

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134967.10 + phase: 0 /pseudo
(624 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
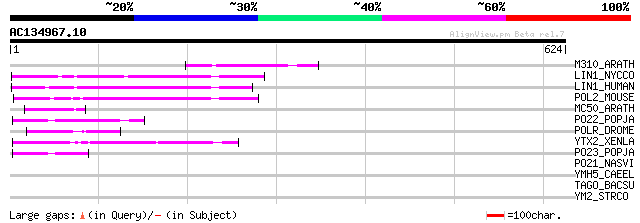
Score E
Sequences producing significant alignments: (bits) Value
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 76 2e-13
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 69 3e-11
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 69 3e-11
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 66 3e-10
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 55 7e-07
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 50 2e-05
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 49 5e-05
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 46 3e-04
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 46 3e-04
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 38 0.070
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 35 0.77
TAGO_BACSU (O34753) Probable undecaprenyl-phosphate N-acetylgluc... 32 6.5
YM2_STRCO (P14706) Mini-circle hypothetical 19.1 kDa protein 31 8.5
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 76.3 bits (186), Expect = 2e-13
Identities = 54/153 (35%), Positives = 74/153 (48%), Gaps = 15/153 (9%)
Query: 198 SLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISWLSWQTVCLKKEC-GGLG 256
+LPVYA+S F+ + + S F+ E+ RKISW++WQ +C KE GGLG
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSS---CENKRKISWVAWQKLCKSKEDDGGLG 58
Query: 257 VRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGRLE--VGGRSVSCWWSEVRR 314
R L FN ALL K +R++ L R+L +RY +E VG R W S +
Sbjct: 59 FRDLGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIH- 117
Query: 315 LRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSDR 347
G +LR +GDG+ T W DR
Sbjct: 118 --------GRELLSRGLLRTIGDGIHTKVWLDR 142
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 69.3 bits (168), Expect = 3e-11
Identities = 70/292 (23%), Positives = 130/292 (43%), Gaps = 25/292 (8%)
Query: 3 KWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQL 62
K I+ S TA++++NG FPL+ G RQG PLSP LF + E L I ++ E +
Sbjct: 627 KLIEAIYSKPTANIILNGVKLKSFPLRSGTRQGCPLSPLLFNIVMEVLAIAIR---EEKA 683
Query: 63 FSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM-L 121
G IG+ + FADD ++ + + L + + +SG K+N HKS+
Sbjct: 684 IKGIHIGSEE---IKLSLFADDMIVYLENTRDSTTKLLEVIKEYSNVSGYKINTHKSVAF 740
Query: 122 VGVNIGSSWLYEAASVLSCKVGIVP--FLYLDMPIGGNPRRLC--FWEPIVNRIKSRLSG 177
+ N + E S +VP YL + + + + L +E + I ++
Sbjct: 741 IYTNNNQA---EKTVKDSIPFTVVPKKMKYLGVYLTKDVKDLYKENYETLRKEIAEDVNK 797
Query: 178 WNSRFLSFGGRLVLLKSVLTSLPVYALSF--FKAPTGIISSIDSLFINFFLGGGGGSEDN 235
W + S+ GR+ ++K + +Y + KAP ++ + ++F N
Sbjct: 798 WKNIPCSWLGRINIVKMSILPKAIYNFNAIPIKAPLSYFKDLEKIILHFIW--------N 849
Query: 236 RKISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRG-GLWYRV 286
+K ++ + K + GG+ + L+ + +++ K W +R +W R+
Sbjct: 850 QKKPQIAKTLLSNKNKAGGITLPDLRLYYKSIVIKTAWYWHKNREVDVWNRI 901
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 69.3 bits (168), Expect = 3e-11
Identities = 63/275 (22%), Positives = 117/275 (41%), Gaps = 18/275 (6%)
Query: 3 KWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQL 62
K I+ TA++++NG + PLK G RQG PLSP LL L ++ +++ + +
Sbjct: 627 KIIRAIYDKPTANIILNGQKLEAPPLKTGTRQGCPLSP---LLPNIVLEVLARAIRQEKE 683
Query: 63 FSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLV 122
G +G V FADD ++ + + L + F +SG K+N KS
Sbjct: 684 IKGIQLGKEE---VKLSLFADDMIVYLENPIVSAQNLLKLISNFSKVSGYKINVQKSQAF 740
Query: 123 GVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLC--FWEPIVNRIKSRLSGWNS 180
+ S L + YL + + + + L ++P++N IK + W +
Sbjct: 741 LYTNNRQTESQIMSELPFTIASKRIKYLGIQLTRDVKDLFKENYKPLLNEIKEDTNKWKN 800
Query: 181 RFLSFGGRLVLLKSVLTSLPVYALSF--FKAPTGIISSIDSLFINFFLGGGGGSEDNRKI 238
S+ GR+ ++K + +Y + K P + ++ + F N+K
Sbjct: 801 IPCSWVGRINIVKMAILPKVIYRFNAIPIKLPMTFFTELEKTTLKFIW--------NQKR 852
Query: 239 SWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCW 273
+ ++ T+ K + GG+ + K + A + K W
Sbjct: 853 AHIAKSTLSQKNKAGGITLPDFKLYYKATVTKTAW 887
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 65.9 bits (159), Expect = 3e-10
Identities = 68/279 (24%), Positives = 115/279 (40%), Gaps = 18/279 (6%)
Query: 5 IKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLFS 64
IK S A++ VNG + PLK G RQG PLSP+LF + L ++ +++ + +
Sbjct: 656 IKAIYSKPVANIKVNGEKLEAIPLKSGTRQGCPLSPYLFNIV---LEVLARAIRQQKEIK 712
Query: 65 GYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLVGV 124
G IG + +S L ADD ++ S + R L + F + G K+N +KSM
Sbjct: 713 GIQIG-KEEVKISLL--ADDMIVYISDPKNSTRELLNLINSFGEVVGYKINSNKSMAFLY 769
Query: 125 NIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLC--FWEPIVNRIKSRLSGWNSRF 182
E + YL + + + L ++ + IK L W
Sbjct: 770 TKNKQAEKEIRETTPFSIVTNNIKYLGVTLTKEVKDLYDKNFKSLKKEIKEDLRRWKDLP 829
Query: 183 LSFGGRLVLLKSVLTSLPVYALSF--FKAPTGIISSIDSLFINFFLGGGGGSEDNRKISW 240
S+ GR+ ++K + +Y + K PT + ++ F N K
Sbjct: 830 CSWIGRINIVKMAILPKAIYRFNAIPIKIPTQFFNELEGAICKFVW--------NNKKPR 881
Query: 241 LSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDR 279
++ + K+ GG+ + LK + A++ K W DR
Sbjct: 882 IAKSLLKDKRTSGGITMPDLKLYYRAIVIKTAWYWYRDR 920
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 54.7 bits (130), Expect = 7e-07
Identities = 29/69 (42%), Positives = 41/69 (59%), Gaps = 1/69 (1%)
Query: 17 LVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLFSGYSIGAANPIVV 76
++NG+P RGLRQGDPLSP+LF+L E L+ + + E G + +P +
Sbjct: 13 IINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSP-RI 71
Query: 77 SHLQFADDT 85
+HL FADDT
Sbjct: 72 NHLLFADDT 80
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 49.7 bits (117), Expect = 2e-05
Identities = 47/149 (31%), Positives = 69/149 (45%), Gaps = 18/149 (12%)
Query: 4 WIKECVSTATASVLVN-GSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQL 62
+I +S +T ++ V GS T + ++RG++QGDPLSPFLF N V+ L+ +
Sbjct: 172 YITGSLSDSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLF-------NAVLDELLCSLQ 224
Query: 63 FSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLV 122
+ G + L FADD LLL L A + F + G+ +N KS+ +
Sbjct: 225 STPGIGGTIGEEKIPVLAFADDLLLLEDNDVLLPTTL-ATVANFFRLRGMSLNAKKSVSI 283
Query: 123 GVNIGSSWLYEAASVLSCKVGIVPFLYLD 151
V AAS C PFL +D
Sbjct: 284 SV---------AASGGVCIPRTKPFLRVD 303
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 48.5 bits (114), Expect = 5e-05
Identities = 35/106 (33%), Positives = 58/106 (54%), Gaps = 10/106 (9%)
Query: 19 NGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLFSGYSIGAANPIVVSH 78
+G ++EF RG++QGDPLSP LF L + L + S + A++ + + AA
Sbjct: 526 DGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRTLPSEIGAKVGNAITNAAA------- 578
Query: 79 LQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLVGV 124
FADD L+L +++ ++ L + F ++ GLK+N K VG+
Sbjct: 579 --FADD-LVLFAETRMGLQVLLDKTLDFLSIVGLKLNADKCFTVGI 621
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 45.8 bits (107), Expect = 3e-04
Identities = 59/257 (22%), Positives = 112/257 (42%), Gaps = 20/257 (7%)
Query: 4 WIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLF 63
++K ++A V +N S T RG+RQG PLS L+ LA E +++ + +
Sbjct: 623 YLKTMYASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSLAIEPFLCLLRKRLTGLVL 682
Query: 64 SGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM-LV 122
+ +V+S +ADD +L+ ++ ++ + ++ A S ++N+ KS L+
Sbjct: 683 KEPDM----RVVLS--AYADDVILV-AQDLVDLERAQECQEVYAAASSARINWSKSSGLL 735
Query: 123 GVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWN--S 180
++ +L A +S + I+ +L + + P F E + + +RL W +
Sbjct: 736 EGSLKVDFLPPAFRDISWESKIIKYLGVYLSAEEYPVSQNFIE-LEECVLTRLGKWKGFA 794
Query: 181 RFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISW 240
+ LS GR +++ ++ S Y L I+ I ++F G W
Sbjct: 795 KVLSMRGRALVINQLVASQIWYRLICLSPTQEFIAKIQRRLLDFLWIGK---------HW 845
Query: 241 LSWQTVCLKKECGGLGV 257
+S L + GG GV
Sbjct: 846 VSAGVSSLPLKEGGQGV 862
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 45.8 bits (107), Expect = 3e-04
Identities = 28/85 (32%), Positives = 43/85 (49%), Gaps = 7/85 (8%)
Query: 4 WIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLF 63
+I ++ A +++V G T++ ++ G++QGDPLSP LF NIV+ LV
Sbjct: 58 FIMSTITDAYTNIVVGGRTTNKIYIRNGVKQGDPLSPVLF-------NIVLDELVTRLND 110
Query: 64 SGYSIGAANPIVVSHLQFADDTLLL 88
++ L FADD LLL
Sbjct: 111 EQPGASMTPACKIASLAFADDLLLL 135
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 38.1 bits (87), Expect = 0.070
Identities = 30/79 (37%), Positives = 42/79 (52%), Gaps = 13/79 (16%)
Query: 30 RGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLFSGYSIGAANPIVVSHLQFADDTLLLG 89
RG+RQGDPLSP LF ++ V++ L E +G+ +GA + L FADD +LL
Sbjct: 511 RGVRQGDPLSPLLFNCV---MDAVLRRLPEN---TGFLMGAEK---IGALVFADDLVLLA 561
Query: 90 SKSWANVRALRAALVLFEA 108
L+A+L EA
Sbjct: 562 ETR----EGLQASLSRIEA 576
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 34.7 bits (78), Expect = 0.77
Identities = 28/90 (31%), Positives = 43/90 (47%), Gaps = 5/90 (5%)
Query: 4 WIKECVSTATASVLVNGS-PTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQL 62
W KE + T SV +N ++ +P+ G+ QG P LF+L L I ++ +
Sbjct: 783 WFKEFLHLRTFSVKINKFVSSNAYPISSGVPQGSVSGPLLFILFINDLLIDLEPNIHVSC 842
Query: 63 FS-GYSIGAANPIVVSHLQFADDTLLLGSK 91
F+ I NP S LQ + DT++ SK
Sbjct: 843 FADDIKIFHHNP---STLQNSIDTIVKWSK 869
>TAGO_BACSU (O34753) Probable undecaprenyl-phosphate
N-acetylglucosaminyl 1-phosphate transferase (EC
2.7.8.-) (UDP-GlcNAc:undecaprenyl-P GlcNAc 1-P
transferase)
Length = 358
Score = 31.6 bits (70), Expect = 6.5
Identities = 37/157 (23%), Positives = 73/157 (45%), Gaps = 24/157 (15%)
Query: 85 TLLLGSKSWANVRALRAALVLFEAMSG-LKVNFHKSMLVGVNIGSSWLYEAASVLSCKVG 143
T+ + + S V L +LV+ + G L NFH + + + GS +L + S+LS +G
Sbjct: 175 TIAVMALSGGKVLILSLSLVVIASTLGFLFYNFHPAKIFMGDTGSLFLGYSISILSL-LG 233
Query: 144 ----------IVPFLYLDMPIGGNP----RRLCFWEPIVNRIKSRLSGWNSRFLSFGGRL 189
++P + L +PI RR+ +PI KS + + R ++FG
Sbjct: 234 LYKSVTLFSIVIPIIILGVPIFDTTFAIIRRILNKQPISAPDKSHI---HHRLMAFG--- 287
Query: 190 VLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFL 226
L ++ L +Y + F + + I+ +++++ F+
Sbjct: 288 --LSHRMSVLVIYLIGFIFSISAIVLKSATIWLSLFI 322
>YM2_STRCO (P14706) Mini-circle hypothetical 19.1 kDa protein
Length = 175
Score = 31.2 bits (69), Expect = 8.5
Identities = 35/126 (27%), Positives = 55/126 (42%), Gaps = 22/126 (17%)
Query: 239 SWLSWQTVCLKKECGGLGVRKLK-----EFNLALLGKWCWRMLVDRGGLWYRVLVAR--- 290
+WL + L +C GL +L+ ++ LLG V+R W++ + A
Sbjct: 23 AWLDFHRATLAVKCSGLKDDQLRLAAAAPSSMTLLGLVQHMAEVERN--WFQRVFAGQNV 80
Query: 291 ---YGEL---GGRLEVG---GRSVSCWWSEVRRLRDGVGDGG---SGWFGDQVLRRVGDG 338
+GE G L+ G +V+ W +EV R R+ + D G SG +Q VGD
Sbjct: 81 PPVFGESNHDGFALKSGRGLDEAVAAWQAEVSRGRELIADAGLDDSGHLSEQEAGHVGDQ 140
Query: 339 VVTSFW 344
V+ W
Sbjct: 141 GVSLRW 146
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.348 0.155 0.570
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 68,112,332
Number of Sequences: 164201
Number of extensions: 2758127
Number of successful extensions: 9393
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 9369
Number of HSP's gapped (non-prelim): 16
length of query: 624
length of database: 59,974,054
effective HSP length: 116
effective length of query: 508
effective length of database: 40,926,738
effective search space: 20790782904
effective search space used: 20790782904
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC134967.10


