

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.3 + phase: 0 /pseudo
(605 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
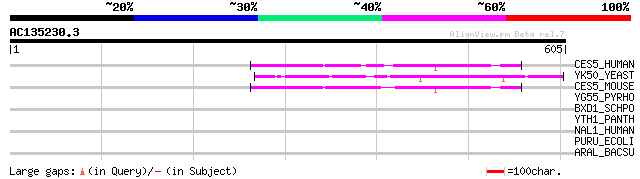
CES5_HUMAN (Q9BXW7) Cat eye syndrome critical region protein 5 p... 151 5e-36
YK50_YEAST (P36151) Hypothetical 39.4 kDa protein in MET1-SIS2 i... 149 2e-35
CES5_MOUSE (Q91WM2) Cat eye syndrome critical region protein 5 h... 144 9e-34
YG55_PYRHO (O59346) Putative HAD-hydrolase PH1655 (EC 3.-.-.-) 34 0.97
BXD1_SCHPO (Q9UUG1) Brix domain containing protein 1 homolog 34 0.97
YTH1_PANTH (P46351) Hypothetical 45.4 kDa protein in thiaminase ... 33 2.8
NAL1_HUMAN (Q9C000) NACHT-, LRR- and PYD-containing protein 2 (D... 32 4.8
PURU_ECOLI (P37051) Formyltetrahydrofolate deformylase (EC 3.5.1... 31 8.2
ARAL_BACSU (P94526) Arabinose operon protein araL 31 8.2
>CES5_HUMAN (Q9BXW7) Cat eye syndrome critical region protein 5
precursor
Length = 423
Score = 151 bits (381), Expect = 5e-36
Identities = 94/301 (31%), Positives = 149/301 (49%), Gaps = 28/301 (9%)
Query: 263 PSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASELS 322
P+FG DIDGV++ G+ + + A R+L N G L+ P VF+TN G I + +A ELS
Sbjct: 45 PTFGFLLDIDGVLVRGHRVIPAALKAFRRLVNSQGQLRVPVVFVTNAGNILQHSKAQELS 104
Query: 323 ELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYASR 382
LLG V A QV+ HSP + L + + +K ++ +G+G GF+NV+++D
Sbjct: 105 ALLGCEVDADQVILSHSPMK-LFSEYHEKRMLVSGQGPVMENAQGLGFRNVVTVDELRMA 163
Query: 383 FENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSRDVQVLCD 442
F PL +L TT P+ D R++ ++ +PV W +Q++ D
Sbjct: 164 F----PLLDMVDLERRLKTTPLPRND---------FPRIEGVLLLGEPVRWETSLQLIMD 210
Query: 443 ILKTGGLPGRNVGTQPHLYF----ANDDLEYQTKFPSERLGMGAFRIALESIFNRTHHHS 498
+L + G PG + T P+ + +N DL + + R G G F + LE+I+ +
Sbjct: 211 VLLSNGSPGAGLATPPYPHLPVLASNMDLLWMAEAKMPRFGHGTFLLCLETIYQKVTGKE 270
Query: 499 LEYT-CFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMIGDNPAVDIRG 557
L Y GKP ++ AE ++++ R A P + LY +GDNP D+ G
Sbjct: 271 LRYEGLMGKPSILTYQYAEDLIRRQAER---------RGWAAPIRKLYAVGDNPMSDVYG 321
Query: 558 A 558
A
Sbjct: 322 A 322
>YK50_YEAST (P36151) Hypothetical 39.4 kDa protein in MET1-SIS2
intergenic region
Length = 352
Score = 149 bits (377), Expect = 2e-35
Identities = 107/351 (30%), Positives = 170/351 (47%), Gaps = 32/351 (9%)
Query: 268 AFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASELSELLGL 327
AFDIDGV+ G P+ G+ AL KL N + K PY+ LTNGGG E R +S L +
Sbjct: 17 AFDIDGVLFRGKKPIAGASDAL-KLLNRN---KIPYILLTNGGGFSERARTEFISSKLDV 72
Query: 328 NVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYASRFENID 387
+VS Q++Q H+P++ LVN++ I+A G V YGF++V+ +I
Sbjct: 73 DVSPLQIIQSHTPYKSLVNKY--SRILAVGTPSVRGVAEGYGFQDVVHQTDIVRYNRDIA 130
Query: 388 PLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSRDVQVLCDILKT- 446
P + L+ Q ++ P D+ +++ A + +DP DW+ D+Q++ D + +
Sbjct: 131 PF-------SGLSDEQVMEYSRDIP--DLTTKKFDAVLVFNDPHDWAADIQIISDAINSE 181
Query: 447 ----GGLPGRNVGTQP-HLYFANDDLEYQTKFPSERLGMGAFRIALESIFNRTHHHSLEY 501
L G +YF+N DL + + R G GAFR+ + ++ + L+
Sbjct: 182 NGMLNTLRNEKSGKPSIPIYFSNQDLLWANPYKLNRFGQGAFRLLVRRLYLELNGEPLQD 241
Query: 502 TCFGKPHPSVFKNAEIVLQKHVPRVD-EDSYDINHK--------NAQPFQTLYMIGDNPA 552
GKP + A VL R+ + + K + PF ++M+GDNPA
Sbjct: 242 YTLGKPTKLTYDFAHHVLIDWEKRLSGKIGQSVKQKLPLLGTKPSTSPFHAVFMVGDNPA 301
Query: 553 VDIRGARQTGHPWFSILTRTGVFKGKENHDKFPADLVVDTVEEAVDYILAK 603
DI GA+ G W S L +TGV+ ++ + L+V+ V +AV L K
Sbjct: 302 SDIIGAQNYG--WNSCLVKTGVYNEGDDLKECKPTLIVNDVFDAVTKTLEK 350
>CES5_MOUSE (Q91WM2) Cat eye syndrome critical region protein 5
homolog precursor
Length = 419
Score = 144 bits (362), Expect = 9e-34
Identities = 91/301 (30%), Positives = 149/301 (49%), Gaps = 29/301 (9%)
Query: 263 PSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASELS 322
P+FG+ FDIDGV++ G+ + + A KL N G L+ P VF+TN G I + +A ELS
Sbjct: 45 PTFGLLFDIDGVLVRGHRVIPAALEAFSKLVNSQGQLRVPVVFVTNAGNILQHNKAQELS 104
Query: 323 ELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYASR 382
+LL V QV+ HSP + L ++ K ++ +G+G GF+NV++ID
Sbjct: 105 DLLRCKVDPDQVILSHSPMK-LFLQYHSKQMLVSGQGPLVENARALGFQNVVTIDELRLA 163
Query: 383 FENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSRDVQVLCD 442
F +D + ++ T + P ++ ++ +PV W ++Q++ D
Sbjct: 164 FPELDMVDLQRRPKTMRLRSDFP--------------AIEGVLLLGEPVRWETNLQLIMD 209
Query: 443 ILKTGGLPGRNVGTQPHLYF----ANDDLEYQTKFPSERLGMGAFRIALESIFNRTHHHS 498
+L + G PG + T P+ + +N DL + + R G G F + LE+I+ + +
Sbjct: 210 VLLSNGHPGTGLATAPYPHLPVLASNMDLLWMAEAKMPRFGHGTFLLCLETIYRKITGNE 269
Query: 499 LEYT-CFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMIGDNPAVDIRG 557
L+Y GKP ++ AE V+++ R A P + LY IGDNP D+ G
Sbjct: 270 LKYEGLMGKPSILTYQYAEDVIRQQAER---------RGWAAPIRKLYAIGDNPMSDVYG 320
Query: 558 A 558
A
Sbjct: 321 A 321
>YG55_PYRHO (O59346) Putative HAD-hydrolase PH1655 (EC 3.-.-.-)
Length = 241
Score = 34.3 bits (77), Expect = 0.97
Identities = 34/123 (27%), Positives = 53/123 (42%), Gaps = 27/123 (21%)
Query: 484 RIALESIFNRTHHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQT 543
R+ L+ F H ++ KPHP +FK A + N +P +
Sbjct: 130 RLELDDFFE--HVIISDFEGVKKPHPKIFKKA-----------------LKAFNVKPEEA 170
Query: 544 LYMIGDNPAVDIRGARQTGHP--WFSILTRTGVFKGKENHDKFPADLVVDTVEEAVDYIL 601
L M+GD DI GA++ G WF R G +E + AD +D +E ++ +L
Sbjct: 171 L-MVGDRLYSDIYGAKRVGMKTVWF----RYGKHSERELEYRKYADYEIDNLESLLE-VL 224
Query: 602 AKE 604
A+E
Sbjct: 225 ARE 227
>BXD1_SCHPO (Q9UUG1) Brix domain containing protein 1 homolog
Length = 317
Score = 34.3 bits (77), Expect = 0.97
Identities = 27/97 (27%), Positives = 43/97 (43%), Gaps = 11/97 (11%)
Query: 334 VLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYASRFENIDPL--AP 391
V H +R + + F D +GEP + G VI + A ++ + PL
Sbjct: 146 VFDAHPTYRHIKSLFLDFF-----RGEPIQKLDSAGLSYVIVVSAAEAQEDETKPLPLVH 200
Query: 392 YKKWTTKLATTQNP----KFDESGPQIDVFSERVQAA 424
++ + TKL T+ + +E GP+ID RVQ A
Sbjct: 201 FRVYGTKLLKTKTNLPRVELEEMGPRIDFNIRRVQPA 237
>YTH1_PANTH (P46351) Hypothetical 45.4 kDa protein in thiaminase I
5'region
Length = 413
Score = 32.7 bits (73), Expect = 2.8
Identities = 21/67 (31%), Positives = 37/67 (54%), Gaps = 5/67 (7%)
Query: 269 FDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASELSELLGLN 328
FD+DGVI +G + G+ AL +L + T++ FLTN + + A+ L+ LG+
Sbjct: 11 FDLDGVIYVGPEALPGAVEALERLRSGGKTIR----FLTNNPCMTREQTAARLNR-LGIE 65
Query: 329 VSASQVL 335
+ +V+
Sbjct: 66 AAKDEVI 72
>NAL1_HUMAN (Q9C000) NACHT-, LRR- and PYD-containing protein 2
(Death effector filament-forming ced-4-like apoptosis
protein) (Nucleotide-binding domain and caspase
recruitment domain) (Caspase recruitment domain protein
7)
Length = 1473
Score = 32.0 bits (71), Expect = 4.8
Identities = 35/114 (30%), Positives = 55/114 (47%), Gaps = 13/114 (11%)
Query: 261 IRPSFGIAFDIDG---VILLGNTPVGGSPAALRKLYNYD-GTL---KFPYVFLTNGGGIP 313
IR FG D VIL G +G S A + + G L +F +VF + +
Sbjct: 314 IRDLFGPGLDTQEPRIVILQGAAGIGKSTLARQVKEAWGRGQLYGDRFQHVFYFSCRELA 373
Query: 314 EAKRASELSELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSE 367
++K S L+EL+G + +A+ +P RQ+++R E L + G EP V+ E
Sbjct: 374 QSKVVS-LAELIGKDGTATP-----APIRQILSRPERLLFILDGVDEPGWVLQE 421
>PURU_ECOLI (P37051) Formyltetrahydrofolate deformylase (EC
3.5.1.10) (Formyl-FH(4) hydrolase)
Length = 280
Score = 31.2 bits (69), Expect = 8.2
Identities = 50/198 (25%), Positives = 79/198 (39%), Gaps = 32/198 (16%)
Query: 316 KRASELSELL------GLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYG 369
K A L +LL GL+V + V+ H R LV RF+ + + +G L +E+
Sbjct: 93 KEAHCLGDLLMKANYGGLDVEIAAVIGNHDTLRSLVERFDIPFELVSHEG---LTRNEHD 149
Query: 370 FKNVISIDAYASRFENIDPLAPYKKWTTKLATTQNPKFDESGP-QIDVFSERVQAAFIVS 428
K +IDAY + LA Y + T P+F P +I AFI +
Sbjct: 150 QKMADAIDAYQPDYV---VLAKYMRVLT-------PEFVARFPNKIINIHHSFLPAFIGA 199
Query: 429 DPVD--WSRDVQVL-------CDILKTGGLPGRNVGTQPHLYFANDDLEYQTKFPSERLG 479
P + R V+++ D L G + ++V H Y A D + L
Sbjct: 200 RPYHQAYERGVKIIGATAHYVNDNLDEGPIIMQDVIHVDHTYTAEDMMRAGRDVEKNVLS 259
Query: 480 MGAFRIALESIF---NRT 494
+++ + +F NRT
Sbjct: 260 RALYKVLAQRVFVYGNRT 277
>ARAL_BACSU (P94526) Arabinose operon protein araL
Length = 272
Score = 31.2 bits (69), Expect = 8.2
Identities = 23/73 (31%), Positives = 36/73 (48%), Gaps = 8/73 (10%)
Query: 257 NDLLIRPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAK 316
+D + P+ GI D+DG + GN + G+ A++ L + VFL+N G I
Sbjct: 7 HDTPVSPA-GILIDLDGTVFRGNELIEGAREAIKTLRRMGKKI----VFLSNRGNI---S 58
Query: 317 RASELSELLGLNV 329
RA +LLG +
Sbjct: 59 RAMCRKKLLGAGI 71
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 72,246,484
Number of Sequences: 164201
Number of extensions: 3134159
Number of successful extensions: 6518
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 6501
Number of HSP's gapped (non-prelim): 10
length of query: 605
length of database: 59,974,054
effective HSP length: 116
effective length of query: 489
effective length of database: 40,926,738
effective search space: 20013174882
effective search space used: 20013174882
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC135230.3


