

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135162.11 + phase: 0 /pseudo
(385 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
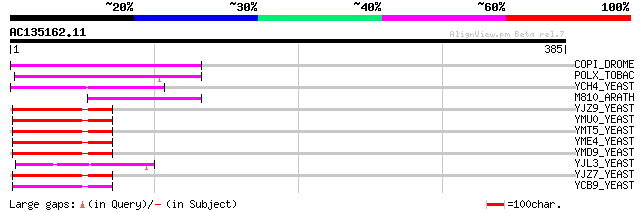
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 99 2e-20
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 97 5e-20
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 69 2e-11
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 61 5e-09
YJZ9_YEAST (P47100) Transposon Ty1 protein B 51 6e-06
YMU0_YEAST (Q04670) Transposon Ty1 protein B 48 5e-05
YMT5_YEAST (Q04214) Transposon Ty1 protein B 48 5e-05
YME4_YEAST (Q04711) Transposon Ty1 protein B 48 5e-05
YMD9_YEAST (Q03434) Transposon Ty1 protein B 48 5e-05
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 48 5e-05
YJZ7_YEAST (P47098) Transposon Ty1 protein B 45 3e-04
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 45 4e-04
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 98.6 bits (244), Expect = 2e-20
Identities = 46/134 (34%), Positives = 78/134 (57%), Gaps = 1/134 (0%)
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNL-VHICL 59
N D + +L+KA+YGLKQ+ R W + L + EFV + +Y+ + + N +++ L
Sbjct: 1027 NSDNVCKLNKAIYGLKQAARCWFEVFEQALKECEFVNSSVDRCIYILDKGNINENIYVLL 1086
Query: 60 YVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDI 119
YVDD+++ ++ + FK ++M+KF MTDL ++ +F+G+ + + Q Y I
Sbjct: 1087 YVDDVVIATGDMTRMNNFKRYLMEKFRMTDLNEIKHFIGIRIEMQEDKIYLSQSAYVKKI 1146
Query: 120 LKRFNMLNCNPAST 133
L +FNM NCN ST
Sbjct: 1147 LSKFNMENCNAVST 1160
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 97.4 bits (241), Expect = 5e-20
Identities = 50/132 (37%), Positives = 77/132 (57%), Gaps = 2/132 (1%)
Query: 4 MIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDD 63
M+ +L+K+LYGLKQ+ R W + +F+ Q ++K ++ VY K ++N + + LYVDD
Sbjct: 952 MVCKLNKSLYGLKQAPRQWYMKFDSFMKSQTYLKTYSDPCVYFKRFSENNFIILLLYVDD 1011
Query: 64 LIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFT--KTSEGLVKHQKKYASDILK 121
+++ + I KG + K F+M DLG LGM+ +TS L Q+KY +L+
Sbjct: 1012 MLIVGKDKGLIAKLKGDLSKSFDMKDLGPAQQILGMKIVRERTSRKLWLSQEKYIERVLE 1071
Query: 122 RFNMLNCNPAST 133
RFNM N P ST
Sbjct: 1072 RFNMKNAKPVST 1083
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 68.9 bits (167), Expect = 2e-11
Identities = 35/107 (32%), Positives = 63/107 (58%), Gaps = 1/107 (0%)
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
N D ++ L+ +YGLKQ+ WN+ I N L + F + + E+ +Y +++ D ++I +Y
Sbjct: 29 NPDYVWELYGGMYGLKQAPLLWNEHINNTLKKIGFCRHEGEHGLYFRSTSDGP-IYIGVY 87
Query: 61 VDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEG 107
VDDL+V + + K + K ++M DLGK+ FLG+ +++ G
Sbjct: 88 VDDLLVAAPSPKIYDRVKQELTKLYSMKDLGKVDKFLGLNIHQSTNG 134
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 60.8 bits (146), Expect = 5e-09
Identities = 30/79 (37%), Positives = 44/79 (54%)
Query: 55 VHICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKK 114
+++ LYVDD+++T S+ + + F+M DLG + YFLG++ GL Q K
Sbjct: 1 MYLLLYVDDILLTGSSNTLLNMLIFQLSSTFSMKDLGPVHYFLGIQIKTHPSGLFLSQTK 60
Query: 115 YASDILKRFNMLNCNPAST 133
YA IL ML+C P ST
Sbjct: 61 YAEQILNNAGMLDCKPMST 79
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 50.8 bits (120), Expect = 6e-06
Identities = 28/69 (40%), Positives = 43/69 (61%), Gaps = 4/69 (5%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + RL K+LYGLKQS W + I ++L+QQ ++ ++ KNS+ V ICL+VD
Sbjct: 1372 DKLIRLKKSLYGLKQSGANWYETIKSYLIQQCGMEEVRGWSCVFKNSQ----VTICLFVD 1427
Query: 63 DLIVTDSNL 71
D+++ NL
Sbjct: 1428 DMVLFSKNL 1436
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 47.8 bits (112), Expect = 5e-05
Identities = 27/69 (39%), Positives = 43/69 (62%), Gaps = 4/69 (5%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + RL K+LYGLKQS W + I ++L++Q ++ ++ KNS+ V ICL+VD
Sbjct: 945 DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVFKNSQ----VTICLFVD 1000
Query: 63 DLIVTDSNL 71
D+I+ +L
Sbjct: 1001 DMILFSKDL 1009
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 47.8 bits (112), Expect = 5e-05
Identities = 26/69 (37%), Positives = 43/69 (61%), Gaps = 4/69 (5%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + RL K+LYGLKQS W + I ++L++Q ++ ++ +NS+ V ICL+VD
Sbjct: 945 DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVFENSQ----VTICLFVD 1000
Query: 63 DLIVTDSNL 71
D+++ NL
Sbjct: 1001 DMVLFSKNL 1009
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 47.8 bits (112), Expect = 5e-05
Identities = 27/69 (39%), Positives = 43/69 (62%), Gaps = 4/69 (5%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + RL K+LYGLKQS W + I ++L++Q ++ ++ KNS+ V ICL+VD
Sbjct: 945 DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVFKNSQ----VTICLFVD 1000
Query: 63 DLIVTDSNL 71
D+I+ +L
Sbjct: 1001 DMILFSKDL 1009
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 47.8 bits (112), Expect = 5e-05
Identities = 27/69 (39%), Positives = 43/69 (62%), Gaps = 4/69 (5%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + RL K+LYGLKQS W + I ++L++Q ++ ++ KNS+ V ICL+VD
Sbjct: 945 DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVFKNSQ----VTICLFVD 1000
Query: 63 DLIVTDSNL 71
D+I+ +L
Sbjct: 1001 DMILFSKDL 1009
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 47.8 bits (112), Expect = 5e-05
Identities = 32/102 (31%), Positives = 50/102 (48%), Gaps = 9/102 (8%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+ +L+KALYGLKQS + WN + +L Y + ++D NL+ I +YVDD
Sbjct: 1409 VVKLNKALYGLKQSPKEWNDHLRQYL--NGIGLKDNSYTPGLYQTEDKNLM-IAVYVDDC 1465
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKL------TYFLGME 100
++ SN ++ F + F + G L T LGM+
Sbjct: 1466 VIAASNEQRLDEFINKLKSNFELKITGTLIDDVLDTDILGMD 1507
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 45.1 bits (105), Expect = 3e-04
Identities = 26/69 (37%), Positives = 42/69 (60%), Gaps = 4/69 (5%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + RL K+ YGLKQS W + I ++L++Q ++ ++ KNS+ V ICL+VD
Sbjct: 1372 DKLIRLKKSHYGLKQSGANWYETIKSYLIKQCGMEEVRGWSCVFKNSQ----VTICLFVD 1427
Query: 63 DLIVTDSNL 71
D+I+ +L
Sbjct: 1428 DMILFSKDL 1436
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 44.7 bits (104), Expect = 4e-04
Identities = 26/69 (37%), Positives = 41/69 (58%), Gaps = 4/69 (5%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + RL K+LYGLKQS W + I ++L+ ++ ++ KNS+ V ICL+VD
Sbjct: 1387 DKLLRLRKSLYGLKQSGANWYETIKSYLINCCDMQEVRGWSCVFKNSQ----VTICLFVD 1442
Query: 63 DLIVTDSNL 71
D+I+ +L
Sbjct: 1443 DMILFSKDL 1451
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.366 0.163 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,544,918
Number of Sequences: 164201
Number of extensions: 1052253
Number of successful extensions: 6533
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 6509
Number of HSP's gapped (non-prelim): 13
length of query: 385
length of database: 59,974,054
effective HSP length: 112
effective length of query: 273
effective length of database: 41,583,542
effective search space: 11352306966
effective search space used: 11352306966
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC135162.11


