

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.16 - phase: 0
(690 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
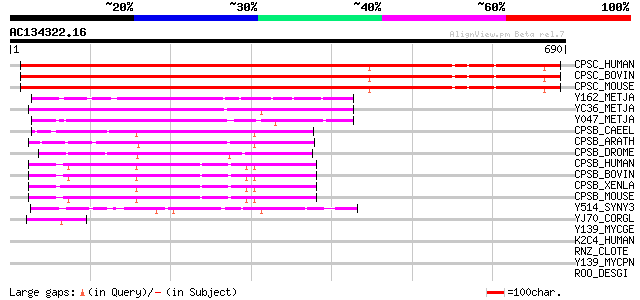
Score E
Sequences producing significant alignments: (bits) Value
CPSC_HUMAN (Q9UKF6) Cleavage and polyadenylation specificity fac... 709 0.0
CPSC_BOVIN (P79101) Cleavage and polyadenylation specificity fac... 709 0.0
CPSC_MOUSE (Q9QXK7) Cleavage and polyadenylation specificity fac... 706 0.0
Y162_METJA (Q57626) Hypothetical protein MJ0162 190 1e-47
YC36_METJA (Q58633) Hypothetical protein MJ1236 189 3e-47
Y047_METJA (Q60355) Hypothetical protein MJ0047 142 3e-33
CPSB_CAEEL (O17403) Probable cleavage and polyadenylation specif... 137 1e-31
CPSB_ARATH (Q9LKF9) Cleavage and polyadenylation specificity fac... 136 2e-31
CPSB_DROME (Q9V3D6) Probable cleavage and polyadenylation specif... 130 2e-29
CPSB_HUMAN (Q9P2I0) Cleavage and polyadenylation specificity fac... 126 2e-28
CPSB_BOVIN (Q10568) Cleavage and polyadenylation specificity fac... 126 2e-28
CPSB_XENLA (Q9W799) Cleavage and polyadenylation specificity fac... 124 8e-28
CPSB_MOUSE (O35218) Cleavage and polyadenylation specificity fac... 124 1e-27
Y514_SYNY3 (Q55470) Hypothetical protein sll0514 80 2e-14
YJ70_CORGL (P54122) Hypothetical UPF0036 protein Cgl1970/cg2160 45 6e-04
Y139_MYCGE (P47385) Hypothetical UPF0036 protein MG139 43 0.003
K2C4_HUMAN (P19013) Keratin, type II cytoskeletal 4 (Cytokeratin... 42 0.004
RNZ_CLOTE (Q892B5) Ribonuclease Z (EC 3.1.26.11) (RNase Z) (tRNA... 42 0.005
Y139_MYCPN (P75497) Hypothetical UPF0036 protein MG139 homolog (... 42 0.007
ROO_DESGI (Q9F0J6) Rubredoxin-oxygen oxidoreductase (EC 1.-.-.-)... 42 0.007
>CPSC_HUMAN (Q9UKF6) Cleavage and polyadenylation specificity
factor, 73 kDa subunit (CPSF 73 kDa subunit)
Length = 684
Score = 709 bits (1830), Expect = 0.0
Identities = 360/685 (52%), Positives = 479/685 (69%), Gaps = 18/685 (2%)
Query: 14 INRETEDQLIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPST 73
I E DQL++ PLGAG EVGRSC+ + +KG+ ++ DCGIHPG GM ALPY D IDP+
Sbjct: 4 IPAEESDQLLIRPLGAGQEVGRSCIILEFKGRKIMLDCGIHPGLEGMDALPYIDLIDPAE 63
Query: 74 VDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLY 133
+D+LLI+HFHLDH +LP+FL+KT+FKGR FMT+ATKAIY+ LLSDYVKVS +S DDMLY
Sbjct: 64 IDLLLISHFHLDHCGALPWFLQKTSFKGRTFMTHATKAIYRWLLSDYVKVSNISADDMLY 123
Query: 134 DEQDINRSMDKIEVIDFHQTVEVNGIRFWCYTAGHVLGAAMFMVDIAGVRVLYTGDYSRE 193
E D+ SMDKIE I+FH+ EV GI+FWCY AGHVLGAAMFM++IAGV++LYTGD+SR+
Sbjct: 124 TETDLEESMDKIETINFHEVKEVAGIKFWCYHAGHVLGAAMFMIEIAGVKLLYTGDFSRQ 183
Query: 194 EDRHLRAAETPQFSPDVCIIESTYGVQHHQPRHTREKRFTDVIHSTISQGGRVLIPAYAL 253
EDRHL AAE P PD+ IIESTYG H+ R RE RF + +H +++GGR LIP +AL
Sbjct: 184 EDRHLMAAEIPNIKPDILIIESTYGTHIHEKREEREARFCNTVHDIVNRGGRGLIPVFAL 243
Query: 254 GRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRI--QNAKSNPF 311
GRAQELLLILDEYW NHPEL +IPIYYAS LAKKC+ VY+TY +MND+I Q +NPF
Sbjct: 244 GRAQELLLILDEYWQNHPELHDIPIYYASSLAKKCMAVYQTYVNAMNDKIRKQININNPF 303
Query: 312 AFKHISALSSIDIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEGTL 371
FKHIS L S+D F D+GPSVVMASPG +QSGLSR+LF+ WC+DK+N +I GY VEGTL
Sbjct: 304 VFKHISNLKSMDHFDDIGPSVVMASPGMMQSGLSRELFESWCTDKRNGVIIAGYCVEGTL 363
Query: 372 AKTILNEPKEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGAANE 431
AK I++EP+E+T M+G PL M V YISFSAH D QTS F+ L PP++ILVHG NE
Sbjct: 364 AKHIMSEPEEITTMSGQKLPLKMSVDYISFSAHTDYQQTSEFIRALKPPHVILVHGEQNE 423
Query: 432 MGRLKQKLMTQFADR---NTKILTPKNCQSVEMYFNSQKMAKTIGKLAEKTPEVGETVSG 488
M RLK L+ ++ D + ++ P+N ++V + F +K+AK +G LA+K PE G+ VSG
Sbjct: 424 MARLKAALIREYEDNDEVHIEVHNPRNTEAVTLNFRGEKLAKVMGFLADKKPEQGQRVSG 483
Query: 489 LLVKKGFTYQIMAPDDLHVFSQLSTANVTQRITIPYSGAFCVIQSRLKQIYESVEPSVDE 548
+LVK+ F Y I++P DL ++ L+ + V Q IPY+G F ++ +L+++ VE +
Sbjct: 484 ILVKRNFNYHILSPCDLSNYTDLAMSTVKQTQAIPYTGPFNLLCYQLQKLTGDVEELEIQ 543
Query: 549 ESGVPMLLVHDRVTVKHESEKHVSLHWASDPINDMVSDSVVALVLNINRDLPKIVAESDA 608
E P L V +TV E V L W ++P NDM +D+V ++L + + PKI +
Sbjct: 544 EK--PALKVFKNITVIQEPGM-VVLEWLANPSNDMYADTVTTVILEVQSN-PKI-RKGAV 598
Query: 609 TKIEEENEKKT-EKVMQALLNSLFGN--VKVGENGKLIINIDGNVAELNKESGEVESE-- 663
K+ ++ E K ++ +L +FG V V ++ L + +DG A LN E+ VE E
Sbjct: 599 QKVSKKLEMHVYSKRLEIMLQDIFGEDCVSVKDDSILSVTVDGKTANLNLETRTVECEEG 658
Query: 664 ---NEGLKERVRTAFRRIQSSVKPI 685
+E L+E V A +R+ ++ P+
Sbjct: 659 SEDDESLREMVELAAQRLYEALTPV 683
>CPSC_BOVIN (P79101) Cleavage and polyadenylation specificity
factor, 73 kDa subunit (CPSF 73 kDa subunit)
Length = 684
Score = 709 bits (1829), Expect = 0.0
Identities = 359/685 (52%), Positives = 478/685 (69%), Gaps = 18/685 (2%)
Query: 14 INRETEDQLIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPST 73
I E DQL++ PLGAG EVGRSC+ + +KG+ ++ DCGIHPG GM ALPY D IDP+
Sbjct: 4 IPAEESDQLLIRPLGAGQEVGRSCIILEFKGRKIMLDCGIHPGLEGMDALPYIDLIDPAE 63
Query: 74 VDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLY 133
+D+LLI+HFHLDH +LP+FL+KT+FKGR FMT+ATKAIY+ LLSDYVKVS +S DDMLY
Sbjct: 64 IDLLLISHFHLDHCGALPWFLQKTSFKGRTFMTHATKAIYRWLLSDYVKVSNISADDMLY 123
Query: 134 DEQDINRSMDKIEVIDFHQTVEVNGIRFWCYTAGHVLGAAMFMVDIAGVRVLYTGDYSRE 193
E D+ SMDKIE I+FH+ EV GI+FWCY AGHVLGAAMFM++IAGV++LYTGD+SR+
Sbjct: 124 TETDLEESMDKIETINFHEVKEVAGIKFWCYHAGHVLGAAMFMIEIAGVKLLYTGDFSRQ 183
Query: 194 EDRHLRAAETPQFSPDVCIIESTYGVQHHQPRHTREKRFTDVIHSTISQGGRVLIPAYAL 253
EDRHL AAE P PD+ IIESTYG H+ R RE RF + +H +++GGR LIP +AL
Sbjct: 184 EDRHLMAAEIPNIKPDILIIESTYGTHIHEKREEREARFCNTVHDIVNRGGRGLIPVFAL 243
Query: 254 GRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRI--QNAKSNPF 311
GRAQELLLILDEYW NHPEL +IPIYYAS LAKKC+ VY+TY +MND+I Q +NPF
Sbjct: 244 GRAQELLLILDEYWQNHPELHDIPIYYASSLAKKCMAVYQTYVNAMNDKIRKQININNPF 303
Query: 312 AFKHISALSSIDIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEGTL 371
FKHIS L S+D F D+GPSVVMASPG +QSGLSR+LF+ WC+DK+N +I GY VEGTL
Sbjct: 304 VFKHISNLKSMDHFDDIGPSVVMASPGMMQSGLSRELFESWCTDKRNGVIIAGYCVEGTL 363
Query: 372 AKTILNEPKEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGAANE 431
AK I++EP+E+T M+G PL M V YISFSAH D QTS F+ L PP++ILVHG NE
Sbjct: 364 AKHIMSEPEEITTMSGQKLPLKMSVDYISFSAHTDYQQTSEFIRALKPPHVILVHGEQNE 423
Query: 432 MGRLKQKLMTQFADR---NTKILTPKNCQSVEMYFNSQKMAKTIGKLAEKTPEVGETVSG 488
M RLK L+ ++ D + ++ P+N ++V + F +K+AK +G LA+K PE G+ VSG
Sbjct: 424 MARLKAALIREYEDNDEVHIEVHNPRNTEAVTLNFRGEKLAKVMGFLADKKPEQGQRVSG 483
Query: 489 LLVKKGFTYQIMAPDDLHVFSQLSTANVTQRITIPYSGAFCVIQSRLKQIYESVEPSVDE 548
+LVK+ F Y I++P DL ++ L+ + V Q IPY+G F ++ +L+++ VE +
Sbjct: 484 ILVKRNFNYHILSPCDLSNYTDLAMSTVKQTQAIPYTGPFNLLYYQLQKLTGDVEELEIQ 543
Query: 549 ESGVPMLLVHDRVTVKHESEKHVSLHWASDPINDMVSDSVVALVLNINRDLPKIVAESDA 608
E P L V +TV E V L W ++P NDM +D+V ++L + + PKI +
Sbjct: 544 EK--PALKVFKNITVIQEPGM-VVLEWLANPSNDMYADTVTTVILEVQSN-PKI-RKGAV 598
Query: 609 TKIEEENEKKT-EKVMQALLNSLFGN--VKVGENGKLIINIDGNVAELNKESGEVESE-- 663
K+ ++ E K ++ +L +FG V V + L + +DG A +N E+ VE E
Sbjct: 599 QKVSKKLEMHVYSKRLEIMLQDIFGEDCVSVKDGSILSVTVDGKTANINLETRTVECEEG 658
Query: 664 ---NEGLKERVRTAFRRIQSSVKPI 685
+E L+E V A +R+ ++ P+
Sbjct: 659 SEDDESLREMVELAAQRLYEALTPV 683
>CPSC_MOUSE (Q9QXK7) Cleavage and polyadenylation specificity
factor, 73 kDa subunit (CPSF 73 kDa subunit)
Length = 684
Score = 706 bits (1822), Expect = 0.0
Identities = 357/685 (52%), Positives = 477/685 (69%), Gaps = 18/685 (2%)
Query: 14 INRETEDQLIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPST 73
I E DQL++ PLGAG EVGRSC+ + +KG+ ++ DCGIHPG GM ALPY D IDP+
Sbjct: 4 IPAEESDQLLIRPLGAGQEVGRSCIILEFKGRKIMLDCGIHPGLEGMDALPYIDLIDPAE 63
Query: 74 VDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLY 133
+D+LLI+HFHLDH +LP+FL+KT+FKGR FMT+ATKAIY+ LLSDYVKVS +S DDMLY
Sbjct: 64 IDLLLISHFHLDHCGALPWFLQKTSFKGRTFMTHATKAIYRWLLSDYVKVSNISADDMLY 123
Query: 134 DEQDINRSMDKIEVIDFHQTVEVNGIRFWCYTAGHVLGAAMFMVDIAGVRVLYTGDYSRE 193
E D+ SMDKIE I+FH+ EV GI+FWCY AGHVLGAAMFM++IAGV++LYTGD+SR+
Sbjct: 124 TETDLEESMDKIETINFHEVKEVAGIKFWCYHAGHVLGAAMFMIEIAGVKLLYTGDFSRQ 183
Query: 194 EDRHLRAAETPQFSPDVCIIESTYGVQHHQPRHTREKRFTDVIHSTISQGGRVLIPAYAL 253
EDRHL AAE P PD+ IIESTYG H+ R RE RF +H +++GGR LIP +AL
Sbjct: 184 EDRHLMAAEIPNIKPDILIIESTYGTHIHEKREEREARFWHTVHDIVNRGGRGLIPVFAL 243
Query: 254 GRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRI--QNAKSNPF 311
GRAQELLLILDEYW NHPEL +IPIYYAS LAKKC+ VY+TY +MND+I Q +NPF
Sbjct: 244 GRAQELLLILDEYWQNHPELHDIPIYYASSLAKKCMAVYQTYVNAMNDKIRKQININNPF 303
Query: 312 AFKHISALSSIDIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEGTL 371
FKHIS L S+D F D+GPSVVMASPG +Q+GLSR+LF+ WC+DK+N +I GY VEGTL
Sbjct: 304 VFKHISNLKSMDHFDDIGPSVVMASPGMIQNGLSRELFESWCTDKRNGVIIAGYCVEGTL 363
Query: 372 AKTILNEPKEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGAANE 431
AK I++EP+E+T M+G PL M V YISFSAH D QTS F+ L PP++ILVHG NE
Sbjct: 364 AKHIMSEPEEITTMSGQKLPLKMSVDYISFSAHTDYQQTSEFIRALKPPHVILVHGEQNE 423
Query: 432 MGRLKQKLMTQFADR---NTKILTPKNCQSVEMYFNSQKMAKTIGKLAEKTPEVGETVSG 488
M RLK L+ ++ D + ++ P+N ++V + F +K+AK +G LA+K PE G+ VSG
Sbjct: 424 MARLKAALIREYEDNDEVHIEVHNPRNTEAVTLNFRGEKLAKVMGFLADKKPEQGQRVSG 483
Query: 489 LLVKKGFTYQIMAPDDLHVFSQLSTANVTQRITIPYSGAFCVIQSRLKQIYESVEPSVDE 548
+LVK+ F Y I++P DL ++ L+ + V Q IPY+G F ++ +L+++ VE +
Sbjct: 484 ILVKRNFNYHILSPCDLSNYTDLAMSTVKQTQAIPYTGPFYLLYYQLQKLTGDVEELDIQ 543
Query: 549 ESGVPMLLVHDRVTVKHESEKHVSLHWASDPINDMVSDSVVALVLNINRDLPKIVAESDA 608
E P L V +TV E V W ++P NDM +D+V ++L + + PKI +
Sbjct: 544 EK--PALKVFKSITVVQEPGM-VGSEWLANPSNDMYADTVTTVILEVQSN-PKI-RKGAV 598
Query: 609 TKIEEENEKKT-EKVMQALLNSLFGN--VKVGENGKLIINIDGNVAELNKESGEVESE-- 663
K+ ++ E K ++ +L +FG V V ++ L + +DG A +N E+ VE E
Sbjct: 599 QKVSKKLEMHVYSKRLEVMLQDIFGEDCVSVKDDSVLSVTVDGKTANINLETRAVECEEG 658
Query: 664 ---NEGLKERVRTAFRRIQSSVKPI 685
+E L+E V A +R+ ++ P+
Sbjct: 659 SEDDESLREMVELAAQRLYEALTPV 683
>Y162_METJA (Q57626) Hypothetical protein MJ0162
Length = 421
Score = 190 bits (482), Expect = 1e-47
Identities = 123/405 (30%), Positives = 211/405 (51%), Gaps = 27/405 (6%)
Query: 28 GAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPSTVDVLLITHFHLDHA 87
G ++G SCV + + VL DCG+ P + ++D VD ++++H HLDH
Sbjct: 8 GGCQQIGMSCVEVETQKGRVLLDCGMSPDTGEIP------KVDDKAVDAVIVSHAHLDHC 61
Query: 88 ASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDINRSMDKIEV 147
++P++ K +++ T+ T + + D + ++K Y E+DI +M+ IE
Sbjct: 62 GAIPFYKFK-----KIYCTHPTADLMFITWRDTLNLTKA------YKEEDIQHAMENIEC 110
Query: 148 IDFHQTVEVN-GIRFWCYTAGHVLGAAMFMVDIAGVRVLYTGDYSREEDRHLRAAETPQF 206
+++++ ++ I+F Y AGH+LG+A +++ G ++LYTGD + R L A+T
Sbjct: 111 LNYYEERQITENIKFKFYNAGHILGSASIYLEVDGKKILYTGDINEGVSRTLLPADTDID 170
Query: 207 SPDVCIIESTYG--VQHHQPRHTREKRFTDVIHSTISQGGRVLIPAYALGRAQELLLILD 264
DV IIESTYG + R T E++ + I TI GG+V+IP +A+GRAQE+LLI++
Sbjct: 171 EIDVLIIESTYGSPLDIKPARKTLERQLIEEISETIENGGKVIIPVFAIGRAQEILLIIN 230
Query: 265 EYWANHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRIQNAKSNPF-AFKHISALSSID 323
Y +L+++PIY L VY +Y +N +I+N N F I
Sbjct: 231 NY-IRSGKLRDVPIYTDGSLI-HATAVYMSYINWLNPKIKNMVENRINPFGEIKKADESL 288
Query: 324 IFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEGTLAKTILNEPKEVT 383
+F + P +++++ G +Q G + + D KN ++ GY EGTL + + KE+
Sbjct: 289 VF-NKEPCIIVSTSGMVQGGPVLKYLKL-LKDPKNKLILTGYQAEGTLGRELEEGAKEIQ 346
Query: 384 LMNGLSAPLHMQVHYISFSAHADSAQTSAFLEEL-NPPNIILVHG 427
P+ +V I FSAH D +++++ P I++HG
Sbjct: 347 PFKN-KIPIRGKVVKIEFSAHGDYNSLVRYIKKIPKPEKAIVMHG 390
>YC36_METJA (Q58633) Hypothetical protein MJ1236
Length = 634
Score = 189 bits (479), Expect = 3e-47
Identities = 139/425 (32%), Positives = 216/425 (50%), Gaps = 25/425 (5%)
Query: 24 VTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFD--EIDPSTVDVLLITH 81
V+ LG EVGRSC+Y+ VL DCGI+ A P+FD E +D +++TH
Sbjct: 182 VSFLGGAREVGRSCLYVQTPDTRVLIDCGINVACEDKA-FPHFDAPEFSIEDLDAVIVTH 240
Query: 82 FHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDINRS 141
HLDH +P L + + G V+ T T+ + LL DY++++K ++ Y +DI
Sbjct: 241 AHLDHCGFIPG-LFRYGYDGPVYCTRPTRDLMTLLQKDYLEIAKKEGKEVPYTSKDIKTC 299
Query: 142 MDKIEVIDFHQTVEVNG-IRFWCYTAGHVLGAAMFMVDIAG--VRVLYTGDYSREEDRHL 198
+ ID+ T +++ I+ + AGHVLG+A+ + I + YTGD E R L
Sbjct: 300 VKHTIPIDYGVTTDISPTIKLTLHNAGHVLGSAIAHLHIGEGLYNLAYTGDIKFETSRLL 359
Query: 199 RAAETPQFSPDVCIIESTYGVQHH--QPRHTREKRFTDVIHSTISQGGRVLIPAYALGRA 256
A + IIESTYG R E+ V+ T +GG+VLIP + +GRA
Sbjct: 360 EPAVCQFPRLETLIIESTYGAYDDVLPEREEAERELLRVVSETTDRGGKVLIPVFGVGRA 419
Query: 257 QELLLILDEYWANHPELQNIPIYYASPL--AKKCLTVYETY-TLSMNDRIQNAKSNPF-- 311
QEL+L+L+E + + + N P+Y + A T Y Y + M +I + NPF
Sbjct: 420 QELMLVLEEGY--NQGIFNAPVYLDGMIWEATAIHTAYPEYLSKEMRQKIFHEGDNPFLS 477
Query: 312 -AFKHISALSS-IDIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEG 369
FK + + + + P V++A+ G L G S + D+KN+ + GY EG
Sbjct: 478 EVFKRVGSTNERRKVIDSDEPCVILATSGMLTGGPSVEYLKHLAPDEKNAIIFVGYQAEG 537
Query: 370 TLAKTILNEPKEVTLM--NG--LSAPLHMQVHYI-SFSAHADSAQTSAFLEEL--NPPNI 422
TL + + + KE+ ++ NG S P+++QV+ I FS H+D Q ++ L +P I
Sbjct: 538 TLGRKVQSGWKEIPIITRNGKTKSIPINLQVYTIEGFSGHSDRKQLIKYIRRLKPSPEKI 597
Query: 423 ILVHG 427
I+VHG
Sbjct: 598 IMVHG 602
>Y047_METJA (Q60355) Hypothetical protein MJ0047
Length = 428
Score = 142 bits (358), Expect = 3e-33
Identities = 108/409 (26%), Positives = 190/409 (46%), Gaps = 26/409 (6%)
Query: 28 GAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPSTVDVLLITHFHLDHA 87
GA EVGRSC+ + +L DCG+ G P D VD + I+H HLDH+
Sbjct: 7 GAALEVGRSCIEIKTDKSKILLDCGVKLGKE--IEYPILDN-SIRDVDKVFISHAHLDHS 63
Query: 88 ASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDINRSMDKIEV 147
+LP + V T +K + K+LL D VK+++ + Y+ D+ ++
Sbjct: 64 GALPVLFHRK-MDVPVITTELSKKLIKVLLKDMVKIAETENKKIPYNNHDVKEAIRHTIP 122
Query: 148 IDFHQTVEVNGIRFWCYTAGHVLGAAMFMVDIAGVR-VLYTGDYSREEDRHLRAAETPQF 206
++++ + ++AGH+ G+A +++ + +LYTGD + R + A+
Sbjct: 123 LNYNDKKYYKDFSYELFSAGHIPGSASILLNYQNNKTILYTGDVKLRDTRLTKGADLSYT 182
Query: 207 SPDV--CIIESTYGVQHHQPRHTREKRFTDVIHSTISQGGRVLIPAYALGRAQELLLILD 264
D+ IIESTYG H R E F + I + +GG LIP +A+ RAQE+LLIL+
Sbjct: 183 KDDIDILIIESTYGNSIHPDRKAVELSFIEKIKEILFRGGVALIPVFAVDRAQEILLILN 242
Query: 265 EYWANHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRIQNAKSNPFAFKHISAL-SSID 323
+Y + P Y +A + + Y +N+ Q K A K++ + S D
Sbjct: 243 DYNIDAP-------IYLDGMAVEVTKLMLNYKHMLNESSQLEK----ALKNVKIIEKSED 291
Query: 324 IFKDV-----GPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEGTLAKTILNE 378
K + +V+ + G L G ++ + KN+ ++ GY V + + ++
Sbjct: 292 RIKAIENLSKNGGIVVTTAGMLDGGPILYYLKLFMHNPKNALLLTGYQVRDSNGRHLIET 351
Query: 379 PKEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHG 427
K + + +++V +FS HA + ++++NP +I+ HG
Sbjct: 352 GKIFIGKDEIKP--NLEVCMYNFSCHAGMDELHEIIKKVNPELLIIQHG 398
>CPSB_CAEEL (O17403) Probable cleavage and polyadenylation
specificity factor, 100 kDa subunit (CPSF 100 kDa
subunit)
Length = 843
Score = 137 bits (344), Expect = 1e-31
Identities = 98/365 (26%), Positives = 176/365 (47%), Gaps = 21/365 (5%)
Query: 28 GAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDP--STVDVLLITHFHLD 85
GA +E G C + G +L DCG + L YF+E+ P + +LI+H
Sbjct: 12 GAKDE-GPLCYLLQVDGDYILLDCG----WDERFGLQYFEELKPFIPKISAVLISHPDPL 66
Query: 86 HAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDML-YDEQDINRSMDK 144
H LPY + K V+ T + ++ + D V S + V++ Y D++ + +K
Sbjct: 67 HLGGLPYLVSKCGLTAPVYATVPVYKMGQMFIYDMV-YSHLDVEEFEHYTLDDVDTAFEK 125
Query: 145 IEVIDFHQTVEV---NGIRFWCYTAGHVLGAAMFMV-DIAGVRVLYTGDYSREEDRHLRA 200
+E + ++QTV + +G+ F AGH+LG +++ + + G ++Y D++ +++RHL
Sbjct: 126 VEQVKYNQTVVLKGDSGVHFTALPAGHMLGGSIWRICRVTGEDIVYCVDFNHKKERHLNG 185
Query: 201 AETPQFSPDVCIIESTYGVQHHQPRHT-REKRFTDVIHSTISQGGRVLIPAYALGRAQEL 259
F+ +I + + Q R R+++ I T+ Q G +I GR EL
Sbjct: 186 CSFDNFNRPHLLITGAHHISLPQMRRKDRDEQLVTKILRTVRQKGDCMIVIDTAGRVLEL 245
Query: 260 LLILDEYWANHPE-LQNIPIYYASPLAKKCLTVYETYTLSMNDRI-----QNAKSNPFAF 313
+LD+ W+N L + S +A + ++ MN+++ +A+ NPF
Sbjct: 246 AHLLDQLWSNADAGLSTYNLVMMSHVASSVVQFAKSQLEWMNEKLFKYDSSSARYNPFTL 305
Query: 314 KHISALSSI-DIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEGTLA 372
KH++ S ++ + P VV+ S ++SG SR+LF WCSD +N ++ TLA
Sbjct: 306 KHVTLCHSHQELMRVRSPKVVLCSSQDMESGFSRELFLDWCSDPRNGVILTARPASFTLA 365
Query: 373 KTILN 377
++N
Sbjct: 366 AKLVN 370
>CPSB_ARATH (Q9LKF9) Cleavage and polyadenylation specificity
factor, 100 kDa subunit (CPSF 100 kDa subunit)
Length = 739
Score = 136 bits (342), Expect = 2e-31
Identities = 106/372 (28%), Positives = 187/372 (49%), Gaps = 24/372 (6%)
Query: 24 VTPL-GAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPSTVDVLLITHF 82
VTPL G NE S + ++ G L DCG + + P ST+D +L++H
Sbjct: 7 VTPLCGVYNENPLSYL-VSIDGFNFLIDCGWNDLFDTSLLEPLSRVA--STIDAVLLSHP 63
Query: 83 HLDHAASLPYFLEKTTFKGRVFMTYATKAIYKL-LLSDYVK-VSKVSVDDM-LYDEQDIN 139
H +LPY +++ V YAT+ +++L LL+ Y + +S+ V D L+ DI+
Sbjct: 64 DTLHIGALPYAMKQLGLSAPV---YATEPVHRLGLLTMYDQFLSRKQVSDFDLFTLDDID 120
Query: 140 RSMDKIEVIDFHQTVEVNG----IRFWCYTAGHVLGAAMFMVDIAGVRVLYTGDYSREED 195
+ + + + Q ++G I + AGH+LG +++ + G V+Y DY+ ++
Sbjct: 121 SAFQNVIRLTYSQNYHLSGKGEGIVIAPHVAGHMLGGSIWRITKDGEDVIYAVDYNHRKE 180
Query: 196 RHLRAAETPQF-SPDVCIIESTYGVQHHQP-RHTREKRFTDVIHSTISQGGRVLIPAYAL 253
RHL F P V I ++ + + +Q R R+K F D I + GG VL+P
Sbjct: 181 RHLNGTVLQSFVRPAVLITDAYHALYTNQTARQQRDKEFLDTISKHLEVGGNVLLPVDTA 240
Query: 254 GRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRI----QNAKSN 309
GR ELLLIL+++W+ + PIY+ + ++ + +++ M+D I + ++ N
Sbjct: 241 GRVLELLLILEQHWSQRG--FSFPIYFLTYVSSSTIDYVKSFLEWMSDSISKSFETSRDN 298
Query: 310 PFAFKHISALSSIDIFKDV--GPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVV 367
F +H++ L + + GP VV+AS L++G +R++F W +D +N +
Sbjct: 299 AFLLRHVTLLINKTDLDNAPPGPKVVLASMASLEAGFAREIFVEWANDPRNLVLFTETGQ 358
Query: 368 EGTLAKTILNEP 379
GTLA+ + + P
Sbjct: 359 FGTLARMLQSAP 370
>CPSB_DROME (Q9V3D6) Probable cleavage and polyadenylation
specificity factor, 100 kDa subunit (CPSF 100 kDa
subunit)
Length = 756
Score = 130 bits (326), Expect = 2e-29
Identities = 95/357 (26%), Positives = 164/357 (45%), Gaps = 23/357 (6%)
Query: 37 CVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPSTVDVLLITHFHLDHAASLPYFLEK 96
C + +L DCG + ++ T+D +L++H H +LPY + K
Sbjct: 20 CYILQIDDVRILLDCGWDEKFDANFIKELKRQVH--TLDAVLLSHPDAYHLGALPYLVGK 77
Query: 97 TTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDM-LYDEQDINRSMDKIEVIDFHQTVE 155
++ T + ++ + D + +S ++ D L+ D++ + +KI + ++QTV
Sbjct: 78 LGLNCPIYATIPVFKMGQMFMYD-LYMSHFNMGDFDLFSLDDVDTAFEKITQLKYNQTVS 136
Query: 156 VN----GIRFWCYTAGHVLGAAMF-MVDIAGVRVLYTGDYSREEDRHLRAAETPQFSPDV 210
+ GI AGH++G ++ +V + ++Y D++ +++RHL E +
Sbjct: 137 LKDKGYGISITPLNAGHMIGGTIWKIVKVGEEDIVYATDFNHKKERHLSGCELDRLQRPS 196
Query: 211 CIIESTYGVQHHQPRH-TREKRFTDVIHSTISQGGRVLIPAYALGRAQELLLILDEYWAN 269
+I Y Q+ Q R R+++ I T+ G VLI GR EL +LD+ W N
Sbjct: 197 LLITDAYNAQYQQARRRARDEKLMTNILQTVRNNGNVLIAVDTAGRVLELAHMLDQLWKN 256
Query: 270 HPE--------LQNIPIYYASPLAKKCLTVYETYTLSMNDRIQNAKSNPFAFKHISALSS 321
L N Y AK + E + + + A++NPF FKHI S
Sbjct: 257 KESGLMAYSLALLNNVSYNVIEFAKSQI---EWMSDKLTKAFEGARNNPFQFKHIQLCHS 313
Query: 322 I-DIFK-DVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEGTLAKTIL 376
+ D++K GP VV+AS L+SG +R LF W S+ NS ++ GTLA ++
Sbjct: 314 LADVYKLPAGPKVVLASTPDLESGFTRDLFVQWASNANNSIILTTRTSPGTLAMELV 370
>CPSB_HUMAN (Q9P2I0) Cleavage and polyadenylation specificity
factor, 100 kDa subunit (CPSF 100 kDa subunit)
Length = 782
Score = 126 bits (316), Expect = 2e-28
Identities = 97/378 (25%), Positives = 169/378 (44%), Gaps = 30/378 (7%)
Query: 24 VTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDP-----STVDVLL 78
+T L E C + L DCG +S D ID +D +L
Sbjct: 7 LTTLSGVQEESALCYLLQVDEFRFLLDCGWDEHFS-------MDIIDSLRKHVHQIDAVL 59
Query: 79 ITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDI 138
++H H +LPY + K ++ T + ++ + D + + D L+ D+
Sbjct: 60 LSHPDPLHLGALPYAVGKLGLNCAIYATIPVYKMGQMFMYDLYQSRHNTEDFTLFTLDDV 119
Query: 139 NRSMDKIEVIDFHQTVEV----NGIRFWCYTAGHVLGAAMFMVDIAGVR-VLYTGDYSRE 193
+ + DKI+ + F Q V + +G+ AGH++G ++ + G ++Y D++ +
Sbjct: 120 DAAFDKIQQLKFSQIVNLKGKGHGLSITPLPAGHMIGGTIWKIVKDGEEEIVYAVDFNHK 179
Query: 194 EDRHLRAAETPQFSPDVCIIESTYGVQHHQPRHTR--EKRFTDVIHSTISQGGRVLIPAY 251
+ HL S +I ++ + QPR + E+ T+V+ T+ G VLI
Sbjct: 180 REIHLNGCSLEMLSRPSLLITDSFNATYVQPRRKQRDEQLLTNVLE-TLRGDGNVLIAVD 238
Query: 252 ALGRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYE---TYTLSMNDRI----Q 304
GR EL +LD+ W + +Y + L V E + M+D++ +
Sbjct: 239 TAGRVLELAQLLDQIWRTKDA--GLGVYSLALLNNVSYNVVEFSKSQVEWMSDKLMRCFE 296
Query: 305 NAKSNPFAFKHISALSSI-DIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIP 363
+ ++NPF F+H+S + D+ + P VV+AS L+ G SR LF WC D KNS ++
Sbjct: 297 DKRNNPFQFRHLSLCHGLSDLARVPSPKVVLASQPDLECGFSRDLFIQWCQDPKNSIILT 356
Query: 364 GYVVEGTLAKTILNEPKE 381
GTLA+ +++ P E
Sbjct: 357 YRTTPGTLARFLIDNPSE 374
>CPSB_BOVIN (Q10568) Cleavage and polyadenylation specificity
factor, 100 kDa subunit (CPSF 100 kDa subunit)
Length = 782
Score = 126 bits (316), Expect = 2e-28
Identities = 97/378 (25%), Positives = 169/378 (44%), Gaps = 30/378 (7%)
Query: 24 VTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDP-----STVDVLL 78
+T L E C + L DCG +S D ID +D +L
Sbjct: 7 LTTLSGVQEESALCYLLQVDEFRFLLDCGWDEHFS-------MDIIDSLRKHVHQIDAVL 59
Query: 79 ITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDI 138
++H H +LPY + K ++ T + ++ + D + + D L+ D+
Sbjct: 60 LSHPDPLHLGALPYAVGKLGLNCAIYATIPVYKMGQMFMYDLYQSRHNTEDFTLFTLDDV 119
Query: 139 NRSMDKIEVIDFHQTVEV----NGIRFWCYTAGHVLGAAMFMVDIAGVR-VLYTGDYSRE 193
+ + DKI+ + F Q V + +G+ AGH++G ++ + G ++Y D++ +
Sbjct: 120 DAAFDKIQQLKFSQIVNLKGKGHGLSITPLPAGHMIGGTIWKIVKDGEEEIVYAVDFNHK 179
Query: 194 EDRHLRAAETPQFSPDVCIIESTYGVQHHQPRHTR--EKRFTDVIHSTISQGGRVLIPAY 251
+ HL S +I ++ + QPR + E+ T+V+ T+ G VLI
Sbjct: 180 REIHLNGCSLEMLSRPSLLITDSFNATYVQPRRKQRDEQLLTNVLE-TLRGDGNVLIAVD 238
Query: 252 ALGRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYE---TYTLSMNDRI----Q 304
GR EL +LD+ W + +Y + L V E + M+D++ +
Sbjct: 239 TAGRVLELAQLLDQIWRTKDA--GLGVYSLALLNNVSYNVVEFSKSQVEWMSDKLMRCFE 296
Query: 305 NAKSNPFAFKHISALSSI-DIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIP 363
+ ++NPF F+H+S + D+ + P VV+AS L+ G SR LF WC D KNS ++
Sbjct: 297 DKRNNPFQFRHLSLCHGLSDLARVPSPKVVLASQPDLECGFSRDLFIQWCQDPKNSIILT 356
Query: 364 GYVVEGTLAKTILNEPKE 381
GTLA+ +++ P E
Sbjct: 357 YRTTPGTLARFLIDNPSE 374
>CPSB_XENLA (Q9W799) Cleavage and polyadenylation specificity
factor, 100 kDa subunit (CPSF 100 kDa subunit)
Length = 783
Score = 124 bits (311), Expect = 8e-28
Identities = 93/375 (24%), Positives = 170/375 (44%), Gaps = 24/375 (6%)
Query: 24 VTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPST--VDVLLITH 81
+T L E C + L DCG +S + D + VD +L++H
Sbjct: 7 LTTLVGAQEESAVCYLLQVDEFRFLLDCGWDENFS----MDIIDSVKKYVHQVDAVLLSH 62
Query: 82 FHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDINRS 141
H +LPY + K ++ T + ++ + D + + D L+ D++ +
Sbjct: 63 PDPLHLGALPYAVGKLGLNCAIYATIPVYKMGQMFMYDLYQSRHNTEDFSLFSLDDVDCA 122
Query: 142 MDKIEVIDFHQTVEV----NGIRFWCYTAGHVLGAAMFMVDIAGVR-VLYTGDYSREEDR 196
DKI+ + ++Q V + +G+ AGH++G ++ + G ++Y D++ + +
Sbjct: 123 FDKIQQLKYNQIVHLKGKGHGLSITPLPAGHMIGGTIWKIVKDGEEEIVYAVDFNHKREI 182
Query: 197 HLRAAETPQFSPDVCIIESTYGVQHHQPRHTR--EKRFTDVIHSTISQGGRVLIPAYALG 254
HL + +I ++ + QPR + E+ T+V+ T+ G VLI G
Sbjct: 183 HLNGCSLEMINRPSLLITDSFNATYVQPRRKQRDEQLLTNVLE-TLRGDGNVLIAVDTAG 241
Query: 255 RAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYE---TYTLSMNDRI----QNAK 307
R EL +LD+ W + +Y + L V E + M+D++ ++ +
Sbjct: 242 RVLELAQLLDQIWRTKDA--GLGVYSLALLNNVSYNVVEFSKSQVEWMSDKLMRCFEDKR 299
Query: 308 SNPFAFKHISALSSI-DIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYV 366
+NPF F+H++ D+ + P VV+AS L+ G SR+LF WC D KNS ++
Sbjct: 300 NNPFQFRHLTLCHGYSDLARVPSPKVVLASQPDLECGFSRELFIQWCQDPKNSVILTYRT 359
Query: 367 VEGTLAKTILNEPKE 381
GTLA+ +++ P E
Sbjct: 360 TPGTLARFLIDHPSE 374
Score = 33.9 bits (76), Expect = 1.5
Identities = 17/72 (23%), Positives = 34/72 (46%), Gaps = 1/72 (1%)
Query: 389 SAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGAANEMGRLKQKLMTQFADRNT 448
S + +V YI + +D + ++ P +I+VHG + L + F ++
Sbjct: 528 SMEIKARVTYIDYEGRSDGDSIKKIINQMKPRQLIIVHGPPDATQDLAEACRA-FGGKDI 586
Query: 449 KILTPKNCQSVE 460
K+ TPK ++V+
Sbjct: 587 KVYTPKLHETVD 598
>CPSB_MOUSE (O35218) Cleavage and polyadenylation specificity
factor, 100 kDa subunit (CPSF 100 kDa subunit)
Length = 782
Score = 124 bits (310), Expect = 1e-27
Identities = 96/378 (25%), Positives = 169/378 (44%), Gaps = 30/378 (7%)
Query: 24 VTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDP-----STVDVLL 78
+T L E C + L DCG +S D ID +D +L
Sbjct: 7 LTTLSGVQEESALCYLLQVDEFRFLLDCGWDEHFS-------VDIIDSLRKHVHQIDAVL 59
Query: 79 ITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDI 138
++H H +LP+ + K ++ T + ++ + D + + D L+ D+
Sbjct: 60 LSHPDPLHLGALPFAVGKLGLNCAIYATIPVYKMGQMFMYDLYQSRHNTEDFTLFTLDDV 119
Query: 139 NRSMDKIEVIDFHQTVEV----NGIRFWCYTAGHVLGAAMFMVDIAGVR-VLYTGDYSRE 193
+ + DKI+ + F Q V + +G+ AGH++G ++ + G ++Y D++ +
Sbjct: 120 DAAFDKIQQLKFSQIVNLKGKGHGLSITPLPAGHMIGGTIWKIVKDGEEEIVYAVDFNHK 179
Query: 194 EDRHLRAAETPQFSPDVCIIESTYGVQHHQPRHTR--EKRFTDVIHSTISQGGRVLIPAY 251
+ HL S +I ++ + QPR + E+ T+V+ T+ G VLI
Sbjct: 180 REIHLNGCSLEMLSRPSLLITDSFNATYVQPRRKQRDEQLLTNVLE-TLRGDGNVLIAVD 238
Query: 252 ALGRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYE---TYTLSMNDRI----Q 304
GR EL +LD+ W + +Y + L V E + M+D++ +
Sbjct: 239 TAGRVLELAQLLDQIWRTKDA--GLGVYSLALLNNVSYNVVEFSKSQVEWMSDKLMRCFE 296
Query: 305 NAKSNPFAFKHISALSSI-DIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIP 363
+ ++NPF F+H+S + D+ + P VV+AS L+ G SR LF WC D KNS ++
Sbjct: 297 DKRNNPFQFRHLSLCHGLSDLARVPSPKVVLASQPDLECGFSRDLFIQWCQDPKNSIILT 356
Query: 364 GYVVEGTLAKTILNEPKE 381
GTLA+ +++ P E
Sbjct: 357 YRTTPGTLARFLIDNPTE 374
>Y514_SYNY3 (Q55470) Hypothetical protein sll0514
Length = 554
Score = 79.7 bits (195), Expect = 2e-14
Identities = 100/427 (23%), Positives = 173/427 (40%), Gaps = 69/427 (16%)
Query: 26 PLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPSTVDVLLITHFHLD 85
P G G G C+ + +L DCG+ +AA DP TVD++ +H H D
Sbjct: 19 PYGVGPRDGGICLELHLGPYRILLDCGLEDLTPLLAA-------DPGTVDLVFCSHAHRD 71
Query: 86 HAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDINRSMDKI 145
H L F ++ A++ +LL ++ D+ +
Sbjct: 72 HGLGLWQFHQQFPH----IPILASEVTQRLLPLNWP-------DEFV---------PPFC 111
Query: 146 EVIDFHQTVEV-NGIRFWCYTAGHVLGAAMFMVDIAG----VRVLYTGDYSREEDRHLRA 200
V+ + EV G+ AGH+ GAA+ +++ RV+YTGDY HL+
Sbjct: 112 RVLPWRSPQEVLPGLTVELLPAGHLPGAALILLEYHNGDRLYRVIYTGDYCLS---HLQL 168
Query: 201 AE----TPQ--FSPDVCIIESTYGVQHHQPRHTREKRFTDVIHSTISQGGRVLIPAYALG 254
+ TP PDV I+E YG + R +EK+F I + +++G +L+P LG
Sbjct: 169 VDGLALTPLRGLKPDVLILEGHYGNRRLPHRRQQEKQFIQAIETVLAKGRNILLPVPPLG 228
Query: 255 RAQELLLILDEYWANHPEL--QNIPIYYASPLAKKCLTVYETYTLSMNDRIQN-AKSNPF 311
AQE+L +L H + + + ++ +A+ C Y+ + D ++N A+ P
Sbjct: 229 LAQEILKLL----RTHHQFTGRQVNLWAGESVARGC-DAYQGIIDHLPDNVRNFAQHQPL 283
Query: 312 -----AFKHISALSSIDIFKDV-GPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGY 365
+ H+ L+ + PS+V+ + L +W +P
Sbjct: 284 FWDDKVYPHLRPLTDDQGELSLSAPSIVITTTWPAFWPSPAALPGLWTVFMPQLLTLPSC 343
Query: 366 VVEGTLAKTILNEPKEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILV 425
+V A L E + L + L A H+D T+ + L P +++ V
Sbjct: 344 LV--NFAWQDLEEFPKYELEDYLLAD------------HSDGRNTTQLIHNLRPQHLVFV 389
Query: 426 HGAANEM 432
HG +++
Sbjct: 390 HGQPSDI 396
>YJ70_CORGL (P54122) Hypothetical UPF0036 protein Cgl1970/cg2160
Length = 718
Score = 45.1 bits (105), Expect = 6e-04
Identities = 26/80 (32%), Positives = 41/80 (50%), Gaps = 6/80 (7%)
Query: 22 LIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAA----LPYFDEIDPST--VD 75
L + LG +E+GR+ Y + ++ DCG+ SG LP F I+ VD
Sbjct: 155 LRIYALGGISEIGRNMTVFEYNNRLLIVDCGVLFPSSGEPGVDLILPDFGPIEDHLHRVD 214
Query: 76 VLLITHFHLDHAASLPYFLE 95
L++TH H DH ++P+ L+
Sbjct: 215 ALVVTHGHEDHIGAIPWLLK 234
>Y139_MYCGE (P47385) Hypothetical UPF0036 protein MG139
Length = 569
Score = 42.7 bits (99), Expect = 0.003
Identities = 36/133 (27%), Positives = 61/133 (45%), Gaps = 15/133 (11%)
Query: 28 GAGNEVGRSCVYMTYKGKTVLFDCGIH------PGYSGMAALPYFDEI--DPSTVDVLLI 79
G EVG++ + Y + ++ DCGI G +G+ +P F+ + + S V L I
Sbjct: 22 GGIQEVGKNMYGIEYDDEIIIIDCGIKFASDDLLGINGI--IPSFEHLIENQSKVKALFI 79
Query: 80 THFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDIN 139
TH H DH +PY L++ + + YA + L+L V K + + + D +
Sbjct: 80 THGHEDHIGGVPYLLKQVD----IPVIYAPRIAASLILKK-VNEHKDAKLNKIVTFDDFS 134
Query: 140 RSMDKIEVIDFHQ 152
K IDF++
Sbjct: 135 EFQTKHFKIDFYR 147
>K2C4_HUMAN (P19013) Keratin, type II cytoskeletal 4 (Cytokeratin 4)
(K4) (CK4)
Length = 534
Score = 42.4 bits (98), Expect = 0.004
Identities = 45/172 (26%), Positives = 77/172 (44%), Gaps = 31/172 (18%)
Query: 531 IQSRLKQIYESVEP---SVDEESGVPMLLVHDRVTVKHESE----KHVSLHWASDPINDM 583
+QS LK + +SVE +EE +D V +K + + V L D +ND
Sbjct: 223 LQSELKTMQDSVEDFKTKYEEEINKRTAAENDFVVLKKDVDAAYLNKVELEAKVDSLNDE 282
Query: 584 ------------------VSDSVVALVLNINR--DLPKIVAESDAT--KIEEENEKKTEK 621
VSD+ V L ++ NR DL I+AE A +I + ++ + E
Sbjct: 283 INFLKVLYDAELSQMQTHVSDTSVVLSMDNNRNLDLDSIIAEVRAQYEEIAQRSKAEAEA 342
Query: 622 VMQALLNSLFGNVKVGENGKLIINIDGNVAELNKESGEVESENEGLKERVRT 673
+ Q + L + V ++G + N +AELN+ + +E E +K++ +T
Sbjct: 343 LYQTKVQQL--QISVDQHGDNLKNTKSEIAELNRMIQRLRAEIENIKKQCQT 392
>RNZ_CLOTE (Q892B5) Ribonuclease Z (EC 3.1.26.11) (RNase Z) (tRNA 3
endonuclease)
Length = 312
Score = 42.0 bits (97), Expect = 0.005
Identities = 26/84 (30%), Positives = 35/84 (40%), Gaps = 10/84 (11%)
Query: 24 VTPLGAGN-----EVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPSTVDVLL 78
+T LG G E S + YKG+ +L DCG G + +D++
Sbjct: 4 ITLLGTGGGMPTPERNLSAAILNYKGRKILIDCG-----EGTQVSMKISKTGFKNIDIIC 58
Query: 79 ITHFHLDHAASLPYFLEKTTFKGR 102
ITH+H DH LP L GR
Sbjct: 59 ITHWHGDHIVGLPGLLATMGNSGR 82
>Y139_MYCPN (P75497) Hypothetical UPF0036 protein MG139 homolog
(A65_orf569)
Length = 569
Score = 41.6 bits (96), Expect = 0.007
Identities = 37/136 (27%), Positives = 60/136 (43%), Gaps = 21/136 (15%)
Query: 28 GAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAAL----PYFDEI--DPSTVDVLLITH 81
G EVG++ + Y + ++ DCGI + + P F+ + + + V L ITH
Sbjct: 22 GGIQEVGKNMYGIEYDDEIIIIDCGIKFASDDLLGIDGIIPSFEYLIENQAKVKALFITH 81
Query: 82 FHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDY-----VKVSKVSVDDMLYDEQ 136
H DH +PY L++ V + YA + L+L K++KV V D
Sbjct: 82 GHEDHIGGVPYLLKQVD----VPVIYAPRIAASLILKKVNEHKDAKLNKVVVYD------ 131
Query: 137 DINRSMDKIEVIDFHQ 152
D + K IDF++
Sbjct: 132 DFSNFETKHFKIDFYR 147
>ROO_DESGI (Q9F0J6) Rubredoxin-oxygen oxidoreductase (EC 1.-.-.-)
(ROO) (Rubredoxin oxidase)
Length = 402
Score = 41.6 bits (96), Expect = 0.007
Identities = 22/57 (38%), Positives = 28/57 (48%), Gaps = 1/57 (1%)
Query: 39 YMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPSTVDVLLITHFHLDHAASLPYFLE 95
Y+ KT LFD + Y G IDP +D L+I H LDHA +LP +E
Sbjct: 38 YLVEDEKTTLFDT-VKAEYKGELLCGIASVIDPKKIDYLVIQHLELDHAGALPALIE 93
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 79,049,070
Number of Sequences: 164201
Number of extensions: 3388022
Number of successful extensions: 9803
Number of sequences better than 10.0: 113
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 82
Number of HSP's that attempted gapping in prelim test: 9640
Number of HSP's gapped (non-prelim): 145
length of query: 690
length of database: 59,974,054
effective HSP length: 117
effective length of query: 573
effective length of database: 40,762,537
effective search space: 23356933701
effective search space used: 23356933701
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC134322.16


