYD A+ + G +A L
Sbjct: 36 DNASCTYKGVRQRTWGKWVAEIREPNRGARLWLGTFDTSREAALAYDSAARKLYGPEAHL 95
Query: 107 NFPENVK-LKQQQSTPTHLNISHSNSALLSHQPRTDP 142
N PE+++ + S+P SN+ S P
Sbjct: 96 NLPESLRSYPKTASSPASQTTPSSNTGGKSSSDSESP 132
>DR2A_ARATH (O82132) Dehydration responsive element binding protein
2A (DREB2A protein)
Length = 335
Score = 83.2 bits (204), Expect = 4e-16
Identities = 41/91 (45%), Positives = 56/91 (61%), Gaps = 1/91 (1%)
Query: 53 YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPENV 112
+RGVRQR WGKW AEIR+P + +R+WLGTF TA++AA AYD+A+ G A+LNFP +
Sbjct: 79 FRGVRQRIWGKWVAEIREPNRGSRLWLGTFPTAQEAASAYDEAAKAMYGPLARLNFPRS- 137
Query: 113 KLKQQQSTPTHLNISHSNSALLSHQPRTDPD 143
+ ST + + + H DPD
Sbjct: 138 DASEVTSTSSQSEVCTVETPGCVHVKTEDPD 168
>DR1B_ARATH (P93835) Dehydration responsive element binding protein
1B (DREB1B protein) (C-repeat binding factor 1)
(C-repeat/dehydration responsive element binding factor
1) (CRT/DRE binding factor 1)
Length = 213
Score = 80.5 bits (197), Expect = 3e-15
Identities = 39/69 (56%), Positives = 52/69 (74%), Gaps = 1/69 (1%)
Query: 53 YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPENV 112
YRGVRQR GKW +E+R+P K TR+WLGTF+TAE AA+A+D A+L RG A LNF ++
Sbjct: 48 YRGVRQRNSGKWVSEVREPNKKTRIWLGTFQTAEMAARAHDVAALALRGRSACLNFADSA 107
Query: 113 -KLKQQQST 120
+L+ +ST
Sbjct: 108 WRLRIPEST 116
>TINY_ARATH (Q39127) Transcriptional factor TINY
Length = 218
Score = 80.1 bits (196), Expect = 4e-15
Identities = 62/188 (32%), Positives = 88/188 (45%), Gaps = 44/188 (23%)
Query: 44 KNEDQPKRK---------YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQ 94
+NE++ K+ YRGVR+R WGKW +EIR+P K +R+WLGTF + E AA+A+D
Sbjct: 18 ENEEEKKKPVKDSGKHPVYRGVRKRNWGKWVSEIREPRKKSRIWLGTFPSPEMAARAHDV 77
Query: 95 ASLRFRGNKAKLNFPENVKLKQQQSTPTHLNISHSNSALLSHQPRTDPDPIVHNETLHTL 154
A+L +G A LNFP+ L P+ L+ A L H ET +
Sbjct: 78 AALSIKGASAILNFPD---LAGSFPRPSSLSPRDIQVAALK---------AAHMETSQSF 125
Query: 155 QSSNKYYDYFNGQNFPMASLQTSVSPSASYSSSYTTTTFASSFSSPQTTSSIPSGLTAWP 214
SS+ S ++SSS ++++ S SS T S + P
Sbjct: 126 SSSS----------------------SLTFSSSQSSSSLESLVSSSATGSEELGEIVELP 163
Query: 215 S-GSSPSG 221
S GSS G
Sbjct: 164 SLGSSYDG 171
>DR1C_ARATH (Q9SYS6) Dehydration responsive element binding protein
1C (DREB1C protein) (C-repeat binding factor 2)
(C-repeat/dehydration responsive element binding factor
2) (CRT/DRE binding factor 2)
Length = 216
Score = 79.0 bits (193), Expect = 8e-15
Identities = 38/69 (55%), Positives = 51/69 (73%), Gaps = 1/69 (1%)
Query: 53 YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPENV 112
YRGVRQR GKW E+R+P K TR+WLGTF+TAE AA+A+D A++ RG A LNF ++
Sbjct: 51 YRGVRQRNSGKWVCELREPNKKTRIWLGTFQTAEMAARAHDVAAIALRGRSACLNFADSA 110
Query: 113 -KLKQQQST 120
+L+ +ST
Sbjct: 111 WRLRIPEST 119
>DR1A_ARATH (Q9M0L0) Dehydration responsive element binding protein
1A (DREB1A protein) (C-repeat binding factor 3)
(C-repeat/dehydration responsive element binding factor
3) (CRT/DRE binding factor 3)
Length = 216
Score = 78.6 bits (192), Expect = 1e-14
Identities = 38/69 (55%), Positives = 51/69 (73%), Gaps = 1/69 (1%)
Query: 53 YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPENV 112
YRGVR+R GKW E+R+P K TR+WLGTF+TAE AA+A+D A+L RG A LNF ++
Sbjct: 51 YRGVRRRNSGKWVCEVREPNKKTRIWLGTFQTAEMAARAHDVAALALRGRSACLNFADSA 110
Query: 113 -KLKQQQST 120
+L+ +ST
Sbjct: 111 WRLRIPEST 119
>DR2C_ARATH (Q8LFR2) Dehydration responsive element binding protein
2C (DREB2C protein)
Length = 341
Score = 77.8 bits (190), Expect = 2e-14
Identities = 39/80 (48%), Positives = 50/80 (61%), Gaps = 2/80 (2%)
Query: 53 YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPENV 112
YRGVRQR WGKW AEIR+P R+WLGTF ++ +AA AYD+A+ G A+LN PE
Sbjct: 72 YRGVRQRRWGKWVAEIREPDGGARLWLGTFSSSYEAALAYDEAAKAIYGQSARLNLPEIT 131
Query: 113 KLKQQQSTPTHLNISHSNSA 132
+ ST +S S +A
Sbjct: 132 --NRSSSTAATATVSGSVTA 149
>DR2E_ARATH (O80917) Dehydration responsive element binding protein
2E (DREB2E protein)
Length = 244
Score = 76.3 bits (186), Expect = 5e-14
Identities = 37/74 (50%), Positives = 50/74 (67%), Gaps = 8/74 (10%)
Query: 47 DQPKRKYRGVRQRPWGKWAAEIRDPF--------KATRVWLGTFETAEDAAKAYDQASLR 98
+ P ++RGVRQR WGKW AEIR+P ++ R+WLGTF TA +AA AYD+A+
Sbjct: 64 ENPVCRFRGVRQRVWGKWVAEIREPVSHRGANSSRSKRLWLGTFATAAEAALAYDRAASV 123
Query: 99 FRGNKAKLNFPENV 112
G A+LNFPE++
Sbjct: 124 MYGPYARLNFPEDL 137
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.310 0.124 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,208,223
Number of Sequences: 164201
Number of extensions: 1127006
Number of successful extensions: 4147
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 64
Number of HSP's successfully gapped in prelim test: 66
Number of HSP's that attempted gapping in prelim test: 3684
Number of HSP's gapped (non-prelim): 409
length of query: 221
length of database: 59,974,054
effective HSP length: 106
effective length of query: 115
effective length of database: 42,568,748
effective search space: 4895406020
effective search space used: 4895406020
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC135232.7


