

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.4 - phase: 0
(360 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
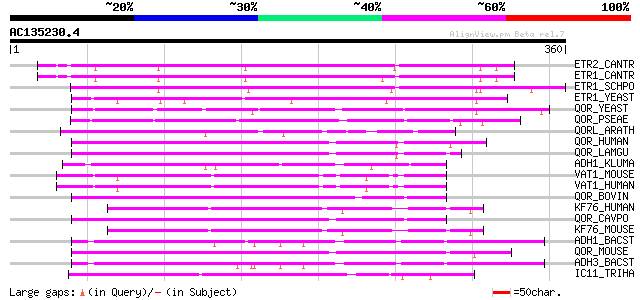
ETR2_CANTR (Q8WZM4) Enoyl-[acyl-carrier protein] reductase [NADP... 211 2e-54
ETR1_CANTR (Q8WZM3) Enoyl-[acyl-carrier protein] reductase [NADP... 210 4e-54
ETR1_SCHPO (Q10488) Enoyl-[acyl-carrier protein] reductase [NADP... 209 9e-54
ETR1_YEAST (P38071) Enoyl-[acyl-carrier protein] reductase [NADP... 160 4e-39
QOR_YEAST (P38230) Probable quinone oxidoreductase (EC 1.6.5.5) ... 77 5e-14
QOR_PSEAE (P43903) Quinone oxidoreductase (EC 1.6.5.5) (NADPH:qu... 77 5e-14
QORL_ARATH (Q9ZUC1) Quinone oxidoreductase-like protein At1g2374... 73 1e-12
QOR_HUMAN (Q08257) Quinone oxidoreductase (EC 1.6.5.5) (NADPH:qu... 71 4e-12
QOR_LAMGU (Q28452) Quinone oxidoreductase (EC 1.6.5.5) (NADPH:qu... 70 6e-12
ADH1_KLUMA (Q07288) Alcohol dehydrogenase 1 (EC 1.1.1.1) 69 1e-11
VAT1_MOUSE (Q62465) Synaptic vesicle membrane protein VAT-1 homo... 68 3e-11
VAT1_HUMAN (Q99536) Synaptic vesicle membrane protein VAT-1 homo... 68 3e-11
QOR_BOVIN (O97764) Zeta-crystallin 68 3e-11
KF76_HUMAN (Q9HCJ6) Probable oxidoreductase KIAA1576 (EC 1.-.-.-) 68 4e-11
QOR_CAVPO (P11415) Quinone oxidoreductase (EC 1.6.5.5) (NADPH:qu... 67 7e-11
KF76_MOUSE (Q80TB8) Probable oxidoreductase KIAA1576 (EC 1.-.-.-) 67 7e-11
ADH1_BACST (P12311) Alcohol dehydrogenase (EC 1.1.1.1) (ADH-T) 67 7e-11
QOR_MOUSE (P47199) Quinone oxidoreductase (EC 1.6.5.5) (NADPH:qu... 67 9e-11
ADH3_BACST (P42328) Alcohol dehydrogenase (EC 1.1.1.1) (ADH-HT) 66 1e-10
IC11_TRIHA (P34055) Protein indc11 65 3e-10
>ETR2_CANTR (Q8WZM4) Enoyl-[acyl-carrier protein] reductase [NADPH,
B-specific] 2, mitochondrial precursor (EC 1.3.1.10)
Length = 386
Score = 211 bits (538), Expect = 2e-54
Identities = 135/348 (38%), Positives = 198/348 (56%), Gaps = 44/348 (12%)
Query: 19 SLRSRILNTHTHAFSSAVSPPSKAIIYESHGQPDAV--TKLVDIPATEVKENDLCVKMLA 76
S+R R+L TH F + ++ ++A++Y HG+P V T+ +I + N++ VK L
Sbjct: 8 SIRPRLLATHNQ-FRTMIT--AQAVLYTQHGEPKDVLFTQSFEIDDDNLAPNEVIVKTLG 64
Query: 77 APINPSDINRIQGVYPVRP---------EPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIP 127
+PINPSDIN+IQGVYP +P EP A G EG+ EV VGS V+ GDWVIP
Sbjct: 65 SPINPSDINQIQGVYPSKPAKTTGFGTAEPAAPCGNEGLFEVIKVGSNVSSLEAGDWVIP 124
Query: 128 SPPSFGTWQTYIVKDQNVWHKVNK-----------GVPMEYAATITVNPLTALLMLEDCV 176
S +FGTW+T+ + + + + K+ G+ + ATI+VNPLTA LML V
Sbjct: 125 SHVNFGTWRTHALGNDDDFIKLPNPAQSKANGKPNGLTINQGATISVNPLTAYLMLTHYV 184
Query: 177 TLNSG-DAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFT 235
L G D +QNG TS VG+ Q+ K ++I++IRDRP + EV LK+LGA +V T
Sbjct: 185 KLTPGKDWFIQNGGTSAVGKYASQIGKLLNFNSISVIRDRPNLDEVVASLKELGATQVIT 244
Query: 236 ESELEVKNVKSLL-------GGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKK 288
E + K + GG E L NCVGG S++ + + L G M+TYGGMS +
Sbjct: 245 EDQNNSKEFGPTIKEWIKQSGG--EAKLALNCVGGKSSTGIARKLNNNGLMLTYGGMSFQ 302
Query: 289 PVTVSTSSFIFKNWLS-----TDKAEEGRRM----IDRLLGLVQDGKL 327
PVT+ TS +IFKN+ S T+ + + + +++++ ++GKL
Sbjct: 303 PVTIPTSLYIFKNFTSAGFWVTELLKNNKELKTSTLNQIIAWYEEGKL 350
>ETR1_CANTR (Q8WZM3) Enoyl-[acyl-carrier protein] reductase [NADPH,
B-specific] 1, mitochondrial precursor (EC 1.3.1.10)
Length = 386
Score = 210 bits (535), Expect = 4e-54
Identities = 134/348 (38%), Positives = 198/348 (56%), Gaps = 44/348 (12%)
Query: 19 SLRSRILNTHTHAFSSAVSPPSKAIIYESHGQPDAV--TKLVDIPATEVKENDLCVKMLA 76
S+R R+L TH F + ++ ++A++Y HG+P V T+ +I + N++ VK L
Sbjct: 8 SIRPRLLATHNQ-FRTMIT--AQAVLYTQHGEPKDVLFTQSFEIDDDNLAPNEVIVKTLG 64
Query: 77 APINPSDINRIQGVYPVRP---------EPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIP 127
+P+NPSDIN+IQGVYP +P EP A G EG+ EV VGS V+ GDWVIP
Sbjct: 65 SPVNPSDINQIQGVYPSKPAKTTGFGTTEPAAPCGNEGLFEVIKVGSNVSSLEAGDWVIP 124
Query: 128 SPPSFGTWQTYIVKDQNVWHKVNK-----------GVPMEYAATITVNPLTALLMLEDCV 176
S +FGTW+T+ + + + + K+ G+ + ATI+VNPLTA LML V
Sbjct: 125 SHVNFGTWRTHALGNDDDFIKLPNPAQSKANGKPNGLTINQGATISVNPLTAYLMLTHYV 184
Query: 177 TLNSG-DAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFT 235
L G D +QNG TS VG+ Q+ K ++I++IRDRP + EV LK+LGA +V T
Sbjct: 185 KLTPGKDWFIQNGGTSAVGKYASQIGKLLNFNSISVIRDRPNLDEVVASLKELGATQVIT 244
Query: 236 ESELE-------VKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKK 288
E + +K GG E L NCVGG S++ + + L G M+TYGGMS +
Sbjct: 245 EDQNNSREFGPTIKEWIKQSGG--EAKLALNCVGGKSSTGIARKLNNNGLMLTYGGMSFQ 302
Query: 289 PVTVSTSSFIFKNWLS-----TDKAEEGRRM----IDRLLGLVQDGKL 327
PVT+ TS +IFKN+ S T+ + + + +++++ ++GKL
Sbjct: 303 PVTIPTSLYIFKNFTSAGFWVTELLKNNKELKTSTLNQIIAWYEEGKL 350
>ETR1_SCHPO (Q10488) Enoyl-[acyl-carrier protein] reductase [NADPH,
B-specific], mitochondrial precursor (EC 1.3.1.10)
(Mitochondrial respiratory function protein homolog)
Length = 372
Score = 209 bits (532), Expect = 9e-54
Identities = 137/352 (38%), Positives = 196/352 (54%), Gaps = 33/352 (9%)
Query: 40 SKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRP---- 95
+KAI Y +G P V + V + +N + V+ LA+PINPSDIN+IQGVYP +P
Sbjct: 21 AKAIAYSEYGNPKEVLRAVSYNVPKCSKNQVNVRFLASPINPSDINQIQGVYPSKPPFTN 80
Query: 96 -----EPPAVGGYEGVGEVHSVGSAVT-CFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKV 149
+P AV G EG+ EV VG FSPG W I + G+W+T + D V
Sbjct: 81 DVCSSKPSAVAGNEGLVEVVDVGDQFKGTFSPGQWAILGSVNLGSWRTEMNIDGRSLVPV 140
Query: 150 NKGV--PMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIH 207
+K + AAT++VNP TA +L+ V LN GD +Q+GA SMVG IQLAK G
Sbjct: 141 DKSAFPSIAEAATLSVNPCTAYCLLQHVVQLNKGDWFIQDGANSMVGIATIQLAKHFGYK 200
Query: 208 NINIIRDRPGVGEVKERLKDLGADEVFTESEL-EVKNVKS-----LLGGIPEPALGFNCV 261
+IN++R+RP + ++KE+LK LGA V T+ EL + K +K + GG E LG +CV
Sbjct: 201 SINVVRNRPDIEKLKEQLKSLGATIVITDEELMDRKTMKQKVPEWIQGG--EVKLGIDCV 258
Query: 262 GGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN------WLS---TDKAEEGR 312
G A+ + K++ +G TM T+GGMS++P+ V S IFKN W++ ++ EE
Sbjct: 259 SGRVAAEMAKYMSKGATMATFGGMSRQPLPVPVSLLIFKNLKFHGFWVTKWKSEHPEEFL 318
Query: 313 RMIDRLLGLVQDGKLK-YKMELTPFN---DFNTALDKALGKLGSQPKQVIKF 360
++I ++ ++G LK EL D T LD L + K++IKF
Sbjct: 319 KIIHKVEDFYRNGTLKTVNTELVSLKEDADEKTFLDTFLNAIEGHGKKIIKF 370
>ETR1_YEAST (P38071) Enoyl-[acyl-carrier protein] reductase [NADPH,
B-specific] (EC 1.3.1.10) (Mitochondrial respiratory
function protein 1)
Length = 371
Score = 160 bits (406), Expect = 4e-39
Identities = 113/312 (36%), Positives = 168/312 (53%), Gaps = 31/312 (9%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKEN---DLCVKMLAAPINPSDINRIQGVYPVRPE- 96
K++IY +H D TK++ + K++ + +K LA PINPSDIN++QGVYP RPE
Sbjct: 12 KSLIYSTHEVEDC-TKVLSVKNYTPKQDLSQSIVLKTLAFPINPSDINQLQGVYPSRPEK 70
Query: 97 --------PPAVGGYEGVGEVHSV--GSAVTCFSPGDWVIPSPPSFGTWQTY-IVKDQNV 145
P A+ G EGV EV S+ GS+ GD VIP + GTW Y + +
Sbjct: 71 TYDYSTDEPAAIAGNEGVFEVVSLPSGSSKGDLKLGDRVIPLQANQGTWSNYRVFSSSSD 130
Query: 146 WHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSG--DAIVQNGATSMVGQCVIQLAKS 203
KVN + + AAT++VN T ++ D + NS + I+QN TS V + V Q+AK+
Sbjct: 131 LIKVND-LDLFSAATVSVNGCTGFQLVSDYIDWNSNGNEWIIQNAGTSSVSKIVTQVAKA 189
Query: 204 RGIHNINIIRDRPGVGEVKERLKD-LGADEVFTESELEVKN-----VKSLLGGIPEPALG 257
+GI +++IRDR EV + L+D GA +V +ES+ K + +LG L
Sbjct: 190 KGIKTLSVIRDRDNFDEVAKVLEDKYGATKVISESQNNDKTFAKEVLSKILGENARVRLA 249
Query: 258 FNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKN------WLSTDKAEEG 311
N VGG S++ + + L M+TYGGMSK+PVT+ TS IFK W++ +
Sbjct: 250 LNSVGGKSSASIARKLENNALMLTYGGMSKQPVTLPTSLHIFKGLTSKGYWVTEKNKKNP 309
Query: 312 RRMIDRLLGLVQ 323
+ ID + ++
Sbjct: 310 QSKIDTISDFIK 321
>QOR_YEAST (P38230) Probable quinone oxidoreductase (EC 1.6.5.5)
(NADPH:quinone reductase)
Length = 334
Score = 77.4 bits (189), Expect = 5e-14
Identities = 93/330 (28%), Positives = 138/330 (41%), Gaps = 31/330 (9%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
K I+ + G D + K D P + E +L +K +N + +G+YP E P V
Sbjct: 10 KVILIDEIGGYDVI-KYEDYPVPSISEEELLIKNKYTGVNYIESYFRKGIYPC--EKPYV 66
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYI-VKDQNVWHKVNKGVPME--- 156
G E G V + G VT F GD V + S T+ Y + Q K+ KG E
Sbjct: 67 LGREASGTVVAKGKGVTNFEVGDQV--AYISNSTFAQYSKISSQGPVMKLPKGTSDEELK 124
Query: 157 -YAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDR 215
YAA + + LTAL + + GD ++ A VG + QL K +G H I +
Sbjct: 125 LYAAGL-LQVLTALSFTNEAYHVKKGDYVLLFAAAGGVGLILNQLLKMKGAHTIAV---- 179
Query: 216 PGVGEVKERLKDLGADEVFTESELEV-KNVKSLLGGIPEPALGFNCVGGNSASLVLKFLR 274
E + K+ GA+ + S+ ++ + V G A F+ VG ++ + L L+
Sbjct: 180 ASTDEKLKIAKEYGAEYLINASKEDILRQVLKFTNGKGVDA-SFDSVGKDTFEISLAALK 238
Query: 275 RGGTMVTYGGMSKKPVTVSTSSFIFKN--------WLSTDKAEEGRRMIDRLLGLVQDGK 326
R G V++G S S + KN + EE + D GLV K
Sbjct: 239 RKGVFVSFGNASGLIPPFSITRLSPKNITLVRPQLYGYIADPEEWKYYSDEFFGLVNSKK 298
Query: 327 LKYKMELT-PFNDFNTAL-----DKALGKL 350
L K+ T P D+ TA K +GKL
Sbjct: 299 LNIKIYKTYPLRDYRTAAADIESRKTVGKL 328
>QOR_PSEAE (P43903) Quinone oxidoreductase (EC 1.6.5.5)
(NADPH:quinone reductase)
Length = 325
Score = 77.4 bits (189), Expect = 5e-14
Identities = 81/304 (26%), Positives = 137/304 (44%), Gaps = 21/304 (6%)
Query: 40 SKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPA 99
+K I + ++G P+ V + D E ++ V+ A +N D G+YP P P+
Sbjct: 2 AKRIQFAAYGGPE-VLEYRDYQPAEPGPREVRVRNRAIGLNFIDTYYRSGLYPA-PGLPS 59
Query: 100 VGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTW-QTYIVKDQNVWHKVNKGVPMEYA 158
G EG GEV +VGS VT F GD V + G + + +++ ++ + H + G+ E A
Sbjct: 60 GLGSEGAGEVEAVGSEVTRFKVGDRVAYATGPLGAYSELHVLAEEKLVH-LPDGIDFEQA 118
Query: 159 ATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGV 218
A + + LT +L L G+ I+ + A VG Q AK+ G+ I +
Sbjct: 119 AAVMLKGLTTQYLLRQTYELRGGETILFHAAAGGVGLFACQWAKALGVQLIGTVSS---- 174
Query: 219 GEVKERLKDLGADEVFTESELEV-KNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGG 277
E + GA E S V + V L G P + ++ VG ++ L + G
Sbjct: 175 PEKARLARQHGAWETIDYSHENVARRVLELTDGKKCPVV-YDSVGKDTWETSLDCVAPRG 233
Query: 278 TMVTYGGMSKKPVT--------VSTSSFIFKNWLST--DKAEEGRRMIDRLLGLVQDGKL 327
+V++G S PVT S ++ + L + D E+ + M D L GL++ G +
Sbjct: 234 LLVSFGNAS-GPVTGVNLGILSQKGSLYVTRPTLGSYADTPEKLQAMADELFGLIERGDI 292
Query: 328 KYKM 331
+ ++
Sbjct: 293 RIEI 296
>QORL_ARATH (Q9ZUC1) Quinone oxidoreductase-like protein At1g23740,
chloroplast precursor (EC 1.-.-.-)
Length = 386
Score = 73.2 bits (178), Expect = 1e-12
Identities = 71/268 (26%), Positives = 115/268 (42%), Gaps = 27/268 (10%)
Query: 34 SAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPV 93
+++ KA +Y +G D + +I E+KE+ + +K++AA +NP D R QG +
Sbjct: 72 ASIPKEMKAWVYSDYGGVDVLKLESNIVVPEIKEDQVLIKVVAAALNPVDAKRRQGKFKA 131
Query: 94 RPEP-PAVGGYEGVGEVHSVGSAVTCFSPGDWV--------IPSPPSFGTWQTYIVKDQN 144
P P V GY+ G V VGSAV GD V + P FG+ Y ++
Sbjct: 132 TDSPLPTVPGYDVAGVVVKVGSAVKDLKEGDEVYANVSEKALEGPKQFGSLAEYTAVEEK 191
Query: 145 VWHKVNKGVPMEYAATITVNPLTALLMLEDCV--TLNSGDAIVQNGATSMVGQCVIQLAK 202
+ K + AA + PL E V ++G +I+ VG VIQLAK
Sbjct: 192 LLALKPKNIDFAQAAGL---PLAIETADEGLVRTEFSAGKSILVLNGAGGVGSLVIQLAK 248
Query: 203 SRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEP-ALGFNCV 261
++ + + E E ++ LGAD L + K + +P+ + F+ +
Sbjct: 249 H--VYGASKVAATAST-EKLELVRSLGAD-------LAIDYTKENIEDLPDKYDVVFDAI 298
Query: 262 GGNSASLVLKFLRRGGTMVTYGGMSKKP 289
G +K ++ GG +V G P
Sbjct: 299 G--MCDKAVKVIKEGGKVVALTGAVTPP 324
>QOR_HUMAN (Q08257) Quinone oxidoreductase (EC 1.6.5.5)
(NADPH:quinone reductase) (Zeta-crystallin)
Length = 329
Score = 71.2 bits (173), Expect = 4e-12
Identities = 64/277 (23%), Positives = 117/277 (42%), Gaps = 15/277 (5%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
+A+ G P+ + DI K++ + +K+ A +NP + G Y +P P
Sbjct: 9 RAVRVFEFGGPEVLKLRSDIAVPIPKDHQVLIKVHACGVNPVETYIRSGTYSRKPLLPYT 68
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAAT 160
G + G + +VG + F GD V S G + Y + + +K+ + + + A
Sbjct: 69 PGSDVAGVIEAVGDNASAFKKGDRVFTSSTISGGYAEYALAADHTVYKLPEKLDFKQGAA 128
Query: 161 ITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGE 220
I + TA L + +G++++ +GA+ VG Q+A++ G+ I G E
Sbjct: 129 IGIPYFTAYRALIHSACVKAGESVLVHGASGGVGLAACQIARAYGLK----ILGTAGTEE 184
Query: 221 VKERLKDLGADEVFTESELE-VKNVKSLLG--GIPEPALGFNCVGGNSASLVLKFLRRGG 277
++ + GA EVF E+ + +K +G GI + + + S L L GG
Sbjct: 185 GQKIVLQNGAHEVFNHREVNYIDKIKKYVGEKGID---IIIEMLANVNLSKDLSLLSHGG 241
Query: 278 TMVTYGG-----MSKKPVTVSTSSFIFKNWLSTDKAE 309
++ G ++ + SS I S+ K E
Sbjct: 242 RVIVVGSRGTIEINPRDTMAKESSIIGVTLFSSTKEE 278
>QOR_LAMGU (Q28452) Quinone oxidoreductase (EC 1.6.5.5)
(NADPH:quinone reductase) (Zeta-crystallin)
Length = 330
Score = 70.5 bits (171), Expect = 6e-12
Identities = 62/256 (24%), Positives = 110/256 (42%), Gaps = 12/256 (4%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
+AI G P+ + D+ +E+ + +K+ A +NP D G Y +P P
Sbjct: 9 RAIRVSEFGGPEVLKLQSDVAVPIPEEHQVLIKVQACGVNPVDTYIRSGTYSRKPRLPYT 68
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAAT 160
G + G + +VG V+ F GD V + G + Y + + +K+ + + A
Sbjct: 69 PGLDVAGLIEAVGERVSAFKKGDRVFTTSTVSGGYAEYALAADHTVYKLPGELDFQKGAA 128
Query: 161 ITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGE 220
I V TA L +G++++ +GA+ VG Q+A++ + G E
Sbjct: 129 IGVPYFTAYRALLHSACAKAGESVLVHGASGGVGLAACQIARACCFK----VLGTAGTEE 184
Query: 221 VKERLKDLGADEVFTESE-LEVKNVKSLLG--GIPEPALGFNCVGGNSASLVLKFLRRGG 277
+ + GA EVF E + + +K +G GI + + + S L L +GG
Sbjct: 185 GQRVVLQNGAHEVFNHREDINIDKIKKSVGEKGID---VIIEMLANVNLSNDLNLLSQGG 241
Query: 278 TMVTYGGMSKKPVTVS 293
++ G SK PV ++
Sbjct: 242 RVIIVG--SKGPVEIN 255
>ADH1_KLUMA (Q07288) Alcohol dehydrogenase 1 (EC 1.1.1.1)
Length = 348
Score = 69.3 bits (168), Expect = 1e-11
Identities = 69/279 (24%), Positives = 119/279 (41%), Gaps = 40/279 (14%)
Query: 35 AVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVR 94
A+ K +I+ HG + DIP + K N+L + + + + +D++ QG +P+
Sbjct: 2 AIPETQKGVIFYEHG---GELQYKDIPVPKPKPNELLINVKYSGVCHTDLHAWQGDWPLD 58
Query: 95 PEPPAVGGYEGVGEVHSVGSAVTCFSPGDWV-------------------IPSPPSF--- 132
+ P VGG+EG G V ++G VT + GD+ P+ P
Sbjct: 59 TKLPLVGGHEGAGIVVAMGENVTGWEIGDYAGIKWLNGSCMSCEECELSNEPNCPKADLS 118
Query: 133 -----GTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQN 187
G++Q Y D ++ K V + A I +T L+ + +GD + +
Sbjct: 119 GYTHDGSFQQYATADAVQAARIPKNVDLAEVAPILCAGVTVYKALKSA-HIKAGDWVAIS 177
Query: 188 GATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERL-KDLGADEV--FTESELEVKNV 244
GA +G IQ AK+ G + I G+ K +L K+LG + FT+++ V V
Sbjct: 178 GACGGLGSLAIQYAKAMGYRVLGI-----DAGDEKAKLFKELGGEYFIDFTKTKDMVAEV 232
Query: 245 KSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYG 283
G+ + + V + S + + R GT+V G
Sbjct: 233 IEATNGVAHAVINVS-VSEAAISTSVLYTRSNGTVVLVG 270
>VAT1_MOUSE (Q62465) Synaptic vesicle membrane protein VAT-1 homolog
(EC 1.-.-.-)
Length = 406
Score = 68.2 bits (165), Expect = 3e-11
Identities = 71/259 (27%), Positives = 113/259 (43%), Gaps = 17/259 (6%)
Query: 31 AFSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKEN--DLCVKMLAAPINPSDINRIQ 88
A +SA +PP + ++ G D V KL PA L +++ A +N +D+ Q
Sbjct: 52 ASASASAPPLRCLVLTGFGGYDKV-KLQSRPAVPPAPGPGQLTLRVRACGLNFADLMGRQ 110
Query: 89 GVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHK 148
G+Y P P G EG G V +VG V GD V+ S G WQ +
Sbjct: 111 GLYDRLPPLPVTPGMEGAGVVVAVGEGVGDRKAGDRVMVLNRS-GMWQEEVTVPSAQTFL 169
Query: 149 VNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHN 208
+ + + E AA + VN +TA ++L D L G +++ + A VG +QL R + N
Sbjct: 170 MPEAMTFEEAAALLVNYITAYMVLFDFGNLRPGHSVLVHMAAGGVGMAALQLC--RTVEN 227
Query: 209 INIIRDRPGVGEVKERLKDLGA----DEVFTESELEVKNVKSLLGGIPEPALGFNCVGGN 264
+ + E LK+ G D T+ E+K + G+ + + +GG+
Sbjct: 228 VTVF--GTASASKHEVLKENGVTHPIDYHTTDYVDEIKKISP--KGVD---IVMDPLGGS 280
Query: 265 SASLVLKFLRRGGTMVTYG 283
+ L+ G +VTYG
Sbjct: 281 DTAKGYHLLKPMGKVVTYG 299
>VAT1_HUMAN (Q99536) Synaptic vesicle membrane protein VAT-1 homolog
(EC 1.-.-.-)
Length = 393
Score = 68.2 bits (165), Expect = 3e-11
Identities = 71/260 (27%), Positives = 114/260 (43%), Gaps = 18/260 (6%)
Query: 31 AFSSAVSPPS-KAIIYESHGQPDAVTKLVDIPATEVKEN--DLCVKMLAAPINPSDINRI 87
A ++A SPP + ++ G D V KL PA L +++ A +N +D+
Sbjct: 38 AAAAAASPPLLRCLVLTGFGGYDKV-KLQSRPAAPPAPGPGQLTLRLRACGLNFADLMAR 96
Query: 88 QGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWH 147
QG+Y P P G EG G V +VG V+ GD V+ S G WQ +
Sbjct: 97 QGLYDRLPPLPVTPGMEGAGVVIAVGEGVSDRKAGDRVMVLNRS-GMWQEEVTVPSVQTF 155
Query: 148 KVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIH 207
+ + + E AA + VN +TA ++L D L G +++ + A VG +QL R +
Sbjct: 156 LIPEAMTFEEAAALLVNYITAYMVLFDFGNLQPGHSVLVHMAAGGVGMAAVQLC--RTVE 213
Query: 208 NINIIRDRPGVGEVKERLKDLGA----DEVFTESELEVKNVKSLLGGIPEPALGFNCVGG 263
N+ + E LK+ G D T+ E+K + G+ + + +GG
Sbjct: 214 NVTVF--GTASASKHEALKENGVTHPIDYHTTDYVDEIKKISP--KGVD---IVMDPLGG 266
Query: 264 NSASLVLKFLRRGGTMVTYG 283
+ + L+ G +VTYG
Sbjct: 267 SDTAKGYNLLKPMGKVVTYG 286
>QOR_BOVIN (O97764) Zeta-crystallin
Length = 330
Score = 68.2 bits (165), Expect = 3e-11
Identities = 54/244 (22%), Positives = 104/244 (42%), Gaps = 6/244 (2%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
+AI G P+ + D+ K++ + +K+ A +NP D G + ++P P
Sbjct: 9 RAIRVFEFGGPEVLKLQSDVAVPIPKDHQVLIKVQACGVNPVDTYIRSGTHNIKPLLPYT 68
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAAT 160
G++ G + +VG +V+ F GD V + G + Y + + + + + + + A
Sbjct: 69 PGFDVAGIIEAVGESVSAFKKGDRVFTTRTISGGYAEYALAADHTVYTLPEKLDFKQGAA 128
Query: 161 ITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGE 220
I + TA L + G++++ +GA+ VG Q+A++ G+ + G
Sbjct: 129 IGIPYFTAYRALLYSAPVKPGESVLVHGASGGVGIAACQIARAYGLKVLGTASTEEGQKI 188
Query: 221 VKERLKDLGADEVFTESELE-VKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTM 279
V E GA +VF E + +K +G + + + S L L GG +
Sbjct: 189 VLEN----GAHKVFNHKEANYIDKIKKSVGEKGVDVI-IEMLANVNLSNDLNLLSHGGRV 243
Query: 280 VTYG 283
+ G
Sbjct: 244 IVVG 247
>KF76_HUMAN (Q9HCJ6) Probable oxidoreductase KIAA1576 (EC 1.-.-.-)
Length = 419
Score = 67.8 bits (164), Expect = 4e-11
Identities = 62/251 (24%), Positives = 108/251 (42%), Gaps = 22/251 (8%)
Query: 64 EVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGD 123
E ++ +L +++ A +N D+ QG P+ P V G+E G V ++G +V + GD
Sbjct: 65 EPQDGELKIRVKACGLNFIDLMVRQGNIDNPPKTPLVPGFECSGIVEALGDSVKGYEIGD 124
Query: 124 WVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDA 183
V+ ++ W + +K+ + AA +N +TA +ML + L G +
Sbjct: 125 RVMAFV-NYNAWAEVVCTPVEFVYKIPDDMSFSEAAAFPMNFVTAYVMLFEVANLREGMS 183
Query: 184 IVQNGATSMVGQCVIQLAKSRGIHNINIIRD-----RPGVGEVKERLKDLGADEVFTESE 238
++ + A VGQ V QL + + N+ + + + L D AD V
Sbjct: 184 VLVHSAGGGVGQAVAQLCST--VPNVTVFGTASTFKHEAIKDSVTHLFDRNADYVQEVKR 241
Query: 239 LEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSF- 297
+ + V +L +C+ G++ L L+ GT + YG S VT T SF
Sbjct: 242 ISAEGVDIVL----------DCLCGDNTGKGLSLLKPLGTYILYG--SSNMVTGETKSFF 289
Query: 298 -IFKNWLSTDK 307
K+W +K
Sbjct: 290 SFAKSWWQVEK 300
>QOR_CAVPO (P11415) Quinone oxidoreductase (EC 1.6.5.5)
(NADPH:quinone reductase) (Zeta-crystallin)
Length = 329
Score = 67.0 bits (162), Expect = 7e-11
Identities = 56/244 (22%), Positives = 104/244 (41%), Gaps = 6/244 (2%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
+AI G P+ + D+ K++ + +K+ A INP + G Y P P
Sbjct: 9 RAIRVFEFGGPEVLKVQSDVAVPIPKDHQVLIKVHACGINPVETYIRSGTYTRIPLLPYT 68
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAAT 160
G + G V S+G+ V+ F GD V + G + Y + + +++ + + A
Sbjct: 69 PGTDVAGVVESIGNDVSAFKKGDRVFTTSTISGGYAEYALASDHTVYRLPEKLDFRQGAA 128
Query: 161 ITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGE 220
I + TA L +G++++ +GA+ VG Q+A++ G+ + G E
Sbjct: 129 IGIPYFTACRALFHSARAKAGESVLVHGASGGVGLAACQIARAYGLK----VLGTAGTEE 184
Query: 221 VKERLKDLGADEVFTESELE-VKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTM 279
++ + GA EVF + + +K +G + + + S LK L GG +
Sbjct: 185 GQKVVLQNGAHEVFNHRDAHYIDEIKKSIGEKGVDVI-IEMLANVNLSNDLKLLSCGGRV 243
Query: 280 VTYG 283
+ G
Sbjct: 244 IIVG 247
>KF76_MOUSE (Q80TB8) Probable oxidoreductase KIAA1576 (EC 1.-.-.-)
Length = 417
Score = 67.0 bits (162), Expect = 7e-11
Identities = 62/251 (24%), Positives = 107/251 (41%), Gaps = 22/251 (8%)
Query: 64 EVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGD 123
E ++ +L +++ A +N D+ QG P+ P V G+E G V ++G +V + GD
Sbjct: 63 EPQDGELKIRVKACGLNFIDLMVRQGNIDNPPKTPLVPGFECSGIVEALGDSVKGYEIGD 122
Query: 124 WVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDA 183
V+ ++ W + +K+ + AA +N +TA ML + L G +
Sbjct: 123 RVMAFV-NYNAWAEVVCTPVEFVYKIPDDMSFSEAAAFPMNFVTAYTMLFEIANLREGMS 181
Query: 184 IVQNGATSMVGQCVIQLAKSRGIHNINIIRD-----RPGVGEVKERLKDLGADEVFTESE 238
++ + A VGQ V QL + + N+ + + + L D AD V
Sbjct: 182 VLVHSAGGGVGQAVAQLCST--VPNVTVFGTASTFKHEAIKDSVTHLFDRNADYVQEVKR 239
Query: 239 LEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSF- 297
+ + V +L +C+ G++ L L+ GT + YG S VT T SF
Sbjct: 240 ISAEGVDIVL----------DCLCGDNTGKGLSLLKPLGTYILYG--SSNMVTGETKSFF 287
Query: 298 -IFKNWLSTDK 307
K+W +K
Sbjct: 288 SFAKSWWQVEK 298
>ADH1_BACST (P12311) Alcohol dehydrogenase (EC 1.1.1.1) (ADH-T)
Length = 337
Score = 67.0 bits (162), Expect = 7e-11
Identities = 79/335 (23%), Positives = 138/335 (40%), Gaps = 40/335 (11%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
KA + E +P ++ ++ ++ ++ V++ A + +D++ G +PV+P+ P +
Sbjct: 2 KAAVVEQFKKP---LQVKEVEKPKISYGEVLVRIKACGVCHTDLHAAHGDWPVKPKLPLI 58
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWV-IPSPPS--------FGTWQTYIVKDQNVWHKVNK 151
G+EGVG + VG VT GD V IP S +T + QN + V+
Sbjct: 59 PGHEGVGVIEEVGPGVTHLKVGDRVGIPWLYSACGHCDYCLSGQETLCERQQNAGYSVDG 118
Query: 152 GVPMEY--AATITVNPLTALLMLED-----CVTLNSGDAIVQNGA----------TSMVG 194
G EY AA V + L E+ C + + A+ GA +G
Sbjct: 119 GY-AEYCRAAADYVVKIPDNLSFEEAAPIFCAGVTTYKALKVTGAKPGEWVAIYGIGGLG 177
Query: 195 QCVIQLAKSRGIHNINIIRDRPGVGEVK-ERLKDLGADEVFTESELEVKN-VKSLLGGIP 252
+Q AK+ G++ + + +G+ K E K LGAD V + +K +GG+
Sbjct: 178 HVAVQYAKAMGLNVVAV-----DLGDEKLELAKQLGADLVVNPKHDDAAQWIKEKVGGV- 231
Query: 253 EPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKNWLSTDKAEEGR 312
A V + K +RRGG V G+ + + + + R
Sbjct: 232 -HATVVTAVSKAAFESAYKSIRRGGACVLV-GLPPEEIPIPIFDTVLNGVKIIGSIVGTR 289
Query: 313 RMIDRLLGLVQDGKLKYKMELTPFNDFNTALDKAL 347
+ + L +GK+K +E+ P + N D+ L
Sbjct: 290 KDLQEALQFAAEGKVKTIVEVQPLENINDVFDRML 324
>QOR_MOUSE (P47199) Quinone oxidoreductase (EC 1.6.5.5)
(NADPH:quinone reductase) (Zeta-crystallin)
Length = 331
Score = 66.6 bits (161), Expect = 9e-11
Identities = 61/289 (21%), Positives = 119/289 (41%), Gaps = 10/289 (3%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
+AI G P+ + D+ + + + +K+ A +NP + G Y +P P
Sbjct: 9 RAIRVFEFGGPEVLKLQSDVVVPVPQSHQVLIKVHACGVNPVETYIRSGAYSRKPALPYT 68
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAAT 160
G + G + SVG V+ F GD V G + + + + + + + + A
Sbjct: 69 PGSDVAGIIESVGDKVSAFKKGDRVFCYSTVSGGYAEFALAADDTIYPLPETLNFRQGAA 128
Query: 161 ITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGE 220
+ + TA L +G++++ +GA+ VG Q+A++ G+ + G E
Sbjct: 129 LGIPYFTACRALFHSARARAGESVLVHGASGGVGLATCQIARAHGLK----VLGTAGSEE 184
Query: 221 VKERLKDLGADEVFTESELE-VKNVKSLLGGIPEPA-LGFNCVGGNSASLVLKFLRRGGT 278
K+ + GA EVF E + +K +G + + + + S LK L GG
Sbjct: 185 GKKLVLQNGAHEVFNHKEANYIDKIKMSVGDKDKGVDVIIEMLANENLSNDLKLLSHGGR 244
Query: 279 MVTYGGMSKKPVTVSTSSFIFK--NWLSTDKAEEGRRMIDRLLGLVQDG 325
+V G + P+ ++ + K + + + + + GL+Q G
Sbjct: 245 VVVVG--CRGPIEINPRDTMAKETSIIGVSLSSSTKEEFQQFAGLLQAG 291
>ADH3_BACST (P42328) Alcohol dehydrogenase (EC 1.1.1.1) (ADH-HT)
Length = 339
Score = 66.2 bits (160), Expect = 1e-10
Identities = 82/334 (24%), Positives = 136/334 (40%), Gaps = 38/334 (11%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
KA + E +P K+ ++ + ++ V++ A + +D++ G +PV+P+ P +
Sbjct: 2 KAAVVEQFKEP---LKIKEVEKPTISYGEVLVRIKACGVCHTDLHAAHGDWPVKPKLPLI 58
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWV-IPSPPSFGTWQTYIVKDQNVW--HKVNKGVPM-- 155
G+EGVG V VG VT GD V IP S Y + Q H+ N G +
Sbjct: 59 PGHEGVGIVEEVGPGVTHLKVGDRVGIPWLYSACGHCDYCLSGQETLCEHQKNAGYSVDG 118
Query: 156 ---EY--AATITVNPLTALLMLED-----CVTLNSGDAIVQNGA----------TSMVGQ 195
EY AA V + L E+ C + + A+ GA +G
Sbjct: 119 GYAEYCRAAADYVVKIPDNLSFEEAAPIFCAGVTTYKALKVTGAKPGEWVAIYGIGGLGH 178
Query: 196 CVIQLAKSRGIHNINIIRDRPGVGEVK-ERLKDLGADEVFTE-SELEVKNVKSLLGGIPE 253
+Q AK+ G++ + + +G+ K E K+LGAD V E K +K +GG+
Sbjct: 179 VAVQYAKAMGLNVVAV-----DIGDEKLELAKELGADLVVNPLKEDAAKFMKEKVGGV-- 231
Query: 254 PALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKNWLSTDKAEEGRR 313
A V + +RRGG V G+ + + + + R+
Sbjct: 232 HAAVVTAVSKPAFQSAYNSIRRGGACVLV-GLPPEEMPIPIFDTVLNGIKIIGSIVGTRK 290
Query: 314 MIDRLLGLVQDGKLKYKMELTPFNDFNTALDKAL 347
+ L +GK+K +E+ P N D+ L
Sbjct: 291 DLQEALQFAAEGKVKTIIEVQPLEKINEVFDRML 324
>IC11_TRIHA (P34055) Protein indc11
Length = 339
Score = 64.7 bits (156), Expect = 3e-10
Identities = 62/269 (23%), Positives = 112/269 (41%), Gaps = 13/269 (4%)
Query: 39 PSKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPP 98
P++ ++ ++G P + V ++ + VK+L A +D+N GVYP++ PP
Sbjct: 4 PNRKVVITAYGPPSTALQFVTEDLPPPPKDHVQVKILYAGFAGADVNMRLGVYPMQSAPP 63
Query: 99 AVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYA 158
GY G V G F PG ++ + + + YI + + GV + A
Sbjct: 64 FTPGYCFAGRVSVNGPGSGKFEPGT-LVTALTKYDSDAEYINIPEKYLLAIPDGVDPKVA 122
Query: 159 ATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGV 218
A + V+ TA M+ ++ G + +G + VGQ V+ L+ +G +R
Sbjct: 123 AALPVDWSTAYGMVHRAAKVSEGQRVFIHGISGAVGQAVMYLSLLQGATVYGTASERNHA 182
Query: 219 GEVKERLKDLGADEVFTESELEVKNVKSLLGGIPE--PALGFNCVGGNSASLVLK----F 272
LK+ GA ++ + +K LGG+ ALGF + + L K
Sbjct: 183 A-----LKEAGAHPYLYTNKDWIAAMKD-LGGVHAVFDALGFESFDESYSILTPKERSVV 236
Query: 273 LRRGGTMVTYGGMSKKPVTVSTSSFIFKN 301
+ G + G ++ + + +FKN
Sbjct: 237 VAYGNNLSNLTGAKRRSPWIPMAKLLFKN 265
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.135 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,007,703
Number of Sequences: 164201
Number of extensions: 1944177
Number of successful extensions: 5095
Number of sequences better than 10.0: 219
Number of HSP's better than 10.0 without gapping: 140
Number of HSP's successfully gapped in prelim test: 79
Number of HSP's that attempted gapping in prelim test: 4879
Number of HSP's gapped (non-prelim): 248
length of query: 360
length of database: 59,974,054
effective HSP length: 111
effective length of query: 249
effective length of database: 41,747,743
effective search space: 10395188007
effective search space used: 10395188007
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 66 (30.0 bits)
Medicago: description of AC135230.4


