

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.1 + phase: 0 /pseudo
(413 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
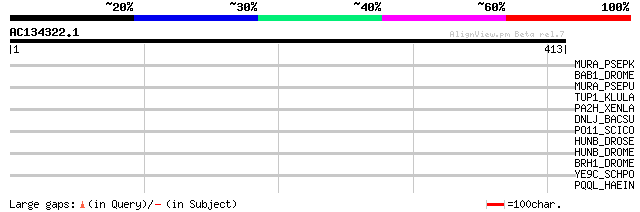
Score E
Sequences producing significant alignments: (bits) Value
MURA_PSEPK (Q88P88) UDP-N-acetylglucosamine 1-carboxyvinyltransf... 35 0.35
BAB1_DROME (Q9W0K7) Bric-a-brac protein 1 35 0.35
MURA_PSEPU (Q9Z3Z6) UDP-N-acetylglucosamine 1-carboxyvinyltransf... 33 1.0
TUP1_KLULA (P56094) Transcriptional repressor TUP1 32 2.3
PA2H_XENLA (P41485) Phospholipase A2 homolog otoconin-22 (Oc22) 32 2.3
DNLJ_BACSU (O31498) DNA ligase (EC 6.5.1.2) (Polydeoxyribonucleo... 32 3.0
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 31 5.1
HUNB_DROSE (O62538) Hunchback protein 31 5.1
HUNB_DROME (P05084) Hunchback protein 31 5.1
BRH1_DROME (Q24255) Homeobox protein B-H1 (Homeobox BarH1 protein) 31 6.6
YE9C_SCHPO (O13773) Hypothetical J-domain protein C17A5.12 in ch... 30 8.7
PQQL_HAEIN (P45181) Probable zinc protease pqqL (EC 3.4.99.-) 30 8.7
>MURA_PSEPK (Q88P88) UDP-N-acetylglucosamine
1-carboxyvinyltransferase (EC 2.5.1.7) (Enoylpyruvate
transferase) (UDP-N-acetylglucosamine enolpyruvyl
transferase) (EPT)
Length = 421
Score = 35.0 bits (79), Expect = 0.35
Identities = 25/84 (29%), Positives = 40/84 (46%), Gaps = 11/84 (13%)
Query: 7 RFIVRGGARW------SIGSGATIPILDTPWLANGESINGNIPGGSYIHDFTVQNLLSQH 60
+ I+ GGAR S A +PIL LA+G GN+P ++HD T ++
Sbjct: 3 KLIITGGARLDGEIRISGAKNAALPILAATLLADGPVTVGNLP---HLHDIT--TMIELF 57
Query: 61 GKSWNEPLVQQMFSNDIAEAILHT 84
G+ EP++ + S +I + T
Sbjct: 58 GRMGIEPVIDEKLSVEIDPRTIKT 81
>BAB1_DROME (Q9W0K7) Bric-a-brac protein 1
Length = 977
Score = 35.0 bits (79), Expect = 0.35
Identities = 23/67 (34%), Positives = 30/67 (44%), Gaps = 8/67 (11%)
Query: 294 PLQLHAQHQ-QSAVPLQAHAHHHNRPAGH-------GNLKWTPPSSGRFKCNVDAAFSAQ 345
P Q AQ Q Q +PL H HHH PA H + +P RF AA +A
Sbjct: 384 PQQQQAQQQGQLPLPLPLHPHHHASPAPHPSQTAGSAHHPASPAGDSRFPLGPAAAMAAA 443
Query: 346 FQRTGIG 352
+ +G+G
Sbjct: 444 RELSGLG 450
>MURA_PSEPU (Q9Z3Z6) UDP-N-acetylglucosamine
1-carboxyvinyltransferase (EC 2.5.1.7) (Enoylpyruvate
transferase) (UDP-N-acetylglucosamine enolpyruvyl
transferase) (EPT)
Length = 419
Score = 33.5 bits (75), Expect = 1.0
Identities = 24/84 (28%), Positives = 40/84 (47%), Gaps = 11/84 (13%)
Query: 7 RFIVRGGA------RWSIGSGATIPILDTPWLANGESINGNIPGGSYIHDFTVQNLLSQH 60
+ I+ GGA R S A +PIL LA+G GN+P ++HD T ++
Sbjct: 3 KLIITGGACLDGEIRISGAKNAALPILAATLLADGPVTVGNLP---HLHDIT--TMIELF 57
Query: 61 GKSWNEPLVQQMFSNDIAEAILHT 84
G+ EP++ + + +I + T
Sbjct: 58 GRMGIEPVIDEKLAVEIDPRTIKT 81
>TUP1_KLULA (P56094) Transcriptional repressor TUP1
Length = 682
Score = 32.3 bits (72), Expect = 2.3
Identities = 24/61 (39%), Positives = 30/61 (48%), Gaps = 3/61 (4%)
Query: 271 EEAITLQPATFLQEDVQQQ*PAVPLQLHAQHQQSAVPLQAHAHHHNRPAGHGNLKWTPPS 330
+ A L P Q+ QQQ P Q Q QQS +P+ A +PAG GNL T P+
Sbjct: 162 QAAANLAPVIQQQQQPQQQLPPQQQQQQQQQQQSNIPVTTAA--PVQPAG-GNLDQTVPN 218
Query: 331 S 331
S
Sbjct: 219 S 219
>PA2H_XENLA (P41485) Phospholipase A2 homolog otoconin-22 (Oc22)
Length = 127
Score = 32.3 bits (72), Expect = 2.3
Identities = 14/27 (51%), Positives = 16/27 (58%)
Query: 91 TNDCLVWKAERNGCYSVRSAYRLYVEE 117
+ DC KAE +GC V YR YVEE
Sbjct: 46 SQDCCYNKAEMSGCNPVTQTYRFYVEE 72
>DNLJ_BACSU (O31498) DNA ligase (EC 6.5.1.2)
(Polydeoxyribonucleotide synthase [NAD+])
Length = 668
Score = 32.0 bits (71), Expect = 3.0
Identities = 29/106 (27%), Positives = 47/106 (43%), Gaps = 18/106 (16%)
Query: 161 QDKGVSCPTNCESCGAD------HEDLNHLLFECPFSIHVWNSAGIWHDVQHAAIHSDSA 214
++K S PT C CG++ L + ECP I G+ H V A++ D
Sbjct: 394 EEKEFSMPTECPECGSELVRIEGEVALRCINPECPAQIR----EGLIHFVSRNAMNIDGL 449
Query: 215 ANTIFSIL--QNLSRNIKQRFAATCWSLWKHRNLKIWENVDENSVQ 258
+ + L +NL RN+ A + L K R +++ E + E S +
Sbjct: 450 GERVITQLFEENLVRNV-----ADLYKLTKERVIQL-ERMGEKSTE 489
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 31.2 bits (69), Expect = 5.1
Identities = 15/30 (50%), Positives = 19/30 (63%), Gaps = 1/30 (3%)
Query: 160 LQDKGVSCPTNCESCGADHEDLNHLLFECP 189
L +G+S C CGA +ED+ HLL ECP
Sbjct: 919 LHKRGLSNTPVC-MCGAPNEDVKHLLGECP 947
>HUNB_DROSE (O62538) Hunchback protein
Length = 757
Score = 31.2 bits (69), Expect = 5.1
Identities = 15/47 (31%), Positives = 27/47 (56%)
Query: 271 EEAITLQPATFLQEDVQQQ*PAVPLQLHAQHQQSAVPLQAHAHHHNR 317
+ A+ L T ++ED QQQ P PL ++ + ++ A PL + ++ R
Sbjct: 538 DSAMDLSQGTPVKEDEQQQQPQQPLAMNLKVEEEATPLMSSSNASRR 584
>HUNB_DROME (P05084) Hunchback protein
Length = 758
Score = 31.2 bits (69), Expect = 5.1
Identities = 15/47 (31%), Positives = 27/47 (56%)
Query: 271 EEAITLQPATFLQEDVQQQ*PAVPLQLHAQHQQSAVPLQAHAHHHNR 317
+ A+ L T ++ED QQQ P PL ++ + ++ A PL + ++ R
Sbjct: 539 DSAMDLSQGTPVKEDEQQQQPQQPLAMNLKVEEEATPLMSSSNASRR 585
>BRH1_DROME (Q24255) Homeobox protein B-H1 (Homeobox BarH1 protein)
Length = 544
Score = 30.8 bits (68), Expect = 6.6
Identities = 20/56 (35%), Positives = 26/56 (45%), Gaps = 3/56 (5%)
Query: 293 VPLQLHAQHQQSAVPLQAHAHHHNRPAGHGNLKWTPPSSGRFKCNVDAAFSAQFQR 348
+P LH H Q P H H H HG L PP++G NV A ++A Q+
Sbjct: 143 LPSALH--HPQPHPPTHPHTHPHALMHPHGKLGHFPPTAGGNGLNV-AQYAAAMQQ 195
>YE9C_SCHPO (O13773) Hypothetical J-domain protein C17A5.12 in
chromosome I
Length = 697
Score = 30.4 bits (67), Expect = 8.7
Identities = 18/59 (30%), Positives = 28/59 (46%)
Query: 196 NSAGIWHDVQHAAIHSDSAANTIFSILQNLSRNIKQRFAATCWSLWKHRNLKIWENVDE 254
N+ G + + + SDS+A T FS Q LS +K + LW K+ + V+E
Sbjct: 223 NAKGQYREGEAYEAFSDSSAKTQFSDFQALSNQLKSQLFEKANDLWNIGRKKLRDAVEE 281
>PQQL_HAEIN (P45181) Probable zinc protease pqqL (EC 3.4.99.-)
Length = 926
Score = 30.4 bits (67), Expect = 8.7
Identities = 23/82 (28%), Positives = 36/82 (43%), Gaps = 6/82 (7%)
Query: 199 GIWHDVQHAAIHSDSA--ANTIFSILQNLSRNIKQRFAATCWSLWKHRNLKIWENVDENS 256
GI H V+H A + N I + L+ L +FA + N N+D N+
Sbjct: 76 GIAHLVEHMAFNGSKKYPENQIINALEKLG----MKFARDINAFTDFENTVYTLNLDSNN 131
Query: 257 VQVVDRARNLIADWEEAITLQP 278
Q ++ A ++I +W IT P
Sbjct: 132 QQKLELAFDVINEWMNNITFLP 153
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.138 0.461
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,987,032
Number of Sequences: 164201
Number of extensions: 2148242
Number of successful extensions: 6242
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 6228
Number of HSP's gapped (non-prelim): 19
length of query: 413
length of database: 59,974,054
effective HSP length: 113
effective length of query: 300
effective length of database: 41,419,341
effective search space: 12425802300
effective search space used: 12425802300
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC134322.1


