

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.2 + phase: 0 /pseudo
(840 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
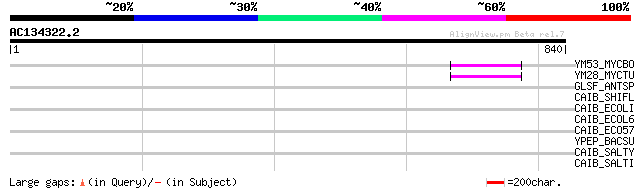
Score E
Sequences producing significant alignments: (bits) Value
YM53_MYCBO (P64956) Hypothetical protein Mb2253c 50 2e-05
YM28_MYCTU (P64955) Hypothetical protein Rv2228c/MT2287 50 2e-05
GLSF_ANTSP (Q06434) Ferredoxin-dependent glutamate synthase (EC ... 36 0.37
CAIB_SHIFL (P59394) Crotonobetainyl-CoA:carnitine CoA-transferas... 33 4.1
CAIB_ECOLI (P31572) Crotonobetainyl-CoA:carnitine CoA-transferas... 33 4.1
CAIB_ECOL6 (Q8FLA4) Crotonobetainyl-CoA:carnitine CoA-transferas... 33 4.1
CAIB_ECO57 (Q8XA32) Crotonobetainyl-CoA:carnitine CoA-transferas... 33 4.1
YPEP_BACSU (P54164) Hypothetical protein ypeP 32 7.0
CAIB_SALTY (Q8ZRX3) Crotonobetainyl-CoA:carnitine CoA-transferas... 32 9.1
CAIB_SALTI (Q8Z9L3) Crotonobetainyl-CoA:carnitine CoA-transferas... 32 9.1
>YM53_MYCBO (P64956) Hypothetical protein Mb2253c
Length = 364
Score = 50.4 bits (119), Expect = 2e-05
Identities = 33/110 (30%), Positives = 51/110 (46%), Gaps = 3/110 (2%)
Query: 668 KWVLSVDGSSNLKGSGAG---VTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLAL 724
K V+ DG S AG V + ++ +S + +++NN AEY L AG+ A+
Sbjct: 2 KVVIEADGGSRGNPGPAGYGAVVWTADHSTVLAESKQAIGRATNNVAEYRGLIAGLDDAV 61
Query: 725 KVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQKFVSFKY 774
K+ E DS+LV Q++G ++ K P LL L +F Y
Sbjct: 62 KLGATEAAVLMDSKLVVEQMSGRWKVKHPDLLKLYVQAQALASQFRRINY 111
>YM28_MYCTU (P64955) Hypothetical protein Rv2228c/MT2287
Length = 364
Score = 50.4 bits (119), Expect = 2e-05
Identities = 33/110 (30%), Positives = 51/110 (46%), Gaps = 3/110 (2%)
Query: 668 KWVLSVDGSSNLKGSGAG---VTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLAL 724
K V+ DG S AG V + ++ +S + +++NN AEY L AG+ A+
Sbjct: 2 KVVIEADGGSRGNPGPAGYGAVVWTADHSTVLAESKQAIGRATNNVAEYRGLIAGLDDAV 61
Query: 725 KVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQKFVSFKY 774
K+ E DS+LV Q++G ++ K P LL L +F Y
Sbjct: 62 KLGATEAAVLMDSKLVVEQMSGRWKVKHPDLLKLYVQAQALASQFRRINY 111
>GLSF_ANTSP (Q06434) Ferredoxin-dependent glutamate synthase (EC
1.4.7.1) (Fd-GOGAT)
Length = 1536
Score = 36.2 bits (82), Expect = 0.37
Identities = 18/51 (35%), Positives = 31/51 (60%), Gaps = 3/51 (5%)
Query: 675 GSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALK 725
G+ N+ G GAGVT++ P+ + I + + F K +NQ+ L G+R+ L+
Sbjct: 61 GADNISGDGAGVTIQIPWDIFISEGINFLPKLQSNQS---ILNYGVRMILR 108
>CAIB_SHIFL (P59394) Crotonobetainyl-CoA:carnitine CoA-transferase
(EC 2.8.3.-)
Length = 405
Score = 32.7 bits (73), Expect = 4.1
Identities = 21/65 (32%), Positives = 36/65 (55%), Gaps = 1/65 (1%)
Query: 688 LEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGN 747
+E PYG L+E+ L + +++ AE + FA + +A V+ E +S+ Q V R+
Sbjct: 284 IECPYGPLVEEKLD-AWLAAHTIAEVKERFAELNIACAKVLTVPELESNPQYVARESITQ 342
Query: 748 FQTKD 752
+QT D
Sbjct: 343 WQTMD 347
>CAIB_ECOLI (P31572) Crotonobetainyl-CoA:carnitine CoA-transferase
(EC 2.8.3.-)
Length = 405
Score = 32.7 bits (73), Expect = 4.1
Identities = 21/65 (32%), Positives = 36/65 (55%), Gaps = 1/65 (1%)
Query: 688 LEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGN 747
+E PYG L+E+ L + +++ AE + FA + +A V+ E +S+ Q V R+
Sbjct: 284 IECPYGPLVEEKLD-AWLATHTIAEVKERFAELNIACAKVLTVPELESNPQYVARESITQ 342
Query: 748 FQTKD 752
+QT D
Sbjct: 343 WQTMD 347
>CAIB_ECOL6 (Q8FLA4) Crotonobetainyl-CoA:carnitine CoA-transferase
(EC 2.8.3.-)
Length = 405
Score = 32.7 bits (73), Expect = 4.1
Identities = 21/65 (32%), Positives = 36/65 (55%), Gaps = 1/65 (1%)
Query: 688 LEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGN 747
+E PYG L+E+ L + +++ AE + FA + +A V+ E +S+ Q V R+
Sbjct: 284 IECPYGPLVEEKLD-AWLAAHTIAEVKERFAELNIACAKVLTVPELESNPQYVARESITQ 342
Query: 748 FQTKD 752
+QT D
Sbjct: 343 WQTMD 347
>CAIB_ECO57 (Q8XA32) Crotonobetainyl-CoA:carnitine CoA-transferase
(EC 2.8.3.-)
Length = 405
Score = 32.7 bits (73), Expect = 4.1
Identities = 21/65 (32%), Positives = 36/65 (55%), Gaps = 1/65 (1%)
Query: 688 LEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGN 747
+E PYG L+E+ L + +++ AE + FA + +A V+ E +S+ Q V R+
Sbjct: 284 IECPYGPLVEEKLD-AWLAAHTIAEVKERFAELNIACAKVLTVPELESNPQYVARESITQ 342
Query: 748 FQTKD 752
+QT D
Sbjct: 343 WQTMD 347
>YPEP_BACSU (P54164) Hypothetical protein ypeP
Length = 120
Score = 32.0 bits (71), Expect = 7.0
Identities = 19/63 (30%), Positives = 32/63 (50%), Gaps = 2/63 (3%)
Query: 707 SNNQAEYEALFAGMR--LALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMY 764
+NN+AEY AL+ +R L + K DS +V Q+ G++ DP +L+ +
Sbjct: 11 TNNEAEYAALYEAIREVRELGASRNSITIKGDSLVVLNQLDGSWPCYDPSHNEWLDKIEA 70
Query: 765 LTQ 767
L +
Sbjct: 71 LLE 73
>CAIB_SALTY (Q8ZRX3) Crotonobetainyl-CoA:carnitine CoA-transferase
(EC 2.8.3.-)
Length = 405
Score = 31.6 bits (70), Expect = 9.1
Identities = 20/65 (30%), Positives = 36/65 (54%), Gaps = 1/65 (1%)
Query: 688 LEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGN 747
+E PYG L+E+ L + +++ A+ +A FA + +A V+ E + + Q V R+
Sbjct: 284 VECPYGPLVEEKLD-AWLATHTIADVQARFAELNIACAKVLTIPELEGNPQYVARESITQ 342
Query: 748 FQTKD 752
+QT D
Sbjct: 343 WQTMD 347
>CAIB_SALTI (Q8Z9L3) Crotonobetainyl-CoA:carnitine CoA-transferase
(EC 2.8.3.-)
Length = 405
Score = 31.6 bits (70), Expect = 9.1
Identities = 20/65 (30%), Positives = 36/65 (54%), Gaps = 1/65 (1%)
Query: 688 LEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGN 747
+E PYG L+E+ L + +++ A+ +A FA + +A V+ E + + Q V R+
Sbjct: 284 VECPYGPLVEEKLD-AWLAAHTIADVQARFAELNIACAKVLTIPELEGNPQYVARESITQ 342
Query: 748 FQTKD 752
+QT D
Sbjct: 343 WQTMD 347
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.358 0.156 0.572
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 78,403,850
Number of Sequences: 164201
Number of extensions: 2655390
Number of successful extensions: 9801
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 9797
Number of HSP's gapped (non-prelim): 10
length of query: 840
length of database: 59,974,054
effective HSP length: 119
effective length of query: 721
effective length of database: 40,434,135
effective search space: 29153011335
effective search space used: 29153011335
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC134322.2


