

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.11 + phase: 0 /pseudo
(954 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
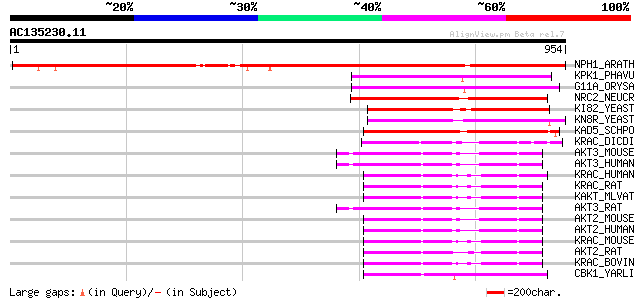
Score E
Sequences producing significant alignments: (bits) Value
NPH1_ARATH (O48963) Nonphototropic hypocotyl protein 1 (EC 2.7.1... 1195 0.0
KPK1_PHAVU (P15792) Protein kinase PVPK-1 (EC 2.7.1.-) 347 9e-95
G11A_ORYSA (P47997) Protein kinase G11A (EC 2.7.1.-) (Fragment) 345 3e-94
NRC2_NEUCR (O42626) Serine/threonine-protein kinase nrc-2 (EC 2.... 305 4e-82
KI82_YEAST (P25341) Probable serine/threonine-protein kinase KIN... 295 5e-79
KN8R_YEAST (P53739) Probable serine/threonine-protein kinase YNR... 294 9e-79
KAD5_SCHPO (Q09831) Probable serine/threonine-protein kinase C4G... 290 2e-77
KRAC_DICDI (P54644) RAC-family serine/threonine-protein kinase h... 219 3e-56
AKT3_MOUSE (Q9WUA6) RAC-gamma serine/threonine-protein kinase (E... 212 3e-54
AKT3_HUMAN (Q9Y243) RAC-gamma serine/threonine-protein kinase (E... 212 3e-54
KRAC_HUMAN (P31749) RAC-alpha serine/threonine-protein kinase (E... 211 1e-53
KRAC_RAT (P47196) RAC-alpha serine/threonine-protein kinase (EC ... 210 1e-53
KAKT_MLVAT (P31748) AKT kinase transforming protein (EC 2.7.1.-) 210 1e-53
AKT3_RAT (Q63484) RAC-gamma serine/threonine-protein kinase (EC ... 210 1e-53
AKT2_MOUSE (Q60823) RAC-beta serine/threonine-protein kinase (EC... 210 2e-53
AKT2_HUMAN (P31751) RAC-beta serine/threonine-protein kinase (EC... 210 2e-53
KRAC_MOUSE (P31750) RAC-alpha serine/threonine-protein kinase (E... 209 3e-53
AKT2_RAT (P47197) RAC-beta serine/threonine-protein kinase (EC 2... 209 3e-53
KRAC_BOVIN (Q01314) RAC-alpha serine/threonine-protein kinase (E... 206 3e-52
CBK1_YARLI (Q6CFS5) Serine/threonine-protein kinase CBK1 (EC 2.7... 204 9e-52
>NPH1_ARATH (O48963) Nonphototropic hypocotyl protein 1 (EC
2.7.1.37) (Phototropin)
Length = 996
Score = 1195 bits (3092), Expect = 0.0
Identities = 644/1014 (63%), Positives = 750/1014 (73%), Gaps = 88/1014 (8%)
Query: 6 KSPSSSSQRPSFPRDPRGSLEVFNPTSNST---SPVRSPSHLKTWT-------------- 48
+ PS+ + PRD RGSLEVFNP++ T +PV P W
Sbjct: 5 EKPSTKPSSRTLPRDTRGSLEVFNPSTQLTRPDNPVFRPEP-PAWQNLSDPRGTSPQPRP 63
Query: 49 ETEEQHKDFISTDE--VTNTSWMAIKEGETG-----------------AAAQRAAEWGLV 89
+ E + + +D+ TSWMA+K+ AA QRAAEWGLV
Sbjct: 64 QQEPAPSNPVRSDQEIAVTTSWMALKDPSPETISKKTITAEKPQKSAVAAEQRAAEWGLV 123
Query: 90 LRTDAETGKPQGVGVRNSG--DDEQNGK-FSGKRNSNNSGRVSGDSSDGGDP---RGFPR 143
L+TD +TGKPQGVGVRNSG +++ NGK + +RNS NS R SG+ SDG P G PR
Sbjct: 124 LKTDTKTGKPQGVGVRNSGGTENDPNGKKTTSQRNSQNSCRSSGEMSDGDVPGGRSGIPR 183
Query: 144 VSEDLKDALSAFQQTFVVSDATKPDYPIMYASAGFFNMTGYTSKEVIGRNCRFLQGADTD 203
VSEDLKDALS FQQTFVVSDATKPDYPIMYASAGFFNMTGYTSKEV+GRNCRFLQG+ TD
Sbjct: 184 VSEDLKDALSTFQQTFVVSDATKPDYPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTD 243
Query: 204 PQDVAKIREALEGGKSYCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLIGMLVEVNKH 263
++AKIRE L G +YCGR+LNYKKDGT FWNLLTI+PIKD+ G VLK IGM VEV+KH
Sbjct: 244 ADELAKIRETLAAGNNYCGRILNYKKDGTSFWNLLTIAPIKDESGKVLKFIGMQVEVSKH 303
Query: 264 TEGSKEKNLRPNGLPESLIRYDARQKEKASSSVSELLQAMKRPRALSESGQ-RPFIIKSG 322
TEG+KEK LRPNGLPESLIRYDARQK+ A++SV+EL++A+KRPRALSES PF+ KS
Sbjct: 304 TEGAKEKALRPNGLPESLIRYDARQKDMATNSVTELVEAVKRPRALSESTNLHPFMTKS- 362
Query: 323 GCSEEDQEIEKVEHKSRRKSDSVA-SFRPKSQRKSRSSMERISELPENANKNSHRHSFMG 381
E E+ K +RR S++V S R S R+SM+RI+E+PE ++ S SFMG
Sbjct: 363 ----ESDELPK--KPARRMSENVVPSGRRNSGGGRRNSMQRINEIPEKKSRKSSL-SFMG 415
Query: 382 LMKALIMKLL*I*ALRVRMMTGMIV-------LNLMTKKS*GKSEKVLILLLHLSV-LRK 433
+ K +L + G I ++ ++ +KV + + L
Sbjct: 416 IKKKSE-------SLDESIDDGFIEYGEEDDEISDRDERPESVDDKVRQKEMRKGIDLAT 468
Query: 434 TLSLLIQGFLT---------ILY----FLELTEYSREEILGRNCRFLQGPETDPATVRKI 480
TL + + F+ I++ FLELTEYSREEILGRNCRFLQGPETD TV+KI
Sbjct: 469 TLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGRNCRFLQGPETDLTTVKKI 528
Query: 481 REAIDNQTEVTVQLINYTRTGKKFWNLFHLQPMRDHKGEVQYFIGVQLDGSQHVEPLHNC 540
R AIDNQTEVTVQLINYT++GKKFWN+FHLQPMRD KGEVQYFIGVQLDGS+HVEP+ N
Sbjct: 529 RNAIDNQTEVTVQLINYTKSGKKFWNIFHLQPMRDQKGEVQYFIGVQLDGSKHVEPVRNV 588
Query: 541 IKEDTAKEGEQLVKQTAENVGEAVRELPDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAI 600
I+E KEGE LVK+TA N+ EAVRELPDAN P+DLW NHSKVVH KPHRKD+ W AI
Sbjct: 589 IEETAVKEGEDLVKKTAVNIDEAVRELPDANMTPEDLWANHSKVVHCKPHRKDSPPWIAI 648
Query: 601 QKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRA 660
QK++E+GE I LKHF+P+KPLGSGDTGSVHLVEL GT Q FAMKAMDK VMLNRNKVHRA
Sbjct: 649 QKVLESGEPIGLKHFKPVKPLGSGDTGSVHLVELVGTDQLFAMKAMDKAVMLNRNKVHRA 708
Query: 661 CTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFY 720
EREILD+LDHPFLPALYASFQTKTH+CLITDYYPGGELF+LLD+QP KVLKEDAVRFY
Sbjct: 709 RAEREILDLLDHPFLPALYASFQTKTHICLITDYYPGGELFMLLDRQPRKVLKEDAVRFY 768
Query: 721 AAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKK 780
AA+V++ALEYLHCQGIIYRDLKPENVLIQ NG +SL+DFDLSCLTSCKPQL+IP+ ++KK
Sbjct: 769 AAQVVVALEYLHCQGIIYRDLKPENVLIQGNGDISLSDFDLSCLTSCKPQLLIPSIDEKK 828
Query: 781 KRKKKKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILL 840
K+K QQK+QQ P FMAEPMRASNSFVGTEEYIAPEII+G+GHTSAVDWWALGIL+
Sbjct: 829 KKK------QQKSQQTPIFMAEPMRASNSFVGTEEYIAPEIISGAGHTSAVDWWALGILM 882
Query: 841 YEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEG 900
YEMLYGYTPFRGKTRQKTF N+L KDLKFP S P S Q KQLI+ LL RDPK RLG EG
Sbjct: 883 YEMLYGYTPFRGKTRQKTFTNVLQKDLKFPASIPASLQVKQLIFRLLQRDPKKRLGCFEG 942
Query: 901 ANEIKSHPFFKNVNWALIRCMKPPELDAPILLENDEKKEAKDIDPGLDDLQKNI 954
ANE+K H FFK +NWALIRC PPEL+ PI E E K +DP L+DLQ N+
Sbjct: 943 ANEVKQHSFFKGINWALIRCTNPPELETPIFSGEAENGE-KVVDPELEDLQTNV 995
>KPK1_PHAVU (P15792) Protein kinase PVPK-1 (EC 2.7.1.-)
Length = 609
Score = 347 bits (890), Expect = 9e-95
Identities = 183/385 (47%), Positives = 232/385 (59%), Gaps = 41/385 (10%)
Query: 588 KPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMD 647
KPH+ ++ W AIQ + + ++HFR +K LG GD GSV+L EL GT FAMK M+
Sbjct: 202 KPHKANDIRWEAIQAVRTRDGMLEMRHFRLLKKLGCGDIGSVYLAELSGTRTSFAMKVMN 261
Query: 648 KGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQ 707
K + NR K+ RA TEREIL LDHPFLP LY F+T+ CL+ ++ PGG+L L +Q
Sbjct: 262 KTELANRKKLLRAQTEREILQSLDHPFLPTLYTHFETEIFSCLVMEFCPGGDLHALRQRQ 321
Query: 708 PTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSC 767
P K E AVRFY AEVL++LEYLH GIIYRDLKPENVL++ +GH+ L+DFDLS S
Sbjct: 322 PGKYFSEHAVRFYVAEVLLSLEYLHMLGIIYRDLKPENVLVREDGHIMLSDFDLSLRCSV 381
Query: 768 KPQLIIPANE--------------------------------------DKKKRKKKKKKG 789
P L+ +N KK KK K K
Sbjct: 382 SPTLVKSSNNLQTKSSGYCVQPSCIEPTCVMQPDCIKPSCFTPRFLSGKSKKDKKSKPKN 441
Query: 790 QQKTQ--QIPTFMAEPMRA-SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYG 846
Q +P +AEP A S SFVGT EY+APEII G GH SAVDWW GI LYE+L+G
Sbjct: 442 DMHNQVTPLPELIAEPTNARSMSFVGTHEYLAPEIIKGEGHGSAVDWWTFGIFLYELLFG 501
Query: 847 YTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKS 906
TPF+G + T N++ + L+FP+S VS A+ LI LL ++P++RL GA EIK
Sbjct: 502 RTPFKGSANRATLFNVIGQPLRFPESPTVSFAARDLIRGLLVKEPQHRLAYRRGATEIKQ 561
Query: 907 HPFFKNVNWALIRCMKPPELDAPIL 931
HPFF+NVNWALIRC PPE+ ++
Sbjct: 562 HPFFQNVNWALIRCATPPEVPRQVI 586
>G11A_ORYSA (P47997) Protein kinase G11A (EC 2.7.1.-) (Fragment)
Length = 531
Score = 345 bits (885), Expect = 3e-94
Identities = 183/401 (45%), Positives = 236/401 (58%), Gaps = 43/401 (10%)
Query: 588 KPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMD 647
KPH+ ++ W AIQ I + L HF+ +K LG GD GSV+L EL GT YFAMK MD
Sbjct: 115 KPHKANDSRWEAIQMIRTRDGILGLSHFKLLKKLGCGDIGSVYLSELSGTESYFAMKVMD 174
Query: 648 KGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQ 707
K + +R K+ RA TE+EIL LDHPFLP LY F+T CL+ ++ PGG+L L +Q
Sbjct: 175 KASLASRKKLLRAQTEKEILQCLDHPFLPTLYTHFETDKFSCLVMEFCPGGDLHTLRQRQ 234
Query: 708 PTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSC 767
K E AV+FY AE+L+A+EYLH GIIYRDLKPENVL++ +GH+ L+DFDLS +
Sbjct: 235 RGKYFPEQAVKFYVAEILLAMEYLHMLGIIYRDLKPENVLVREDGHIMLSDFDLSLRCAV 294
Query: 768 KPQLIIPANEDKK---------------------------------------KRKKKKKK 788
P LI +N D + K KK +K
Sbjct: 295 SPTLIRSSNPDAEALRKNNQAYCVQPACVEPSCMIQPSCATPTTCFGPRFFSKSKKDRKP 354
Query: 789 GQQKTQQI---PTFMAEPMRA-SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEML 844
+ Q+ P +AEP A S SFVGT EY+APEII G GH SAVDWW GI LYE+L
Sbjct: 355 KPEVVNQVSPWPELIAEPSDARSMSFVGTHEYLAPEIIKGEGHGSAVDWWTFGIFLYELL 414
Query: 845 YGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEI 904
+G TPF+G + T N++ + L+FP+ VS A+ LI LL ++P+ RLG GA EI
Sbjct: 415 FGKTPFKGSGNRATLFNVIGQPLRFPEYPVVSFSARDLIRGLLVKEPQQRLGCKRGATEI 474
Query: 905 KSHPFFKNVNWALIRCMKPPELDAPILLENDEKKEAKDIDP 945
K HPFF+ VNWALIRC PPE+ P+ +E K+ +P
Sbjct: 475 KQHPFFEGVNWALIRCASPPEVPRPVEIERPPKQPVSTSEP 515
>NRC2_NEUCR (O42626) Serine/threonine-protein kinase nrc-2 (EC
2.7.1.37) (Nonrepressible conidiation protein 2)
Length = 623
Score = 305 bits (781), Expect = 4e-82
Identities = 168/349 (48%), Positives = 218/349 (62%), Gaps = 24/349 (6%)
Query: 587 PKPHRKDNDAWR---AIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAM 643
P P R+ A+R + I ++ + F IK +G GD G V+LV+ + +G+ +AM
Sbjct: 211 PPPIRQSPLAFRRTYSSNSIKVRNVEVGPQSFDKIKLIGKGDVGKVYLVKEKKSGRLYAM 270
Query: 644 KAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLL 703
K + K M+ RNK+ RA E+EIL +HPF+ LY SFQ++ ++ L +Y GGE F
Sbjct: 271 KVLSKKEMIKRNKIKRALAEQEILATSNHPFIVTLYHSFQSEDYLYLCMEYCSGGEFFRA 330
Query: 704 LDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSC 763
L +P K + ED RFYAAEV ALEYLH G IYRDLKPEN+L+ ++GH+ L+DFDLS
Sbjct: 331 LQTRPGKCIPEDDARFYAAEVTAALEYLHLMGFIYRDLKPENILLHQSGHIMLSDFDLSK 390
Query: 764 LT--SCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMAEPMRA---SNSFVGTEEYIA 818
+ KP +II K T +PT + A +NSFVGTEEYIA
Sbjct: 391 QSDPGGKPTMII-------------GKNGTSTSSLPTIDTKSCIANFRTNSFVGTEEYIA 437
Query: 819 PEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPK---SKPV 875
PE+I GSGHTSAVDWW LGIL+YEMLYG TPF+GK R TFANIL +D+ FP + +
Sbjct: 438 PEVIKGSGHTSAVDWWTLGILIYEMLYGTTPFKGKNRNATFANILREDIPFPDHAGAPQI 497
Query: 876 SPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPP 924
S K LI LL +D RLG+ GA++IK+HPFF+ WALIR MKPP
Sbjct: 498 SNLCKSLIRKLLIKDENRRLGARAGASDIKTHPFFRTTQWALIRHMKPP 546
>KI82_YEAST (P25341) Probable serine/threonine-protein kinase KIN82
(EC 2.7.1.37)
Length = 720
Score = 295 bits (754), Expect = 5e-79
Identities = 153/314 (48%), Positives = 199/314 (62%), Gaps = 17/314 (5%)
Query: 615 FRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPF 674
F I+ LG GD G V+L+ T Q FA+K ++K M+ R K+ R TE+EIL DHPF
Sbjct: 324 FEKIRLLGQGDVGKVYLMRERDTNQIFALKVLNKHEMIKRKKIKRVLTEQEILATSDHPF 383
Query: 675 LPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQ 734
+ LY SFQTK ++ L +Y GGE F L + +K + E+ +FYA+EV+ ALEYLH
Sbjct: 384 IVTLYHSFQTKDYLYLCMEYCMGGEFFRALQTRKSKCIAEEDAKFYASEVVAALEYLHLL 443
Query: 735 GIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQ 794
G IYRDLKPEN+L+ ++GHV L+DFDLS I A KK K +
Sbjct: 444 GFIYRDLKPENILLHQSGHVMLSDFDLS----------IQATGSKKPTMKD-------ST 486
Query: 795 QIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKT 854
+ T + +NSFVGTEEY+APE+I G+GHT+AVDWW LGIL+YEML+G TPF+G
Sbjct: 487 YLDTKICSDGFRTNSFVGTEEYLAPEVIRGNGHTAAVDWWTLGILIYEMLFGCTPFKGDN 546
Query: 855 RQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVN 914
+TF+NIL KD+KFP K VS K LI LL+++ RLGS GA +IK HPFFK V
Sbjct: 547 SNETFSNILTKDVKFPHDKEVSKNCKDLIKKLLNKNEAKRLGSKSGAADIKRHPFFKKVQ 606
Query: 915 WALIRCMKPPELDA 928
W+ +R PP + A
Sbjct: 607 WSFLRNQDPPLIPA 620
>KN8R_YEAST (P53739) Probable serine/threonine-protein kinase
YNR047W (EC 2.7.1.37)
Length = 893
Score = 294 bits (752), Expect = 9e-79
Identities = 153/348 (43%), Positives = 211/348 (59%), Gaps = 22/348 (6%)
Query: 615 FRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPF 674
F I+ LG GD G V LV + T + +A+K + K M+ RNK+ R TE+EIL +HPF
Sbjct: 496 FEKIRLLGQGDVGKVFLVREKKTNRVYALKVLSKDEMIKRNKIKRVLTEQEILATSNHPF 555
Query: 675 LPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQ 734
+ LY SFQ++ ++ L +Y GGE F L + TK + ED RFYA+EV ALEYLH
Sbjct: 556 IVTLYHSFQSEDYLYLCMEYCMGGEFFRALQTRKTKCICEDDARFYASEVTAALEYLHLL 615
Query: 735 GIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQ 794
G IYRDLKPEN+L+ ++GH+ L+DFDLS K K KG ++
Sbjct: 616 GFIYRDLKPENILLHQSGHIMLSDFDLSI--------------QAKDSKVPVVKGSAQST 661
Query: 795 QIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKT 854
+ T + +NSFVGTEEYIAPE+I G+GHT+AVDWW LGIL+YEML+G+TPF+G
Sbjct: 662 LVDTKICSDGFRTNSFVGTEEYIAPEVIRGNGHTAAVDWWTLGILIYEMLFGFTPFKGDN 721
Query: 855 RQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVN 914
+TF NIL ++ FP + +S K LI LL ++ RLG GA ++K HPFFK V
Sbjct: 722 TNETFTNILKNEVSFPNNNEISRTCKDLIKKLLTKNESKRLGCKMGAADVKKHPFFKKVQ 781
Query: 915 WALIRCMKPPEL--------DAPILLENDEKKEAKDIDPGLDDLQKNI 954
W+L+R +PP + D L N +++ ++D LD+ +KN+
Sbjct: 782 WSLLRNQEPPLIPVLSEDGYDFAKLSSNKKRQTSQDSHKHLDEQEKNM 829
>KAD5_SCHPO (Q09831) Probable serine/threonine-protein kinase
C4G8.05 (EC 2.7.1.37)
Length = 566
Score = 290 bits (741), Expect = 2e-77
Identities = 161/348 (46%), Positives = 210/348 (60%), Gaps = 23/348 (6%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
++ F + LG GD G V+LV + +G+++AMK + K M+ RNK RA E+ IL
Sbjct: 189 EVGPSSFEKVFLLGKGDVGRVYLVREKKSGKFYAMKVLSKQEMIKRNKSKRAFAEQHILA 248
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
+HPF+ LY SFQ+ ++ L +Y GGE F L ++P + L E+ +FY AEV AL
Sbjct: 249 TSNHPFIVTLYHSFQSDEYLYLCMEYCMGGEFFRALQRRPGRCLSENEAKFYIAEVTAAL 308
Query: 729 EYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLT-SCKPQLIIPANEDKKKRKKKKK 787
EYLH G IYRDLKPEN+L+ +GH+ L+DFDLS + S +I A +
Sbjct: 309 EYLHLMGFIYRDLKPENILLHESGHIMLSDFDLSKQSNSAGAPTVIQA---------RNA 359
Query: 788 KGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGY 847
Q + T +NSFVGTEEYIAPE+I G GHTSAVDWW LGIL YEMLY
Sbjct: 360 PSAQNAYALDTKSCIADFRTNSFVGTEEYIAPEVIKGCGHTSAVDWWTLGILFYEMLYAT 419
Query: 848 TPFRGKTRQKTFANILHKDLKFPK---SKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEI 904
TPF+GK R TF+NILHKD+ FP+ + +S K LI LL +D +RLGS GA ++
Sbjct: 420 TPFKGKNRNMTFSNILHKDVIFPEYADAPSISSLCKNLIRKLLVKDENDRLGSQAGAADV 479
Query: 905 KSHPFFKNVNWALIRCMKPPELDAPILLENDEK--------KEAKDID 944
K HPFFKNV WAL+R +PP + P L DEK KE+K +D
Sbjct: 480 KLHPFFKNVQWALLRHTEPPII--PKLAPIDEKGNPNISHLKESKSLD 525
>KRAC_DICDI (P54644) RAC-family serine/threonine-protein kinase
homolog (EC 2.7.1.37)
Length = 444
Score = 219 bits (558), Expect = 3e-56
Identities = 128/346 (36%), Positives = 183/346 (51%), Gaps = 46/346 (13%)
Query: 605 ENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTER 664
+ E++ + F + +G G G V V + TG+ +AMK + K ++ N+V +ER
Sbjct: 110 KKSEKVGVADFELLNLVGKGSFGKVIQVRKKDTGEVYAMKVLSKKHIVEHNEVEHTLSER 169
Query: 665 EILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEV 724
IL ++HPFL L SFQT+ + I DY GGELF L Q K ED VR+Y AE+
Sbjct: 170 NILQKINHPFLVNLNYSFQTEDKLYFILDYVNGGELFYHL--QKDKKFTEDRVRYYGAEI 227
Query: 725 LIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKK 784
++ALE+LH G+IYRDLKPEN+L+ GH+ +TDF L CK L+ P ++
Sbjct: 228 VLALEHLHLSGVIYRDLKPENLLLTNEGHICMTDFGL-----CKEGLLTPTDK------- 275
Query: 785 KKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEML 844
+ +F GT EY+APE++ G+G+ VDWW+ G LLYEML
Sbjct: 276 ----------------------TGTFCGTPEYLAPEVLQGNGYGKQVDWWSFGSLLYEML 313
Query: 845 YGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEI 904
G PF + Q+ + I+ + L FP +SP A+ L+ LL RDP+ RL N I
Sbjct: 314 TGLPPFYNQDVQEMYRKIMMEKLSFPHF--ISPDARSLLEQLLERDPEKRLAD---PNLI 368
Query: 905 KSHPFFKNVNW-ALIRCMKPPELDAPILLENDEKKEAKDIDPGLDD 949
K HPFF++++W L + PP P + + IDP D
Sbjct: 369 KRHPFFRSIDWEQLFQKNIPP----PFIPNVKGSADTSQIDPVFTD 410
>AKT3_MOUSE (Q9WUA6) RAC-gamma serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-gamma) (Protein kinase Akt-3) (Protein
kinase B, gamma) (PKB gamma)
Length = 479
Score = 212 bits (540), Expect = 3e-54
Identities = 125/355 (35%), Positives = 190/355 (53%), Gaps = 45/355 (12%)
Query: 562 EAVRELPDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPL 621
EA++ + D Q+ ++ +N S + DN + + ++ ++ F +K L
Sbjct: 100 EAIQAVADRLQRQEEERMNCSPT-----SQIDNIGEEEMDASTTHHKRKTMNDFDYLKLL 154
Query: 622 GSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYAS 681
G G G V LV + +G+Y+AMK + K V++ +++V TE +L HPFL +L S
Sbjct: 155 GKGTFGKVILVREKASGKYYAMKILKKEVIIAKDEVAHTLTESRVLKNTRHPFLTSLKYS 214
Query: 682 FQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDL 741
FQTK +C + +Y GGELF L ++ +V ED RFY AE++ AL+YLH I+YRDL
Sbjct: 215 FQTKDRLCFVMEYVNGGELFFHLSRE--RVFSEDRTRFYGAEIVSALDYLHSGKIVYRDL 272
Query: 742 KPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMA 801
K EN+++ ++GH+ +TDF L CK + A
Sbjct: 273 KLENLMLDKDGHIKITDFGL-----CKEGITDAA-------------------------- 301
Query: 802 EPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFAN 861
+F GT EY+APE++ + + AVDWW LG+++YEM+ G PF + +K F
Sbjct: 302 ----TMKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGRLPFYNQDHEKLFEL 357
Query: 862 ILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKSHPFFKNVNW 915
IL +D+KFP++ +S AK L+ LL +DP RL G + A EI H FF VNW
Sbjct: 358 ILMEDIKFPRT--LSSDAKSLLSGLLIKDPNKRLGGGPDDAKEIMRHSFFSGVNW 410
>AKT3_HUMAN (Q9Y243) RAC-gamma serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-gamma) (Protein kinase Akt-3) (Protein
kinase B, gamma) (PKB gamma) (STK-2)
Length = 479
Score = 212 bits (540), Expect = 3e-54
Identities = 125/355 (35%), Positives = 190/355 (53%), Gaps = 45/355 (12%)
Query: 562 EAVRELPDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPL 621
EA++ + D Q+ ++ +N S + DN + + ++ ++ F +K L
Sbjct: 100 EAIQAVADRLQRQEEERMNCSPT-----SQIDNIGEEEMDASTTHHKRKTMNDFDYLKLL 154
Query: 622 GSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYAS 681
G G G V LV + +G+Y+AMK + K V++ +++V TE +L HPFL +L S
Sbjct: 155 GKGTFGKVILVREKASGKYYAMKILKKEVIIAKDEVAHTLTESRVLKNTRHPFLTSLKYS 214
Query: 682 FQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDL 741
FQTK +C + +Y GGELF L ++ +V ED RFY AE++ AL+YLH I+YRDL
Sbjct: 215 FQTKDRLCFVMEYVNGGELFFHLSRE--RVFSEDRTRFYGAEIVSALDYLHSGKIVYRDL 272
Query: 742 KPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMA 801
K EN+++ ++GH+ +TDF L CK + A
Sbjct: 273 KLENLMLDKDGHIKITDFGL-----CKEGITDAA-------------------------- 301
Query: 802 EPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFAN 861
+F GT EY+APE++ + + AVDWW LG+++YEM+ G PF + +K F
Sbjct: 302 ----TMKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGRLPFYNQDHEKLFEL 357
Query: 862 ILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKSHPFFKNVNW 915
IL +D+KFP++ +S AK L+ LL +DP RL G + A EI H FF VNW
Sbjct: 358 ILMEDIKFPRT--LSSDAKSLLSGLLIKDPNKRLGGGPDDAKEIMRHSFFSGVNW 410
>KRAC_HUMAN (P31749) RAC-alpha serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-alpha) (Protein kinase B) (PKB)
(C-AKT)
Length = 480
Score = 211 bits (536), Expect = 1e-53
Identities = 119/320 (37%), Positives = 180/320 (56%), Gaps = 43/320 (13%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
++++ F +K LG G G V LV+ + TG+Y+AMK + K V++ +++V TE +L
Sbjct: 144 RVTMNEFEYLKLLGKGTFGKVILVKEKATGRYYAMKILKKEVIVAKDEVAHTLTENRVLQ 203
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
HPFL AL SFQT +C + +Y GGELF L ++ +V ED RFY AE++ AL
Sbjct: 204 NSRHPFLTALKYSFQTHDRLCFVMEYANGGELFFHLSRE--RVFSEDRARFYGAEIVSAL 261
Query: 729 EYLHCQ-GIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKK 787
+YLH + ++YRDLK EN+++ ++GH+ +TDF L CK + K
Sbjct: 262 DYLHSEKNVVYRDLKLENLMLDKDGHIKITDFGL-----CKEGI--------------KD 302
Query: 788 KGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGY 847
KT F GT EY+APE++ + + AVDWW LG+++YEM+ G
Sbjct: 303 GATMKT----------------FCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGR 346
Query: 848 TPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKS 906
PF + +K F IL ++++FP++ + P+AK L+ LL +DPK RL G E A EI
Sbjct: 347 LPFYNQDHEKLFELILMEEIRFPRT--LGPEAKSLLSGLLKKDPKQRLGGGSEDAKEIMQ 404
Query: 907 HPFFKNVNWALI--RCMKPP 924
H FF + W + + + PP
Sbjct: 405 HRFFAGIVWQHVYEKKLSPP 424
>KRAC_RAT (P47196) RAC-alpha serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-alpha) (Protein kinase B) (PKB)
Length = 480
Score = 210 bits (535), Expect = 1e-53
Identities = 117/309 (37%), Positives = 175/309 (55%), Gaps = 41/309 (13%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
++++ F +K LG G G V LV+ + TG+Y+AMK + K V++ +++V TE +L
Sbjct: 144 RVTMNEFEYLKLLGKGTFGKVILVKEKATGRYYAMKILKKEVIVAKDEVAHTLTENRVLQ 203
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
HPFL AL SFQT +C + +Y GGELF L ++ +V ED RFY AE++ AL
Sbjct: 204 NSRHPFLTALKYSFQTHDRLCFVMEYANGGELFFHLSRE--RVFSEDRARFYGAEIVSAL 261
Query: 729 EYLHCQ-GIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKK 787
+YLH + ++YRDLK EN+++ ++GH+ +TDF L CK + K
Sbjct: 262 DYLHSEKNVVYRDLKLENLMLDKDGHIKITDFGL-----CKEGI--------------KD 302
Query: 788 KGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGY 847
KT F GT EY+APE++ + + AVDWW LG+++YEM+ G
Sbjct: 303 GATMKT----------------FCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGR 346
Query: 848 TPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKS 906
PF + +K F IL ++++FP++ + P+AK L+ LL +DP RL G E A EI
Sbjct: 347 LPFYNQDHEKLFELILMEEIRFPRT--LGPEAKSLLSGLLKKDPTQRLGGGSEDAKEIMQ 404
Query: 907 HPFFKNVNW 915
H FF N+ W
Sbjct: 405 HRFFANIVW 413
>KAKT_MLVAT (P31748) AKT kinase transforming protein (EC 2.7.1.-)
Length = 501
Score = 210 bits (535), Expect = 1e-53
Identities = 117/309 (37%), Positives = 175/309 (55%), Gaps = 41/309 (13%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
++++ F +K LG G G V LV+ + TG+Y+AMK + K V++ +++V TE +L
Sbjct: 165 RVTMNEFEYLKLLGKGTFGKVILVKEKATGRYYAMKILKKEVIVAKDEVAHTLTENRVLQ 224
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
HPFL AL SFQT +C + +Y GGELF L ++ +V ED RFY AE++ AL
Sbjct: 225 NSRHPFLTALKYSFQTHDRLCFVMEYANGGELFFHLSRE--RVFSEDRARFYGAEIVSAL 282
Query: 729 EYLHCQ-GIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKK 787
+YLH + ++YRDLK EN+++ ++GH+ +TDF L CK + K
Sbjct: 283 DYLHSEKNVVYRDLKLENLMLDKDGHIKITDFGL-----CKEGI--------------KD 323
Query: 788 KGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGY 847
KT F GT EY+APE++ + + AVDWW LG+++YEM+ G
Sbjct: 324 GATMKT----------------FCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGR 367
Query: 848 TPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKS 906
PF + +K F IL ++++FP++ + P+AK L+ LL +DP RL G E A EI
Sbjct: 368 LPFYNQDHEKLFELILMEEIRFPRT--LGPEAKSLLSGLLKKDPTQRLGGGSEDAKEIMQ 425
Query: 907 HPFFKNVNW 915
H FF N+ W
Sbjct: 426 HRFFANIVW 434
>AKT3_RAT (Q63484) RAC-gamma serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-gamma) (Protein kinase Akt-3) (Protein
kinase B, gamma) (PKB gamma)
Length = 454
Score = 210 bits (535), Expect = 1e-53
Identities = 124/355 (34%), Positives = 189/355 (52%), Gaps = 45/355 (12%)
Query: 562 EAVRELPDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPL 621
EA++ + D Q+ ++ +N S + DN + + ++ ++ F +K L
Sbjct: 100 EAIQAVADRLQRQEEERMNCSPT-----SQIDNIGEEEMDASTTHHKRKTMNDFDYLKLL 154
Query: 622 GSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYAS 681
G G G V LV + +G+Y+AMK + K V++ +++V TE +L HPFL +L S
Sbjct: 155 GKGTFGKVILVREKASGKYYAMKILKKEVIIAKDEVAHTLTESRVLKNTRHPFLTSLKYS 214
Query: 682 FQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDL 741
FQTK +C + +Y GGELF L ++ +V ED RFY AE++ AL+YLH I+YRDL
Sbjct: 215 FQTKDRLCFVMEYVNGGELFFHLSRE--RVFSEDRTRFYGAEIVSALDYLHSGKIVYRDL 272
Query: 742 KPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMA 801
K EN+++ ++GH+ +TDF L CK + A
Sbjct: 273 KLENLMLDKDGHIKITDFGL-----CKEGITDAA-------------------------- 301
Query: 802 EPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFAN 861
+F GT EY+APE++ + + AVDWW LG+++YEM+ G PF + +K F
Sbjct: 302 ----TMKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGRLPFYNQDHEKLFEL 357
Query: 862 ILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKSHPFFKNVNW 915
IL +D+KFP++ +S AK L+ LL +DP RL G + EI H FF VNW
Sbjct: 358 ILMEDIKFPRT--LSSDAKSLLSGLLIKDPNKRLGGGPDDPKEIMRHSFFSGVNW 410
>AKT2_MOUSE (Q60823) RAC-beta serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-beta) (Protein kinase Akt-2) (Protein
kinase B, beta) (PKB beta)
Length = 481
Score = 210 bits (534), Expect = 2e-53
Identities = 112/308 (36%), Positives = 174/308 (56%), Gaps = 40/308 (12%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
++++ F +K LG G G V LV + TG+Y+AMK + K V++ +++V TE +L
Sbjct: 146 KVTMNDFDYLKLLGKGTFGKVILVREKATGRYYAMKILRKEVIIAKDEVAHTVTESRVLQ 205
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
HPFL AL +FQT +C + +Y GGELF L ++ +V ED RFY AE++ AL
Sbjct: 206 NTRHPFLTALKYAFQTHDRLCFVMEYANGGELFFHLSRE--RVFTEDRARFYGAEIVSAL 263
Query: 729 EYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKK 788
EYLH + ++YRD+K EN+++ ++GH+ +TDF L CK + A
Sbjct: 264 EYLHSRDVVYRDIKLENLMLDKDGHIKITDFGL-----CKEGISDGA------------- 305
Query: 789 GQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYT 848
+F GT EY+APE++ + + AVDWW LG+++YEM+ G
Sbjct: 306 -----------------TMKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGRL 348
Query: 849 PFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKSH 907
PF + ++ F IL ++++FP++ + P+AK L+ LL +DPK RL G A E+ H
Sbjct: 349 PFYNQDHERLFELILMEEIRFPRT--LGPEAKSLLAGLLKKDPKQRLGGGPSDAKEVMEH 406
Query: 908 PFFKNVNW 915
FF ++NW
Sbjct: 407 RFFLSINW 414
>AKT2_HUMAN (P31751) RAC-beta serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-beta) (Protein kinase Akt-2) (Protein
kinase B, beta) (PKB beta)
Length = 481
Score = 210 bits (534), Expect = 2e-53
Identities = 112/308 (36%), Positives = 175/308 (56%), Gaps = 40/308 (12%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
++++ F +K LG G G V LV + TG+Y+AMK + K V++ +++V TE +L
Sbjct: 146 KVTMNDFDYLKLLGKGTFGKVILVREKATGRYYAMKILRKEVIIAKDEVAHTVTESRVLQ 205
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
HPFL AL +FQT +C + +Y GGELF L ++ +V E+ RFY AE++ AL
Sbjct: 206 NTRHPFLTALKYAFQTHDRLCFVMEYANGGELFFHLSRE--RVFTEERARFYGAEIVSAL 263
Query: 729 EYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKK 788
EYLH + ++YRD+K EN+++ ++GH+ +TDF L CK + A
Sbjct: 264 EYLHSRDVVYRDIKLENLMLDKDGHIKITDFGL-----CKEGISDGA------------- 305
Query: 789 GQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYT 848
+F GT EY+APE++ + + AVDWW LG+++YEM+ G
Sbjct: 306 -----------------TMKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGRL 348
Query: 849 PFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKSH 907
PF + ++ F IL ++++FP++ +SP+AK L+ LL +DPK RL G A E+ H
Sbjct: 349 PFYNQDHERLFELILMEEIRFPRT--LSPEAKSLLAGLLKKDPKQRLGGGPSDAKEVMEH 406
Query: 908 PFFKNVNW 915
FF ++NW
Sbjct: 407 RFFLSINW 414
>KRAC_MOUSE (P31750) RAC-alpha serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-alpha) (AKT1 kinase) (Protein kinase
B) (PKB) (C-AKT) (Thymoma viral proto-oncogene)
Length = 480
Score = 209 bits (532), Expect = 3e-53
Identities = 117/309 (37%), Positives = 174/309 (55%), Gaps = 41/309 (13%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
++++ F +K LG G G V LV+ + TG+Y+AMK + K V++ +++V TE +L
Sbjct: 144 RVTMNEFEYLKLLGKGTFGKVILVKEKATGRYYAMKILKKEVIVAKDEVAHTLTENRVLQ 203
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
HPFL AL SFQT +C + +Y GGELF L ++ +V ED RFY AE++ AL
Sbjct: 204 NSRHPFLTALKYSFQTHDRLCFVMEYANGGELFFHLSRE--RVFSEDRARFYGAEIVSAL 261
Query: 729 EYLHCQ-GIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKK 787
+YLH + ++YRDLK EN+++ ++GH+ +TDF L CK + K
Sbjct: 262 DYLHSEKNVVYRDLKLENLMLDKDGHIKITDFGL-----CKEGI--------------KD 302
Query: 788 KGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGY 847
KT F GT EY+APE++ + + AVDWW LG+++YEM+ G
Sbjct: 303 GATMKT----------------FCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGR 346
Query: 848 TPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKS 906
PF + +K F IL +++ FP++ + P+AK L+ LL +DP RL G E A EI
Sbjct: 347 LPFYNQDHEKLFELILMEEIAFPRT--LGPEAKSLLSGLLKKDPTQRLGGGSEDAKEIMQ 404
Query: 907 HPFFKNVNW 915
H FF N+ W
Sbjct: 405 HRFFANIVW 413
>AKT2_RAT (P47197) RAC-beta serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-beta) (Protein kinase Akt-2) (Protein
kinase B, beta) (PKB beta)
Length = 481
Score = 209 bits (532), Expect = 3e-53
Identities = 112/308 (36%), Positives = 174/308 (56%), Gaps = 40/308 (12%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
++++ F +K LG G G V LV + TG+Y+AMK + K V++ +++V TE +L
Sbjct: 146 KVTMNDFDYLKLLGKGTFGKVILVREKATGRYYAMKILRKEVIIAKDEVAHTVTESRVLQ 205
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
HPFL AL +FQT +C + +Y GG+LF L ++ +V ED RFY AE++ AL
Sbjct: 206 NTRHPFLTALKYAFQTHDRLCFVMEYANGGDLFFHLSRE--RVFTEDRARFYGAEIVSAL 263
Query: 729 EYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKK 788
EYLH ++YRD+K EN+++ ++GH+ +TDF LS K+
Sbjct: 264 EYLHSTDVVYRDIKLENLMLDKDGHIKITDFGLS------------------------KE 299
Query: 789 GQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYT 848
G + T F GT EY+APE++ + + AVDWW LG+++YEM+ G
Sbjct: 300 GISDGATMKT-----------FCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGRL 348
Query: 849 PFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKSH 907
PF + ++ F IL ++++FP++ + P+AK L+ LL +DPK RL G A E+ H
Sbjct: 349 PFYNQDHERLFELILMEEIRFPRT--LGPEAKSLLAGLLKKDPKQRLGGGPSDAKEVMEH 406
Query: 908 PFFKNVNW 915
FF ++NW
Sbjct: 407 RFFLSINW 414
>KRAC_BOVIN (Q01314) RAC-alpha serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-alpha) (Protein kinase B) (PKB)
Length = 480
Score = 206 bits (523), Expect = 3e-52
Identities = 116/309 (37%), Positives = 174/309 (55%), Gaps = 41/309 (13%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
++++ F +K LG G G V LV+ + T Y+AMK + K V++ +++V TE +L
Sbjct: 144 RVTMNEFEYVKLLGKGTFGKVILVKEKATAAYYAMKILKKEVIVAKDEVAHTLTENRVLQ 203
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
HP L AL SFQT +C + +Y GGELF L ++ +V ED RFY AE++ AL
Sbjct: 204 NSRHPSLTALKYSFQTHDRLCFVMEYANGGELFFHLSRE--RVFSEDRARFYGAEIVSAL 261
Query: 729 EYLHCQ-GIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKK 787
+YLH + ++YRDLK EN+++ ++GH+ +TDF L CK + K
Sbjct: 262 DYLHSEKEVVYRDLKLENLMLDKDGHIKITDFGL-----CKEGI--------------KD 302
Query: 788 KGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGY 847
KT F GT EY+APE++ + + AVDWW LG+++YEM+ G
Sbjct: 303 GATMKT----------------FCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGR 346
Query: 848 TPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL-GSLEGANEIKS 906
PF + +K F IL ++++FP++ +SP+AK L+ LL +DPK RL G E A EI
Sbjct: 347 LPFYNQDHEKLFELILMEEIRFPRT--LSPEAKSLLSGLLKKDPKQRLGGGSEDAKEIMQ 404
Query: 907 HPFFKNVNW 915
H FF ++ W
Sbjct: 405 HRFFASIVW 413
>CBK1_YARLI (Q6CFS5) Serine/threonine-protein kinase CBK1 (EC
2.7.1.37)
Length = 588
Score = 204 bits (519), Expect = 9e-52
Identities = 119/334 (35%), Positives = 184/334 (54%), Gaps = 21/334 (6%)
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
+++L+ F +K +G G G V LV+ G+ +AMK M K M + + ER++L
Sbjct: 201 RMALEDFVTVKVIGKGAFGEVRLVQKRDNGKIYAMKTMLKKEMDMKEQWAHVKAERDVLA 260
Query: 669 MLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIAL 728
D P++ +LY SFQ ++ LI ++ PGG+L +L + V ED RFY AE ++A+
Sbjct: 261 DSDSPWIVSLYFSFQDDLYLYLIMEFLPGGDLMTMLIKYD--VFSEDITRFYIAECVLAI 318
Query: 729 EYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSC----------------LTSCKPQLI 772
E +H G I+RD+KP+N+LI + GH+ L+DF LS TS P
Sbjct: 319 EAIHKLGFIHRDIKPDNILIDKTGHIKLSDFGLSTGFHKTHSSAYWKKLKDGTSSNPATQ 378
Query: 773 IPANEDKKKRKKKKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVD 832
+ ++ ++ QQI T+ + S VGT +YIAPEI G+ D
Sbjct: 379 MGPPQNTNRQSTYDSIHLTMRQQISTWRKNRRLMAYSTVGTPDYIAPEIFVHQGYGQECD 438
Query: 833 WWALGILLYEMLYGYTPFRGKTRQKTFANILH--KDLKFPKSKPVSPQAKQLIYWLLHRD 890
WW+LG +++E L G+ PF + ++T+ I++ + L+FP +SP+++ LI LL
Sbjct: 439 WWSLGAIMFECLVGWPPFCSEQPRETYHKIINWRETLQFPDDVHLSPESEDLIRRLL-TS 497
Query: 891 PKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPP 924
+NRLG + GANEIKSHPFF+ V+W+ IR P
Sbjct: 498 SENRLGRIGGANEIKSHPFFRGVDWSSIREFNAP 531
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.135 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 117,933,424
Number of Sequences: 164201
Number of extensions: 5373227
Number of successful extensions: 20843
Number of sequences better than 10.0: 1653
Number of HSP's better than 10.0 without gapping: 1318
Number of HSP's successfully gapped in prelim test: 335
Number of HSP's that attempted gapping in prelim test: 16263
Number of HSP's gapped (non-prelim): 3272
length of query: 954
length of database: 59,974,054
effective HSP length: 120
effective length of query: 834
effective length of database: 40,269,934
effective search space: 33585124956
effective search space used: 33585124956
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC135230.11


