

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135160.2 + phase: 0 /pseudo
(319 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
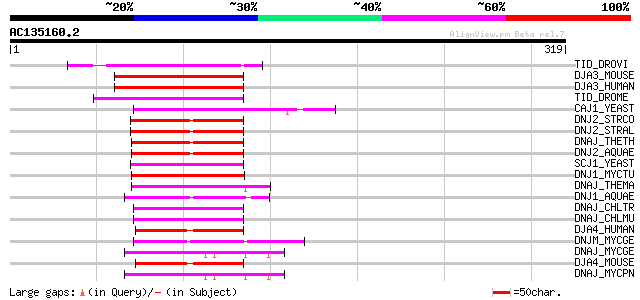
Score E
Sequences producing significant alignments: (bits) Value
TID_DROVI (Q24331) Tumorous imaginal discs protein, mitochondria... 71 4e-12
DJA3_MOUSE (Q99M87) DnaJ homolog subfamily A member 3, mitochond... 70 5e-12
DJA3_HUMAN (Q96EY1) DnaJ homolog subfamily A member 3, mitochond... 70 5e-12
TID_DROME (Q27237) Tumorous imaginal discs protein, mitochondria... 66 1e-10
CAJ1_YEAST (P39101) CAJ1 protein 64 7e-10
DNJ2_STRCO (Q9RDD7) Chaperone protein dnaJ2 63 9e-10
DNJ2_STRAL (O52164) Chaperone protein dnaJ2 63 9e-10
DNAJ_THETH (Q56237) Chaperone protein dnaJ 63 1e-09
DNJ2_AQUAE (O66921) Chaperone protein dnaJ-2 59 1e-08
SCJ1_YEAST (P25303) DnaJ-related protein SCJ1 59 2e-08
DNJ1_MYCTU (P07881) Chaperone protein dnaJ1 58 4e-08
DNAJ_THEMA (Q9WZV3) Chaperone protein dnaJ 58 4e-08
DNJ1_AQUAE (O67623) Chaperone protein dnaJ-1 57 5e-08
DNAJ_CHLTR (O84345) Chaperone protein dnaJ 57 5e-08
DNAJ_CHLMU (Q9PK53) Chaperone protein dnaJ 57 5e-08
DJA4_HUMAN (Q8WW22) DnaJ homolog subfamily A member 4 57 5e-08
DNJM_MYCGE (P47442) DnaJ-like protein MG200 57 6e-08
DNAJ_MYCGE (P47265) Chaperone protein dnaJ 57 6e-08
DJA4_MOUSE (Q9JMC3) DnaJ homolog subfamily A member 4 (MmDjA4) 57 6e-08
DNAJ_MYCPN (P78004) Chaperone protein dnaJ 57 8e-08
>TID_DROVI (Q24331) Tumorous imaginal discs protein, mitochondrial
precursor (Lethal(2)tumorous imaginal discs protein)
(TID58)
Length = 529
Score = 70.9 bits (172), Expect = 4e-12
Identities = 40/112 (35%), Positives = 63/112 (55%), Gaps = 8/112 (7%)
Query: 34 AVSPWRFATPSCKRRGFGRVRVATEQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYY 93
AVS W+ T + + + ++ S +S + A+DYYA LG+ +A + IKKAYY
Sbjct: 49 AVSAWQQRTTAKQHQ-------QQQRRSLFSSSRMQAKDYYATLGVAKNANAKDIKKAYY 101
Query: 94 NCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRVYDDIHGYSLTSIN 145
K HPD + +DP+ + ++E YEVLSD +RR Y D +G + ++N
Sbjct: 102 ELAKKYHPDTNKDDPDASKKFQDVSEAYEVLSDDQKRREY-DTYGQTTENMN 152
>DJA3_MOUSE (Q99M87) DnaJ homolog subfamily A member 3,
mitochondrial precursor (Tumorous imaginal discs protein
Tid56 homolog) (DnaJ protein Tid-1) (mTid-1)
Length = 480
Score = 70.5 bits (171), Expect = 5e-12
Identities = 34/74 (45%), Positives = 48/74 (63%)
Query: 61 SFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEV 120
SF TS S+ +DYY +LG+ +A+ + IKKAYY K HPD + +DP+ + + E
Sbjct: 82 SFHTSASLAKDDYYQILGVPRNASQKDIKKAYYQLAKKYHPDTNKDDPKAKEKFSQLAEA 141
Query: 121 YEVLSDPVQRRVYD 134
YEVLSD V+R+ YD
Sbjct: 142 YEVLSDEVKRKQYD 155
>DJA3_HUMAN (Q96EY1) DnaJ homolog subfamily A member 3,
mitochondrial precursor (Tumorous imaginal discs protein
Tid56 homolog) (DnaJ protein Tid-1) (hTid-1)
Length = 480
Score = 70.5 bits (171), Expect = 5e-12
Identities = 34/74 (45%), Positives = 48/74 (63%)
Query: 61 SFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEV 120
SF TS + EDYY +LG+ +A+ ++IKKAYY K HPD + +DP+ + + E
Sbjct: 82 SFHTSAPLAKEDYYQILGVPRNASQKEIKKAYYQLAKKYHPDTNKDDPKAKEKFSQLAEA 141
Query: 121 YEVLSDPVQRRVYD 134
YEVLSD V+R+ YD
Sbjct: 142 YEVLSDEVKRKQYD 155
>TID_DROME (Q27237) Tumorous imaginal discs protein, mitochondrial
precursor (Lethal(2)tumorous imaginal discs protein)
(TID56) (TID50)
Length = 520
Score = 65.9 bits (159), Expect = 1e-10
Identities = 36/87 (41%), Positives = 48/87 (54%), Gaps = 1/87 (1%)
Query: 49 GFGRVRV-ATEQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGND 107
G G R A + T+ + A+DYYA LG+ +A + IKKAYY K HPD + D
Sbjct: 41 GSGSTRADAPQVRRLHTTRDLLAKDYYATLGVAKNANGKDIKKAYYQLAKKYHPDTNKED 100
Query: 108 PETTNFCTFINEVYEVLSDPVQRRVYD 134
P+ ++E YEVLSD +RR YD
Sbjct: 101 PDAGRKFQEVSEAYEVLSDEQKRREYD 127
>CAJ1_YEAST (P39101) CAJ1 protein
Length = 391
Score = 63.5 bits (153), Expect = 7e-10
Identities = 42/124 (33%), Positives = 57/124 (45%), Gaps = 11/124 (8%)
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRR 131
+YY +LG+ P+ATP +IKKAY HPD +DP+ + E Y+VLSDP R
Sbjct: 6 EYYDILGIKPEATPTEIKKAYRRKAMETHPDKHPDDPDAQAKFQAVGEAYQVLSDPGLRS 65
Query: 132 VYDDIHGYSLTSINPFMDDSSPKDHVF--------VDEFSCIGCKNCANVACDVFGIEED 183
YD F D S +F + EFS N A ++FG E++
Sbjct: 66 KYDQFGKEDAVPQQGFEDASEYFTAIFGGDGFKDWIGEFSLF---KELNEATEMFGKEDE 122
Query: 184 FGRA 187
G A
Sbjct: 123 EGTA 126
>DNJ2_STRCO (Q9RDD7) Chaperone protein dnaJ2
Length = 378
Score = 63.2 bits (152), Expect = 9e-10
Identities = 32/65 (49%), Positives = 44/65 (67%), Gaps = 1/65 (1%)
Query: 70 AEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQ 129
A DYYAVLG+ DA+ ++IKKA+ + HPD++ DP+T IN YEVLSDP +
Sbjct: 2 ATDYYAVLGVRRDASQDEIKKAFRRLARELHPDVN-PDPKTQERFKEINAAYEVLSDPQK 60
Query: 130 RRVYD 134
++VYD
Sbjct: 61 KQVYD 65
>DNJ2_STRAL (O52164) Chaperone protein dnaJ2
Length = 379
Score = 63.2 bits (152), Expect = 9e-10
Identities = 32/65 (49%), Positives = 44/65 (67%), Gaps = 1/65 (1%)
Query: 70 AEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQ 129
A DYYAVLG+ DA+ ++IKKA+ + HPD++ DP+T IN YEVLSDP +
Sbjct: 2 ATDYYAVLGVRRDASQDEIKKAFRRLARELHPDVN-PDPKTQERFKEINAAYEVLSDPQK 60
Query: 130 RRVYD 134
++VYD
Sbjct: 61 KQVYD 65
>DNAJ_THETH (Q56237) Chaperone protein dnaJ
Length = 280
Score = 62.8 bits (151), Expect = 1e-09
Identities = 31/64 (48%), Positives = 42/64 (65%), Gaps = 1/64 (1%)
Query: 71 EDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQR 130
+DYYA+LG+ +AT E+IK+AY + HPD++ PE INE Y VLSDP +R
Sbjct: 5 KDYYAILGVPRNATQEEIKRAYKRLARQYHPDVN-KSPEAEEKFKEINEAYAVLSDPEKR 63
Query: 131 RVYD 134
R+YD
Sbjct: 64 RIYD 67
>DNJ2_AQUAE (O66921) Chaperone protein dnaJ-2
Length = 376
Score = 59.3 bits (142), Expect = 1e-08
Identities = 29/64 (45%), Positives = 42/64 (65%), Gaps = 1/64 (1%)
Query: 71 EDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQR 130
+DYY +LG+ +A+ E+IKKAY ++ HPD+ PE INE Y+VLSDP +R
Sbjct: 7 KDYYEILGVPRNASQEEIKKAYRRLVRKYHPDIC-KKPECEEKFKEINEAYQVLSDPEKR 65
Query: 131 RVYD 134
++YD
Sbjct: 66 KLYD 69
>SCJ1_YEAST (P25303) DnaJ-related protein SCJ1
Length = 404
Score = 58.9 bits (141), Expect = 2e-08
Identities = 27/65 (41%), Positives = 39/65 (59%)
Query: 70 AEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQ 129
A+DYYA+L + DAT ++IK AY K HPD + E + E Y+VLSDP +
Sbjct: 48 AQDYYAILEIDKDATEKEIKSAYRQLSKKYHPDKNAGSEEAHQKFIEVGEAYDVLSDPEK 107
Query: 130 RRVYD 134
+++YD
Sbjct: 108 KKIYD 112
>DNJ1_MYCTU (P07881) Chaperone protein dnaJ1
Length = 395
Score = 57.8 bits (138), Expect = 4e-08
Identities = 25/65 (38%), Positives = 40/65 (61%)
Query: 71 EDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQR 130
+D+Y LG+ DA+PE+IK+AY + HPD + +P ++E + VLSDP +R
Sbjct: 9 KDFYQELGVSSDASPEEIKRAYRKLARDLHPDANPGNPAAGERFKAVSEAHNVLSDPAKR 68
Query: 131 RVYDD 135
+ YD+
Sbjct: 69 KEYDE 73
>DNAJ_THEMA (Q9WZV3) Chaperone protein dnaJ
Length = 369
Score = 57.8 bits (138), Expect = 4e-08
Identities = 36/87 (41%), Positives = 45/87 (51%), Gaps = 7/87 (8%)
Query: 71 EDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDL-SGNDPETTNFCTFINEVYEVLSDPVQ 129
+DYY +LG+ DAT E+IK+AY +K HPD N E I E YEVLSDP +
Sbjct: 6 KDYYEILGVPRDATQEEIKRAYKRLVKEWHPDRHPENRKEAEQRFKEIQEAYEVLSDPQK 65
Query: 130 RRVYD------DIHGYSLTSINPFMDD 150
R +YD + Y T F DD
Sbjct: 66 RAMYDRFGYVGEQPTYQETESGGFFDD 92
>DNJ1_AQUAE (O67623) Chaperone protein dnaJ-1
Length = 364
Score = 57.4 bits (137), Expect = 5e-08
Identities = 34/83 (40%), Positives = 44/83 (52%), Gaps = 3/83 (3%)
Query: 67 SVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSD 126
S +DYY +LG+ DAT E+IKKAY ++ HPD++ DP INE Y VL D
Sbjct: 2 SQAVKDYYEILGVNRDATKEEIKKAYRKLVRIYHPDIN-PDPSAQEKFKEINEAYHVLID 60
Query: 127 PVQRRVYDDIHGYSLTSINPFMD 149
+R YD I S + F D
Sbjct: 61 DERRSEYDAI--LSRNDVGKFRD 81
>DNAJ_CHLTR (O84345) Chaperone protein dnaJ
Length = 392
Score = 57.4 bits (137), Expect = 5e-08
Identities = 28/63 (44%), Positives = 35/63 (55%)
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRR 131
DYY +LG+ ATPE+IKKAY HPD + D E ++E YEVL D +R
Sbjct: 2 DYYTILGVAKTATPEEIKKAYRKLAVKYHPDKNPGDAEAERRFKEVSEAYEVLGDAQKRE 61
Query: 132 VYD 134
YD
Sbjct: 62 SYD 64
>DNAJ_CHLMU (Q9PK53) Chaperone protein dnaJ
Length = 392
Score = 57.4 bits (137), Expect = 5e-08
Identities = 28/63 (44%), Positives = 35/63 (55%)
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRR 131
DYY +LG+ ATPE+IKKAY HPD + D E ++E YEVL D +R
Sbjct: 2 DYYTILGVAKTATPEEIKKAYRKLAVKYHPDKNPGDAEAERRFKEVSEAYEVLGDAQKRE 61
Query: 132 VYD 134
YD
Sbjct: 62 SYD 64
>DJA4_HUMAN (Q8WW22) DnaJ homolog subfamily A member 4
Length = 397
Score = 57.4 bits (137), Expect = 5e-08
Identities = 29/62 (46%), Positives = 38/62 (60%), Gaps = 3/62 (4%)
Query: 73 YYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRV 132
YY +LG+ P A+PE+IKKAY HPD +P+ I++ YEVLSDP +R V
Sbjct: 7 YYDILGVKPSASPEEIKKAYRKLALKYHPD---KNPDEGEKFKLISQAYEVLSDPKKRDV 63
Query: 133 YD 134
YD
Sbjct: 64 YD 65
>DNJM_MYCGE (P47442) DnaJ-like protein MG200
Length = 601
Score = 57.0 bits (136), Expect = 6e-08
Identities = 35/98 (35%), Positives = 48/98 (48%), Gaps = 2/98 (2%)
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRR 131
DYY VLG+ PDA +IKKA+ K HPD N P+ INE +VLS+P +R
Sbjct: 7 DYYEVLGITPDADQSEIKKAFRKLAKKYHPD-RNNAPDAAKIFAEINEANDVLSNPKKRA 65
Query: 132 VYDDIHGYSLTSINPFMDDSSPKDHVFVDEFSCIGCKN 169
YD +G+ P + + F +E + G N
Sbjct: 66 NYDK-YGFDGVDGEPAFNFQADVFQSFFEEIAKSGVFN 102
>DNAJ_MYCGE (P47265) Chaperone protein dnaJ
Length = 389
Score = 57.0 bits (136), Expect = 6e-08
Identities = 40/118 (33%), Positives = 55/118 (45%), Gaps = 26/118 (22%)
Query: 67 SVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETT---NFCTF--INEVY 121
+ G DYY VLG+ +A+ + IK+A+ HPD + ETT N F +NE Y
Sbjct: 2 AAGKRDYYEVLGISKNASSQDIKRAFRKLAMQYHPDRHKAENETTQKQNEEKFKEVNEAY 61
Query: 122 EVLSDPVQRRVYD-------DIHGYSLTSINPF--------------MDDSSPKDHVF 158
EVLSD +R++YD + G+ NPF MD SP D +F
Sbjct: 62 EVLSDEEKRKLYDQFGHEGLNASGFHEAGFNPFDIFNSVFGEGFSFGMDGDSPFDFIF 119
>DJA4_MOUSE (Q9JMC3) DnaJ homolog subfamily A member 4 (MmDjA4)
Length = 397
Score = 57.0 bits (136), Expect = 6e-08
Identities = 28/62 (45%), Positives = 38/62 (61%), Gaps = 3/62 (4%)
Query: 73 YYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRV 132
YY +LG+ P A+PE+IKKAY HPD +P+ I++ YEVLSDP +R +
Sbjct: 7 YYDILGVKPSASPEEIKKAYRKLALKYHPD---KNPDEGEKFKLISQAYEVLSDPKKRDI 63
Query: 133 YD 134
YD
Sbjct: 64 YD 65
>DNAJ_MYCPN (P78004) Chaperone protein dnaJ
Length = 390
Score = 56.6 bits (135), Expect = 8e-08
Identities = 41/118 (34%), Positives = 53/118 (44%), Gaps = 26/118 (22%)
Query: 67 SVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETT---NFCTF--INEVY 121
+ G DYY VLG+ AT + IK+A+ HPD + ET N F +NE Y
Sbjct: 2 AAGKRDYYEVLGVSRSATAQDIKRAFRKLAMQYHPDRHKGEGETVQKQNEEKFKEVNEAY 61
Query: 122 EVLSDPVQRRVYD-------DIHGYSLTSINPF--------------MDDSSPKDHVF 158
EVLSD +R +YD + G+ T NPF MD SP D +F
Sbjct: 62 EVLSDTEKRGMYDRFGHEGLNASGFHETGFNPFDIFNSVFGEGFSFDMDGGSPFDFIF 119
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.135 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,672,305
Number of Sequences: 164201
Number of extensions: 1544310
Number of successful extensions: 4529
Number of sequences better than 10.0: 246
Number of HSP's better than 10.0 without gapping: 200
Number of HSP's successfully gapped in prelim test: 46
Number of HSP's that attempted gapping in prelim test: 4202
Number of HSP's gapped (non-prelim): 260
length of query: 319
length of database: 59,974,054
effective HSP length: 110
effective length of query: 209
effective length of database: 41,911,944
effective search space: 8759596296
effective search space used: 8759596296
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC135160.2


