

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134049.3 - phase: 0
(1113 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
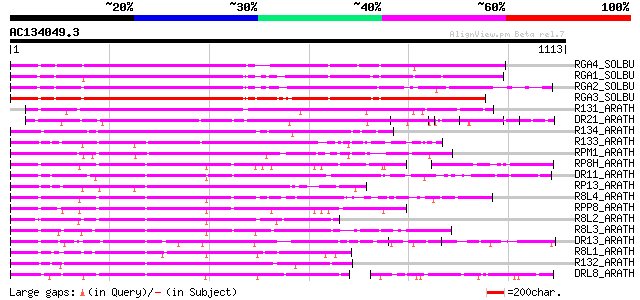
Score E
Sequences producing significant alignments: (bits) Value
RGA4_SOLBU (Q7XA39) Putative disease resistance protein RGA4 (RG... 705 0.0
RGA1_SOLBU (Q7XA42) Putative disease resistance protein RGA1 (RG... 691 0.0
RGA2_SOLBU (Q7XBQ9) Disease resistance protein RGA2 (RGA2-blb) (... 689 0.0
RGA3_SOLBU (Q7XA40) Putative disease resistance protein RGA3 (RG... 688 0.0
R131_ARATH (Q9LRR4) Putative disease resistance RPP13-like prote... 465 e-130
DR21_ARATH (Q9LRR5) Putative disease resistance protein At3g14460 388 e-107
R134_ARATH (Q38834) Putative disease resistance RPP13-like prote... 243 3e-63
R133_ARATH (Q9STE7) Putative disease resistance RPP13-like prote... 232 5e-60
RPM1_ARATH (Q39214) Disease resistance protein RPM1 (Resistance ... 231 1e-59
RP8H_ARATH (P59584) Disease resistance protein RPH8A (RPP8 homol... 223 2e-57
DR11_ARATH (Q94HW3) Probable disease resistance protein RDL6/RF9 219 3e-56
RP13_ARATH (Q9M667) Disease resistance protein RPP13 (Resistance... 219 4e-56
R8L4_ARATH (Q9FJK8) Probable disease resistance RPP8-like protein 4 214 1e-54
RPP8_ARATH (Q8W4J9) Disease resistance protein RPP8 (Resistance ... 214 1e-54
R8L2_ARATH (Q9MAG6) Probable disease resistance RPP8-like protein 2 211 9e-54
R8L3_ARATH (Q9FJB5) Probable disease resistance RPP8-like protein 3 207 1e-52
DR13_ARATH (Q9XIF0) Putative disease resistance protein At1g59780 204 1e-51
R8L1_ARATH (O04093) Putative disease resistance protein RPP8-lik... 199 5e-50
R132_ARATH (Q9STE5) Putative disease resistance RPP13-like prote... 194 1e-48
DRL8_ARATH (Q8W3K3) Putative disease resistance protein At1g58400 192 3e-48
>RGA4_SOLBU (Q7XA39) Putative disease resistance protein RGA4
(RGA4-blb)
Length = 988
Score = 705 bits (1820), Expect = 0.0
Identities = 427/1007 (42%), Positives = 598/1007 (58%), Gaps = 56/1007 (5%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA L+++L +L+ I ++ L GF++E +L+S+ +TI+A L+DA+EKQ D I
Sbjct: 1 MAEAFLQVLLENLTSFIGDKLVLIFGFEKECEKLSSVFSTIQAVLQDAQEKQLKDKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIA-FRYKLAKKMKRIGV 119
++WL KL AAY +DDI+ EC EA+ E S+ G H I FR+K+ ++MK I
Sbjct: 59 --ENWLQKLNSAAYEVDDILGECKNEAIRFEQ--SRLGFYHPGIINFRHKIGRRMKEIME 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD I+ E+ KFH E + ER R+T ++T+P VYGR++++D+IV L+ + +
Sbjct: 115 KLDAISEERRKFHFLEKITERQAAAAT-RETGFVLTEPKVYGRDKEEDEIVKILINNVNV 173
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
E+L V+PI+G+GGLGKTTLAQ++FN +++ HF KIWVCVS+DF KR+ K II
Sbjct: 174 AEELPVFPIIGMGGLGKTTLAQMIFNDERVTKHFNPKIWVCVSDDFDEKRLIKTIIGNIE 233
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
+ S DL Q+KLQ+LL KRYLLVLDDVWND E W +L++VL G +GASIL TT
Sbjct: 234 RSSPHVEDLASFQKKLQELLNGKRYLLVLDDVWNDDLEKWAKLRAVLTVGARGASILATT 293
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL KV IMGT+ + LS LS D LF QRAFG + LV +GKEI+KKCGG PL
Sbjct: 294 RLEKVGSIMGTLQPYHLSNLSPHDSLLLFMQRAFGQQKEANPNLVAIGKEIVKKCGGVPL 353
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKREE EW +V+++++W+L Q E+ ++PALRLSY HLP+ LRQCF++CA+
Sbjct: 354 AAKTLGGLLRFKREESEWEHVRDNEIWSLPQDESSILPALRLSYHHLPLDLRQCFAYCAV 413
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD + K+ LI LW A+GF+ S LE +D+GNEVWNELY RSFF+ E T
Sbjct: 414 FPKDTKMIKENLITLWMAHGFLLSKGNLELEDVGNEVWNELYLRSFFQEIEAKSGN--TY 471
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FK+HDL+HDLA S+ ++ A +I+ +VK K
Sbjct: 472 FKIHDLIHDLATSLF---------------------------SASASCGNIREINVKDYK 504
Query: 539 TYMEFNFDVYEAGQLSPQVLNCY-SLRVL-LSH-RLNNLSSSIGRLKYLRYLDISEGRFK 595
+ F SP +L + SLRVL LS+ +L L SSIG L +LRYLD+S F+
Sbjct: 505 HTVSIGFAAV-VSSYSPSLLKKFVSLRVLNLSYSKLEQLPSSIGDLLHLRYLDLSCNNFR 563
Query: 596 NLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSL 655
+LP LCKL NL+ L + C SL LP ++L L++L + C LTS P +IG LT L
Sbjct: 564 SLPERLCKLQNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDGC-PLTSTPPRIGLLTCL 622
Query: 656 NTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSW 714
TL +IVG ++G+ L EL LNL G + I +LER+K+ TDA +AN+S K L L +SW
Sbjct: 623 KTLGFFIVGSKKGYQLGELKNLNLCGSISITHLERVKNDTDA-EANLSAKANLQSLSMSW 681
Query: 715 ERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCK 774
+ + ++ + ++LEAL+P+ Y + + G FP WI+ L + S+ + CK
Sbjct: 682 DNDGPNRYESKEVKVLEALKPHPNLKY-LEIIAFGGFRFPSWINHSVLEKVISVRIKSCK 740
Query: 775 SCLNLPELWKLPSLKYLKLSN-MIHVIYLFHESYDG-----EGLMALKTLFLEKLPNLIG 828
+CL LP +LP L+ L+L N V Y+ + +LK L + +L G
Sbjct: 741 NCLCLPPFGELPCLENLELQNGSAEVEYVEEDDVHSRFSTRRSFPSLKKLRIWFFRSLKG 800
Query: 829 LSREE-RVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLH 887
L +EE FP L+ + I CP L P L S+ L + G N + SSI L +L SL
Sbjct: 801 LMKEEGEEKFPMLEEMAILYCP-LFVFPTLSSVKKLEVHGNTNTRGLSSISNLSTLTSLR 859
Query: 888 FSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEEL 947
N P+ + +L + L+ L F LK LPT + ++AL++L I C ++E
Sbjct: 860 IGANYRATSLPEEMFTSLTN-LEFLSFFDFKNLKDLPTSLTSLNALKRLQIESCDSLESF 918
Query: 948 PNEVMQRLHSLKELDIVGCDKLK-LSSDFQYLTCLETLAIGSCSEVE 993
P + ++ L SL +L + C LK L Q+LT L L + C EVE
Sbjct: 919 PEQGLEGLTSLTQLFVKYCKMLKCLPEGLQHLTALTNLGVSGCPEVE 965
Score = 40.8 bits (94), Expect = 0.021
Identities = 39/160 (24%), Positives = 71/160 (44%), Gaps = 16/160 (10%)
Query: 874 PSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKL-----KMLPTEMI 928
PS + K SL L+ S ++ L L S + L R+ L + LP +
Sbjct: 520 PSLLKKFVSLRVLNLSYSK---------LEQLPSSIGDLLHLRYLDLSCNNFRSLPERLC 570
Query: 929 HIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGS 988
+ LQ L +++C ++ LP + +L SL+ L + GC LTCL+TL
Sbjct: 571 KLQNLQTLDVHNCYSLNCLPKQT-SKLSSLRHLVVDGCPLTSTPPRIGLLTCLKTLGFFI 629
Query: 989 CSEVEGFH-EALQHMTTLKSLTLSDLPNLEYLPECIGNLT 1027
+G+ L+++ S++++ L ++ + NL+
Sbjct: 630 VGSKKGYQLGELKNLNLCGSISITHLERVKNDTDAEANLS 669
Score = 39.7 bits (91), Expect = 0.046
Identities = 26/83 (31%), Positives = 44/83 (52%), Gaps = 3/83 (3%)
Query: 983 TLAIGSCSEVEGFHEAL-QHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLA 1041
T++IG + V + +L + +L+ L LS LE LP IG+L L +++ SC
Sbjct: 506 TVSIGFAAVVSSYSPSLLKKFVSLRVLNLS-YSKLEQLPSSIGDLLHLRYLDL-SCNNFR 563
Query: 1042 CLPTSIQQISGLEILSIHDCSKL 1064
LP + ++ L+ L +H+C L
Sbjct: 564 SLPERLCKLQNLQTLDVHNCYSL 586
Score = 37.7 bits (86), Expect = 0.17
Identities = 32/121 (26%), Positives = 52/121 (42%), Gaps = 5/121 (4%)
Query: 941 CRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQ 1000
C NI E+ + + S+ +V L F L L S S++E ++
Sbjct: 492 CGNIREINVKDYKHTVSIGFAAVVSSYSPSLLKKFVSLRVLNL----SYSKLEQLPSSIG 547
Query: 1001 HMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHD 1060
+ L+ L LS N LPE + L L +++++C L CLP ++S L L +
Sbjct: 548 DLLHLRYLDLS-CNNFRSLPERLCKLQNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDG 606
Query: 1061 C 1061
C
Sbjct: 607 C 607
>RGA1_SOLBU (Q7XA42) Putative disease resistance protein RGA1
(RGA3-blb)
Length = 1025
Score = 691 bits (1782), Expect = 0.0
Identities = 438/1042 (42%), Positives = 601/1042 (57%), Gaps = 98/1042 (9%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA +++VL +L+ ++ E+ L GF EF RL+S+ +TI+A LEDA+EKQ +D +
Sbjct: 1 MAEAFIQVVLDNLTSFLKGELVLLFGFQDEFQRLSSMFSTIQAVLEDAQEKQLND----K 56
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
+++WL KL A Y +DDI+DE T+A + S+ G H K I FR+K+ K+M ++
Sbjct: 57 PLENWLQKLNAATYEVDDILDEYKTKATR--FLQSEYGRYHPKVIPFRHKVGKRMDQVMK 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPD--------------------------------- 146
L+ IA E+ KFHL E + ER +
Sbjct: 115 KLNAIAEERKKFHLQEKIIERQAATRETVSMFSTCSSSHPYLFIVQNPSLFNYFLPTPKE 174
Query: 147 --------WRQTT--SIVTQPLVYGRNEDKDKIVDFLVGDASEQEDLSVYPIVGLGGLGK 196
W S++T+P VYGR+++KD+IV L+ AS+ + LSV PI+G+GGLGK
Sbjct: 175 LECNNILIWTLLAPGSVLTEPQVYGRDKEKDEIVKILINTASDAQKLSVLPILGMGGLGK 234
Query: 197 TTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSCEDLDLELLQRKLQ 256
TTL+Q+VFN ++ F KIW+C+S+DF KR+ KAI+E KS D+DL LQ+KLQ
Sbjct: 235 TTLSQMVFNDQRVTERFYPKIWICISDDFNEKRLIKAIVESIEGKSLSDMDLAPLQKKLQ 294
Query: 257 DLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIMGTIPHHEL 316
+LL KRY LVLDDVWN+ Q W L++VL G GA +L TTRL KV IMGT+ +EL
Sbjct: 295 ELLNGKRYFLVLDDVWNEDQHKWANLRAVLKVGASGAFVLTTTRLEKVGSIMGTLQPYEL 354
Query: 317 SRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKE 376
S LS EDCW LF QRAFG E L+ +GKEI+KKCGG PLAA LG +LRFKREE+E
Sbjct: 355 SNLSPEDCWFLFMQRAFGHQEEINPNLMAIGKEIVKKCGGVPLAAKTLGGILRFKREERE 414
Query: 377 WLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWT 435
W +V++S +WNL Q E+ ++PALRLSY HLP+ LRQCF +CA+FPKD ++K+ LI W
Sbjct: 415 WEHVRDSPIWNLPQDESSILPALRLSYHHLPLDLRQCFVYCAVFPKDTKMAKENLIAFWM 474
Query: 436 ANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQD 495
A+GF+ S LE +D+GNEVWNELY RSFF+ E V G+ T FKMHDL+HDLA S+
Sbjct: 475 AHGFLLSKGNLELEDVGNEVWNELYLRSFFQEIE-VESGK-TYFKMHDLIHDLATSLF-- 530
Query: 496 VCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSP 555
S T S R + N N SI V S SP
Sbjct: 531 --------SANTSSSNIREI---NANYDGYMMSIGFAEVVS---------------SYSP 564
Query: 556 QVLNCY-SLRV--LLSHRLNNLSSSIGRLKYLRYLDISEG-RFKNLPNSLCKLCNLEVLK 611
+L + SLRV L + LN L SSIG L +LRYLD+S R +NLP LCKL NL+ L
Sbjct: 565 SLLQKFVSLRVLNLRNSNLNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCKLQNLQTLD 624
Query: 612 LDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLL 671
L C SL LP ++L L+NL L C SLTS P +IG LT L +LS +++G+ +G L
Sbjct: 625 LHYCDSLSCLPKQTSKLGSLRNLLLDGC-SLTSTPPRIGLLTCLKSLSCFVIGKRKGHQL 683
Query: 672 EELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSWERNEVSQLQENVEQIL 730
EL LNL G + I L+R+K TDAK+AN+S K L+ L LSW+ + + ++L
Sbjct: 684 GELKNLNLYGSISITKLDRVKKDTDAKEANLSAKANLHSLCLSWDLDGKHRYD---SEVL 740
Query: 731 EALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLNLPELWKLPSLKY 790
EAL+P++ Y + G+ G P W++ L ++ S+ + C++C LP +LP L+
Sbjct: 741 EALKPHSNLKY-LEINGFGGIRLPDWMNQSVLKNVVSIRIRGCENCSCLPPFGELPCLES 799
Query: 791 LKL-SNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSR-EERVMFPRLKALEITEC 848
L+L + V Y+ + G +L+ L + NL GL + E FP L+ + C
Sbjct: 800 LELHTGSADVEYVEDNVHPGR-FPSLRKLVIWDFSNLKGLLKMEGEKQFPVLEEMTFYWC 858
Query: 849 PNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASP 908
P + +P L S+ L + + + SI L +L SL SDN E P+ + ++LA+
Sbjct: 859 P-MFVIPTLSSVKTLKVI-VTDATVLRSISNLRALTSLDISDNVEATSLPEEMFKSLAN- 915
Query: 909 LKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDK 968
LK L LK LPT + ++AL+ L C +E LP E ++ L SL EL + C
Sbjct: 916 LKYLKISFFRNLKELPTSLASLNALKSLKFEFCDALESLPEEGVKGLTSLTELSVSNCMM 975
Query: 969 LK-LSSDFQYLTCLETLAIGSC 989
LK L Q+LT L TL I C
Sbjct: 976 LKCLPEGLQHLTALTTLTITQC 997
Score = 35.4 bits (80), Expect = 0.86
Identities = 23/82 (28%), Positives = 41/82 (49%), Gaps = 2/82 (2%)
Query: 984 LAIGSCSEVEGFHEAL-QHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLAC 1042
++IG V + +L Q +L+ L L + NL LP IG+L L +++ ++
Sbjct: 551 MSIGFAEVVSSYSPSLLQKFVSLRVLNLRN-SNLNQLPSSIGDLVHLRYLDLSGNFRIRN 609
Query: 1043 LPTSIQQISGLEILSIHDCSKL 1064
LP + ++ L+ L +H C L
Sbjct: 610 LPKRLCKLQNLQTLDLHYCDSL 631
Score = 33.1 bits (74), Expect = 4.3
Identities = 18/73 (24%), Positives = 34/73 (45%)
Query: 990 SEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQ 1049
S + ++ + L+ L LS + LP+ + L L ++++ C L+CLP +
Sbjct: 581 SNLNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCKLQNLQTLDLHYCDSLSCLPKQTSK 640
Query: 1050 ISGLEILSIHDCS 1062
+ L L + CS
Sbjct: 641 LGSLRNLLLDGCS 653
>RGA2_SOLBU (Q7XBQ9) Disease resistance protein RGA2 (RGA2-blb)
(Blight resistance protein RPI)
Length = 970
Score = 689 bits (1777), Expect = 0.0
Identities = 441/1099 (40%), Positives = 618/1099 (56%), Gaps = 143/1099 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA ++++L +L+ ++ E+ L GF EF RL+S+ +TI+A LEDA+EKQ ++ +
Sbjct: 1 MAEAFIQVLLDNLTSFLKGELVLLFGFQDEFQRLSSMFSTIQAVLEDAQEKQLNN----K 56
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
+++WL KL A Y +DDI+DE T+A + S+ G H K I FR+K+ K+M ++
Sbjct: 57 PLENWLQKLNAATYEVDDILDEYKTKATR--FSQSEYGRYHPKVIPFRHKVGKRMDQVMK 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
L IA E+ FHL E + ER V R+T S++T+P VYGR+++KD+IV L+ + S+
Sbjct: 115 KLKAIAEERKNFHLHEKIVERQAVR---RETGSVLTEPQVYGRDKEKDEIVKILINNVSD 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+ LSV PI+G+GGLGKTTLAQ+VFN ++ HF KIW+CVSEDF KR+ KAI+E
Sbjct: 172 AQHLSVLPILGMGGLGKTTLAQMVFNDQRVTEHFHSKIWICVSEDFDEKRLIKAIVESIE 231
Query: 240 KKSC-EDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVT 298
+ ++DL LQ+KLQ+LL KRYLLVLDDVWN+ Q+ W L++VL G GAS+L T
Sbjct: 232 GRPLLGEMDLAPLQKKLQELLNGKRYLLVLDDVWNEDQQKWANLRAVLKVGASGASVLTT 291
Query: 299 TRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFP 358
TRL KV IMGT+ +ELS LS EDCW LF QRAFG E LV +GKEI+KK GG P
Sbjct: 292 TRLEKVGSIMGTLQPYELSNLSQEDCWLLFMQRAFGHQEEINPNLVAIGKEIVKKSGGVP 351
Query: 359 LAAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCA 417
LAA LG +L FKREE+ W +V++S +WNL Q E+ ++PALRLSY LP+ L+QCF++CA
Sbjct: 352 LAAKTLGGILCFKREERAWEHVRDSPIWNLPQDESSILPALRLSYHQLPLDLKQCFAYCA 411
Query: 418 LFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQIT 477
+FPKD + K+ LI LW A+GF+ S +E +D+G+EVW ELY RSFF+ E V G+ T
Sbjct: 412 VFPKDAKMEKEKLISLWMAHGFLLSKGNMELEDVGDEVWKELYLRSFFQEIE-VKDGK-T 469
Query: 478 IFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSL 537
FKMHDL+HDLA S+ S T S R + N++S+ SI V
Sbjct: 470 YFKMHDLIHDLATSLF----------SANTSSSNIREI---NKHSYTHMMSIGFAEVVFF 516
Query: 538 KTYMEFNFDVYEAGQLSPQVLNCYSLRVLL--SHRLNNLSSSIGRLKYLRYLDISEGRFK 595
T P + SLRVL N L SSIG L +LRYL++ +
Sbjct: 517 YTL--------------PPLEKFISLRVLNLGDSTFNKLPSSIGDLVHLRYLNLYGSGMR 562
Query: 596 NLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSL 655
+LP LCKL NL+ L L C L LP ++L L+NL L SLT +P +IG LT L
Sbjct: 563 SLPKQLCKLQNLQTLDLQYCTKLCCLPKETSKLGSLRNLLLDGSQSLTCMPPRIGSLTCL 622
Query: 656 NTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSW 714
TL +++VG ++G+ L ELG LNL G + I +LER+K+ DAK+AN+S K L+ L +SW
Sbjct: 623 KTLGQFVVGRKKGYQLGELGNLNLYGSIKISHLERVKNDKDAKEANLSAKGNLHSLSMSW 682
Query: 715 ERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCK 774
+ ++LEAL+P++ L S + G+ G + P+W++ L ++ S+ + + +
Sbjct: 683 NNFGPHIYESEEVKVLEALKPHS-NLTSLKIYGFRGIHLPEWMNHSVLKNIVSILISNFR 741
Query: 775 SCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREER 834
+C LP LP L+ L+ L S D E ++E++ + R
Sbjct: 742 NCSCLPPFGDLPCLESLE---------LHWGSADVE--------YVEEVDIDVHSGFPTR 784
Query: 835 VMFPRLKALEITECPNLLGL------PCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHF 888
+ FP L+ L+I + +L GL P L ++ I L S+ L +L SL
Sbjct: 785 IRFPSLRKLDIWDFGSLKGLLKKEGEEQFPVLEEMIIHECPFLTLSSN---LRALTSLRI 841
Query: 889 SDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELP 948
N+ FP+ + +NLA+ LK L R + LK LPT + ++AL+ L I C +E LP
Sbjct: 842 CYNKVATSFPEEMFKNLAN-LKYLTISRCNNLKELPTSLASLNALKSLKIQLCCALESLP 900
Query: 949 NEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSL 1008
E ++ L SL EL + C+ LK CL E LQH+TTL SL
Sbjct: 901 EEGLEGLSSLTELFVEHCNMLK---------CLP--------------EGLQHLTTLTSL 937
Query: 1009 TLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRC 1068
I CP+L KRC
Sbjct: 938 ------------------------KIRGCPQLI------------------------KRC 949
Query: 1069 QKEIGEDWPKIVHVQYIEI 1087
+K IGEDW KI H+ + I
Sbjct: 950 EKGIGEDWHKISHIPNVNI 968
>RGA3_SOLBU (Q7XA40) Putative disease resistance protein RGA3
(RGA1-blb) (Blight resistance protein B149)
Length = 947
Score = 688 bits (1775), Expect = 0.0
Identities = 417/959 (43%), Positives = 587/959 (60%), Gaps = 38/959 (3%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA L+++L +L+ I+ E+ L GF++EF +L+S+ + I+A LEDA+EKQ I
Sbjct: 1 MAEAFLQVLLDNLTFFIQGELGLVFGFEKEFKKLSSMFSMIQAVLEDAQEKQLKYKAI-- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-KHIAFRYKLAKKMKRIGV 119
K+WL KL AAY +DDI+D+C TEA +K + G H + I F YK+ K+MK +
Sbjct: 59 --KNWLQKLNVAAYEVDDILDDCKTEAAR--FKQAVLGRYHPRTITFCYKVGKRMKEMME 114
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
LD IA E+ FHL E + ER RQT ++T+P VYGR +++D+IV L+ + S
Sbjct: 115 KLDAIAEERRNFHLDERIIERQAAR---RQTGFVLTEPKVYGREKEEDEIVKILINNVSY 171
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
E++ V PI+G+GGLGKTTLAQ+VFN +I HF LKIWVCVS+DF KR+ KAI+E
Sbjct: 172 SEEVPVLPILGMGGLGKTTLAQMVFNDQRITEHFNLKIWVCVSDDFDEKRLIKAIVESIE 231
Query: 240 KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
KS D+DL LQ+KLQ+LL KRY LVLDDVWN+ QE W L++VL G GASIL+TT
Sbjct: 232 GKSLGDMDLAPLQKKLQELLNGKRYFLVLDDVWNEDQEKWDNLRAVLKIGASGASILITT 291
Query: 300 RLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPL 359
RL K+ IMGT+ ++LS LS EDCW LFKQRAF +L+ +GKEI+KKCGG PL
Sbjct: 292 RLEKIGSIMGTLQLYQLSNLSQEDCWLLFKQRAFCHQTETSPKLMEIGKEIVKKCGGVPL 351
Query: 360 AAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCAL 418
AA LG LLRFKREE EW +V++S++WNL Q E V+PALRLSY HLP+ LRQCF++CA+
Sbjct: 352 AAKTLGGLLRFKREESEWEHVRDSEIWNLPQDENSVLPALRLSYHHLPLDLRQCFAYCAV 411
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITI 478
FPKD I K+ LI LW A+ F+ S +E +D+GNEVWNELY RSFF+ E V G+ T
Sbjct: 412 FPKDTKIEKEYLIALWMAHSFLLSKGNMELEDVGNEVWNELYLRSFFQEIE-VKSGK-TY 469
Query: 479 FKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK 538
FKMHDL+HDLA S+ S+R ++ + +++ ++ + SI V S
Sbjct: 470 FKMHDLIHDLATSM---FSASASSRSIRQINVKDDEDMMFIVTNYKDMMSIGFSEVVS-- 524
Query: 539 TYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLSSSIGRLKYLRYLDISEGRFKNLP 598
+Y F + +S +VLN L + L SS+G L +LRYLD+S + +LP
Sbjct: 525 SYSPSLFKRF----VSLRVLN------LSNSEFEQLPSSVGDLVHLRYLDLSGNKICSLP 574
Query: 599 NSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTL 658
LCKL NL+ L L C SL LP ++L L+NL L C LTS+P +IG LT L TL
Sbjct: 575 KRLCKLQNLQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHC-PLTSMPPRIGLLTCLKTL 633
Query: 659 SKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMSRK-KLNQLWLSWERN 717
++VGE +G+ L EL LNL+G + I +LER+K+ +AK+AN+S K L+ L +SW+R
Sbjct: 634 GYFVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRP 693
Query: 718 EVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCL 777
+ +E ++LEAL+P+ Y + + G P W++ L ++ S+ + C++C
Sbjct: 694 NRYESEE--VKVLEALKPHPNLKY-LEIIDFCGFCLPDWMNHSVLKNVVSILISGCENCS 750
Query: 778 NLPELWKLPSLKYLKLSN-MIHVIYLFHESY-DGEGLMALKTLFLEKLPNLIGLSREERV 835
LP +LP L+ L+L + + V Y+ + +L+ L + NL GL R +
Sbjct: 751 CLPPFGELPCLESLELQDGSVEVEYVEDSGFLTRRRFPSLRKLHIGGFCNLKGLQRMKGA 810
Query: 836 -MFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEEL 894
FP L+ ++I++CP + P L S+ L I G+ + SSI L +L SL N +
Sbjct: 811 EQFPVLEEMKISDCP-MFVFPTLSSVKKLEIWGEADAGGLSSISNLSTLTSLKIFSNHTV 869
Query: 895 IYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQ 953
+ + +NL + L L LK LPT + ++ L+ L I C +E LP E ++
Sbjct: 870 TSLLEEMFKNLEN-LIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESLPEEGLE 927
Score = 35.0 bits (79), Expect = 1.1
Identities = 36/155 (23%), Positives = 68/155 (43%), Gaps = 16/155 (10%)
Query: 914 FHRHSKLKMLPTEMIHIHA----LQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKL 969
F H + L T M A ++Q+ + D ++M + + K++ +G ++
Sbjct: 470 FKMHDLIHDLATSMFSASASSRSIRQINVKD-------DEDMMFIVTNYKDMMSIGFSEV 522
Query: 970 KLS---SDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNL 1026
S S F+ L L + + SE E ++ + L+ L LS + LP+ + L
Sbjct: 523 VSSYSPSLFKRFVSLRVLNLSN-SEFEQLPSSVGDLVHLRYLDLSG-NKICSLPKRLCKL 580
Query: 1027 TLLHEINIYSCPKLACLPTSIQQISGLEILSIHDC 1061
L +++Y+C L+CLP ++ L L + C
Sbjct: 581 QNLQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHC 615
Score = 33.9 bits (76), Expect = 2.5
Identities = 26/91 (28%), Positives = 43/91 (46%), Gaps = 2/91 (2%)
Query: 945 EELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTT 1004
E+LP+ V +H L+ LD+ G L L L+TL + +C + + + +
Sbjct: 548 EQLPSSVGDLVH-LRYLDLSGNKICSLPKRLCKLQNLQTLDLYNCQSLSCLPKQTSKLCS 606
Query: 1005 LKSLTLSDLPNLEYLPECIGNLTLLHEINIY 1035
L++L L P L +P IG LT L + +
Sbjct: 607 LRNLVLDHCP-LTSMPPRIGLLTCLKTLGYF 636
>R131_ARATH (Q9LRR4) Putative disease resistance RPP13-like protein 1
Length = 1054
Score = 465 bits (1197), Expect = e-130
Identities = 327/997 (32%), Positives = 534/997 (52%), Gaps = 85/997 (8%)
Query: 33 RLASLLTTIKATLEDAEEKQFSDSEIGRDVKDWLLKLKDAAYTLDDIMDECATEALEMEY 92
RL++ L TI A L DAEEKQ ++ V+ W+ +L+D Y +D +D+ ATEAL +
Sbjct: 41 RLSTALLTITAVLIDAEEKQITNPV----VEKWVNELRDVVYHAEDALDDIATEALRLNI 96
Query: 93 KASKCGLSH-KHIAFRYKLAK-----------KMKRIGVWLDDIAAEKNKFHLTEIVRER 140
A + + + R L +++++ + L+ +A+++N L E+
Sbjct: 97 GAESSSSNRLRQLRGRMSLGDFLDGNSEHLETRLEKVTIRLERLASQRNILGLKEL---- 152
Query: 141 SGVVPDWR-QTTSIVTQPLVYGRNEDKDKIVDFLVGDASEQEDLSVYPIVGLGGLGKTTL 199
+ ++P R TTS+V + V+GR++DKD+I+ FL+ + + ++V IVG+GG+GKTTL
Sbjct: 153 TAMIPKQRLPTTSLVDESEVFGRDDDKDEIMRFLIPENGKDNGITVVAIVGIGGVGKTTL 212
Query: 200 AQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSCEDLDLELLQRKLQDLL 259
+QL++N + ++F K+W VSE+F + ++TK + E T + CE DL++LQ KL++ L
Sbjct: 213 SQLLYNDQHVRSYFGTKVWAHVSEEFDVFKITKKVYESVTSRPCEFTDLDVLQVKLKERL 272
Query: 260 RRK--RYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIMGTIPHHELS 317
+LLVLDD+WN+ +W L+ +G+ ILVTTR +VA IM + H L
Sbjct: 273 TGTGLPFLLVLDDLWNENFADWDLLRQPFIHAAQGSQILVTTRSQRVASIMCAVHVHNLQ 332
Query: 318 RLSDEDCWELFKQRAFGPNE-VQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKE 376
LSD DCW LF + FG E +E+ + + I+ KC G PLA LG +LRF+ + E
Sbjct: 333 PLSDGDCWSLFMKTVFGNQEPCLNREIGDLAERIVHKCRGLPLAVKTLGGVLRFEGKVIE 392
Query: 377 WLYVKESKLWNLQGE-AYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWT 435
W V S++W+L + + ++P LR+SY +LP L++CF++C++FPK K ++ LW
Sbjct: 393 WERVLSSRIWDLPADKSNLLPVLRVSYYYLPAHLKRCFAYCSIFPKGHAFEKDKVVLLWM 452
Query: 436 ANGFISSNQMLE-ADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQ 494
A GF+ + + +++GNE ++EL RS + T+ T + MHD +++LA +
Sbjct: 453 AEGFLQQTRSSKNLEELGNEYFSELESRSLLQKTK-------TRYIMHDFINELAQFASG 505
Query: 495 DVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQ-LHHVKSLKTYMEFNFDVYEAGQL 553
+ +D +SE TR+ L Y R+++AE + L VK L+T++ +
Sbjct: 506 EFSSKFEDGCKLQVSERTRY-LSYLRDNYAEPMEFEALREVKFLRTFLPLSLTNSSRSCC 564
Query: 554 SPQVLNCYSLRVLLSHRLNNLSS-SIGRL--------KYLRYLDISEGRFKNLPNSLCKL 604
Q+++ L L R+ +LS I RL + R+LD+S + LP SLC +
Sbjct: 565 LDQMVSEKLLPTLTRLRVLSLSHYKIARLPPDFFKNISHARFLDLSRTELEKLPKSLCYM 624
Query: 605 CNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVG 664
NL+ L L C SL++LP ++ L L+ L L L +PR+ G+L SL TL+ + V
Sbjct: 625 YNLQTLLLSYCSSLKELPTDISNLINLRYLDLIG-TKLRQMPRRFGRLKSLQTLTTFFVS 683
Query: 665 EERGFLLEELGQL-NLKGQLHIKNLERLKSVTDAKKANM-SRKKLNQLWLSW-------E 715
G + ELG L +L G+L I L+R+ V DA +AN+ S+K L ++ W E
Sbjct: 684 ASDGSRISELGGLHDLHGKLKIVELQRVVDVADAAEANLNSKKHLREIDFVWRTGSSSSE 743
Query: 716 RNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKS 775
N +N ++ E L+P+ + + + Y G FP W+S PS + + + L +C+
Sbjct: 744 NNTNPHRTQNEAEVFEKLRPH-RHIEKLAIERYKGRRFPDWLSDPSFSRIVCIRLRECQY 802
Query: 776 CLNLPELWKLPSLKYLKLSNMIHVIYLFHESY---------DGEGLMALKTLFLEKLPNL 826
C +LP L +LP LK L +S M+ + + + Y D + +L+TL + LP+
Sbjct: 803 CTSLPSLGQLPCLKELHISGMVGLQSIGRKFYFSDQQLRDQDQQPFRSLETLRFDNLPDW 862
Query: 827 -----IGLSREERVMFPRLKALEITECPNLLG-LPC-LPSLSDLYIQ--GKYNQQLPSSI 877
+ ++R + +FP LK L I CP L G LP LPSL L+I G + Q
Sbjct: 863 QEWLDVRVTRGD--LFPSLKKLFILRCPELTGTLPTFLPSLISLHIYKCGLLDFQPDHHE 920
Query: 878 HKLGSLESLHF-SDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHI---HAL 933
+ +L++L S + L+ FP L + A+ L L + + L L H+ +AL
Sbjct: 921 YSYRNLQTLSIKSSCDTLVKFP---LNHFAN-LDKLEVDQCTSLYSLELSNEHLRGPNAL 976
Query: 934 QQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLK 970
+ L INDC+N++ LP + L ++ I C L+
Sbjct: 977 RNLRINDCQNLQLLPK--LNALPQNLQVTITNCRYLR 1011
Score = 41.2 bits (95), Expect = 0.016
Identities = 31/125 (24%), Positives = 65/125 (51%), Gaps = 7/125 (5%)
Query: 870 NQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIH 929
+++L ++ +L L H+ ++ P +N+ S + L R ++L+ LP + +
Sbjct: 570 SEKLLPTLTRLRVLSLSHY----KIARLPPDFFKNI-SHARFLDLSR-TELEKLPKSLCY 623
Query: 930 IHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSC 989
++ LQ L ++ C +++ELP ++ L +L+ LD++G ++ F L L+TL
Sbjct: 624 MYNLQTLLLSYCSSLKELPTDI-SNLINLRYLDLIGTKLRQMPRRFGRLKSLQTLTTFFV 682
Query: 990 SEVEG 994
S +G
Sbjct: 683 SASDG 687
Score = 32.0 bits (71), Expect = 9.5
Identities = 27/115 (23%), Positives = 51/115 (43%), Gaps = 6/115 (5%)
Query: 921 KMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTC 980
K+LPT + L+ L ++ + I LP + + + + LD+ + KL Y+
Sbjct: 572 KLLPT----LTRLRVLSLSHYK-IARLPPDFFKNISHARFLDLSRTELEKLPKSLCYMYN 626
Query: 981 LETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIY 1035
L+TL + CS ++ + ++ L+ L L L +P G L L + +
Sbjct: 627 LQTLLLSYCSSLKELPTDISNLINLRYLDLIG-TKLRQMPRRFGRLKSLQTLTTF 680
>DR21_ARATH (Q9LRR5) Putative disease resistance protein At3g14460
Length = 1424
Score = 388 bits (997), Expect = e-107
Identities = 282/866 (32%), Positives = 443/866 (50%), Gaps = 70/866 (8%)
Query: 33 RLASLLTTIKATLEDAEEKQFSDSEIGRDVKDWLLKLKDAAYTLDDIMDECATEALEMEY 92
RL L T L DA+++ +E R+VK WL +KDA + +DI+DE TEAL
Sbjct: 38 RLKVALVTANPVLADADQR----AEHVREVKHWLTGIKDAFFQAEDILDELQTEALRRRV 93
Query: 93 KASKCGLSHK-------HIAFRYKLAKKMKRIGVWLDDIAAEKNKFHLTEIVRERSGVVP 145
A GL A + K+ KM+++ L+ L E R P
Sbjct: 94 VAEAGGLGGLFQNLMAGREAIQKKIEPKMEKVVRLLEHHVKHIEVIGLKEYSETRE---P 150
Query: 146 DWRQTTSI----VTQPLVYGRNEDKDKIVDFLVGDASEQEDLS-----VYPIVGLGGLGK 196
WRQ + + Q + GR EDK +V+ L+ D +++S V +VG+ G+GK
Sbjct: 151 QWRQASRSRPDDLPQGRLVGRVEDKLALVNLLLSD----DEISIGKPAVISVVGMPGVGK 206
Query: 197 TTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSCEDLDLELLQRKLQ 256
TTL ++VFN ++ HFE+K+W+ +F + +TKA+++ T + DL LQ +L+
Sbjct: 207 TTLTEIVFNDYRVTEHFEVKMWISAGINFNVFTVTKAVLQDITSSAVNTEDLPSLQIQLK 266
Query: 257 DLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIMGTIPHHEL 316
L KR+LLVLDD W++ W+ + +G+ I++TTR V+ + +++
Sbjct: 267 KTLSGKRFLLVLDDFWSESDSEWESFQVAFTDAEEGSKIVLTTRSEIVSTVAKAEKIYQM 326
Query: 317 SRLSDEDCWELFKQRAFGPNEVQ--QKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREE 374
+++E+CWEL + AFG V +EL +GK I ++C G PLAA A+ S LR K
Sbjct: 327 KLMTNEECWELISRFAFGNISVGSINQELEGIGKRIAEQCKGLPLAARAIASHLRSKPNP 386
Query: 375 KEWLYVKESKLWNLQGEAYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLW 434
+W V SK ++ + ++P L+LSY LP +L++CF+ C++FPK + ++ L+ LW
Sbjct: 387 DDWYAV--SKNFSSYTNS-ILPVLKLSYDSLPPQLKRCFALCSIFPKGHVFDREELVLLW 443
Query: 435 TANGFI---SSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGS 491
A + S++ LE DIGN+ +L +SFF+ + +T F MHDL++DLA +
Sbjct: 444 MAIDLLYQPRSSRRLE--DIGNDYLGDLVAQSFFQRLDIT----MTSFVMHDLMNDLAKA 497
Query: 492 VTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDV-YEA 550
V+ D C +D+++ + TRH A + + L+T + FN E+
Sbjct: 498 VSGDFCFRLEDDNIPEIPSTTRHFSFSRSQCDASVAFRSICGAEFLRTILPFNSPTSLES 557
Query: 551 GQLSPQVLN-----CYSLRVL-LSH-RLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCK 603
QL+ +VLN LR+L LSH ++ NL S+ LK LRYLD+S + K LP +C
Sbjct: 558 LQLTEKVLNPLLNALSGLRILSLSHYQITNLPKSLKGLKLLRYLDLSSTKIKELPEFVCT 617
Query: 604 LCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIV 663
LCNL+ L L C L LP + L L+ L L L +P I KL SL LS +++
Sbjct: 618 LCNLQTLLLSNCRDLTSLPKSIAELINLRLLDLVG-TPLVEMPPGIKKLRSLQKLSNFVI 676
Query: 664 GEERGFLLEELGQL-NLKGQLHIKNLERLKSVTDAKKANMSRKK-LNQLWLSWE------ 715
G G L EL +L +L+G L I L+ + ++AK A + RK L+ L L W
Sbjct: 677 GRLSGAGLHELKELSHLRGTLRISELQNVAFASEAKDAGLKRKPFLDGLILKWTVKGSGF 736
Query: 716 -RNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCK 774
+ L + +++L L+P+ L +F + Y G FP+W+ S + S+ L C
Sbjct: 737 VPGSFNALACDQKEVLRMLEPHPH-LKTFCIESYQGGAFPKWLGDSSFFGITSVTLSSCN 795
Query: 775 SCLNLPELWKLPSLKYLKLSNM-----IHVIYLFHESYDG----EGLMALKTLFLEKLPN 825
C++LP + +LPSLKYL + + + + F E+ + L LK + +
Sbjct: 796 LCISLPPVGQLPSLKYLSIEKFNILQKVGLDFFFGENNSRGVPFQSLQILKFYGMPRWDE 855
Query: 826 LIGLSREERVMFPRLKALEITECPNL 851
I E+ + FP L+ L I CP+L
Sbjct: 856 WICPELEDGI-FPCLQKLIIQRCPSL 880
Score = 55.8 bits (133), Expect = 6e-07
Identities = 67/257 (26%), Positives = 105/257 (40%), Gaps = 32/257 (12%)
Query: 840 LKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPD 899
L++L I C L LP +L++ Y ++ L + H L S H + +Y D
Sbjct: 1093 LQSLHIDSCDGLTSLP--ENLTESY--PNLHELLIIACHSLESFPGSHPPTTLKTLYIRD 1148
Query: 900 GILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYI-NDCRNIEELPNEVMQRLHSL 958
N L+ PT L+ L+I + C N+ P + +L SL
Sbjct: 1149 CKKLNFTESLQ-------------PTRSYS--QLEYLFIGSSCSNLVNFPLSLFPKLRSL 1193
Query: 959 KELDIVGCDKLKLSSDFQYL----TCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLP 1014
D C+ K S L LE+L I C +E F + L S+ LS+
Sbjct: 1194 SIRD---CESFKTFSIHAGLGDDRIALESLEIRDCPNLETFPQGGLPTPKLSSMLLSNCK 1250
Query: 1015 NLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGE 1074
L+ LPE + LT L + I CP++ +P S L L I C KL R + + +
Sbjct: 1251 KLQALPEKLFGLTSLLSLFIIKCPEIETIPGG-GFPSNLRTLCISLCDKLTPRIEWGLRD 1309
Query: 1075 DWPKIVHVQYIEIENDN 1091
+ +++ +EI+ N
Sbjct: 1310 ----LENLRNLEIDGGN 1322
Score = 49.7 bits (117), Expect = 4e-05
Identities = 76/271 (28%), Positives = 106/271 (39%), Gaps = 62/271 (22%)
Query: 765 LKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLP 824
LK+L + DCK LN E + P+ Y +L YLF G L L P
Sbjct: 1141 LKTLYIRDCKK-LNFTESLQ-PTRSYSQLE------YLFI----GSSCSNLVNFPLSLFP 1188
Query: 825 NLIGLSREERVMFPR-------------LKALEITECPNLL-----GLPCLPSLSDLYIQ 866
L LS + F L++LEI +CPNL GLP P LS + +
Sbjct: 1189 KLRSLSIRDCESFKTFSIHAGLGDDRIALESLEIRDCPNLETFPQGGLPT-PKLSSMLLS 1247
Query: 867 G-KYNQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKL----- 920
K Q LP + L SL SL E+ P G S L+TL KL
Sbjct: 1248 NCKKLQALPEKLFGLTSLLSLFIIKCPEIETIPGG---GFPSNLRTLCISLCDKLTPRIE 1304
Query: 921 ---------------------KMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLK 959
+ P E + ++ L I+ N++ L + +++
Sbjct: 1305 WGLRDLENLRNLEIDGGNEDIESFPEEGLLPKSVFSLRISRFENLKTLNRKGFHDTKAIE 1364
Query: 960 ELDIVGCDKLKLSSDFQYLTCLETLAIGSCS 990
++I GCDKL++S D + L L L I SCS
Sbjct: 1365 TMEISGCDKLQISID-EDLPPLSCLRISSCS 1394
Score = 45.4 bits (106), Expect = 8e-04
Identities = 37/128 (28%), Positives = 65/128 (49%), Gaps = 10/128 (7%)
Query: 902 LRNLASPLKTLGFHRH-----SKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLH 956
+ NL LK L R+ +K+K LP + + LQ L +++CR++ LP + + L
Sbjct: 585 ITNLPKSLKGLKLLRYLDLSSTKIKELPEFVCTLCNLQTLLLSNCRDLTSLPKSIAE-LI 643
Query: 957 SLKELDIVGCDKLKLSSDFQYLTCLETLA---IGSCSEVEGFHEALQHMTTLKSLTLSDL 1013
+L+ LD+VG +++ + L L+ L+ IG S G HE + +L +S+L
Sbjct: 644 NLRLLDLVGTPLVEMPPGIKKLRSLQKLSNFVIGRLSGA-GLHELKELSHLRGTLRISEL 702
Query: 1014 PNLEYLPE 1021
N+ + E
Sbjct: 703 QNVAFASE 710
Score = 34.7 bits (78), Expect = 1.5
Identities = 20/57 (35%), Positives = 33/57 (57%), Gaps = 2/57 (3%)
Query: 607 LEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIV 663
LE L++ C +L+ P G +L ++ L +C L +LP ++ LTSL LS +I+
Sbjct: 1217 LESLEIRDCPNLETFPQGGLPTPKLSSMLLSNCKKLQALPEKLFGLTSL--LSLFII 1271
>R134_ARATH (Q38834) Putative disease resistance RPP13-like protein
4
Length = 852
Score = 243 bits (619), Expect = 3e-63
Identities = 223/794 (28%), Positives = 365/794 (45%), Gaps = 47/794 (5%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M +AV+ + L ++ ++ + ++ L S L +++ L+DAE ++ ++ +
Sbjct: 1 MVDAVVTVFLEKTLNILEEKGRTVSDYRKQLEDLQSELKYMQSFLKDAERQKRTNETLRT 60
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALE--MEYKASKCGLSHKH---IAFRYKLAKKMK 115
V D L++ Y +DI+ +C + E ++S LS H + +YK +K+++
Sbjct: 61 LVAD----LRELVYEAEDILVDCQLADGDDGNEQRSSNAWLSRLHPARVPLQYKKSKRLQ 116
Query: 116 RIGVWLDDIAAEKNKFH--LTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFL 173
I + I ++ + +T R W TQ V G DK KI ++L
Sbjct: 117 EINERITKIKSQVEPYFEFITPSNVGRDNGTDRWSSPVYDHTQ--VVGLEGDKRKIKEWL 174
Query: 174 VGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKA 233
S L + VG+GGLGKTT+AQ VFN +I + FE +IWV VS+ FT +++ ++
Sbjct: 175 F--RSNDSQLLIMAFVGMGGLGKTTIAQEVFNDKEIEHRFERRIWVSVSQTFTEEQIMRS 232
Query: 234 IIEGATKKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGA 293
I+ S D D+ L RK+Q L KRYL+V+DDVW+ W ++ L G+G
Sbjct: 233 ILRNLGDASVGD-DIGTLLRKIQQYLLGKRYLIVMDDVWDKNLSWWDKIYQGLP-RGQGG 290
Query: 294 SILVTTRLPKVAKIMGTIPH--HELSRLSDEDCWELFKQRAFGPNE--VQQKELVIVGKE 349
S++VTTR VAK + H LS ++ W LF AF N+ ++ EL VGKE
Sbjct: 291 SVIVTTRSESVAKRVQARDDKTHRPELLSPDNSWLLFCNVAFAANDGTCERPELEDVGKE 350
Query: 350 IIKKCGGFPLAAIALGSLLRFK-REEKEWLYVKESKLWNLQGEA----YVMPALRLSYLH 404
I+ KC G PL A+G LL K EW + E L+G VM +L+LSY
Sbjct: 351 IVTKCKGLPLTIKAVGGLLLCKDHVYHEWRRIAEHFQDELRGNTSETDNVMSSLQLSYDE 410
Query: 405 LPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSF 464
LP L+ C +L+P+D +I KQ L+ W GF+ A + G + ++ L R
Sbjct: 411 LPSHLKSCILTLSLYPEDCVIPKQQLVHGWIGEGFVMWRNGRSATESGEDCFSGLTNRCL 470
Query: 465 FENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFA 524
E + G I K+HD+V DL + + D+ RHL I +F
Sbjct: 471 IEVVDKTYSGTIITCKIHDMVRDLVIDIAK------KDSFSNPEGLNCRHLGI--SGNFD 522
Query: 525 EANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVL------LSHRLNNLSSSI 578
E H ++ + + + L+ + +C LRVL L+ + I
Sbjct: 523 EKQIKVNHKLRGVVSTTKTGEVNKLNSDLAKKFTDCKYLRVLDISKSIFDAPLSEILDEI 582
Query: 579 GRLKYLRYLDISEGR-FKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLR 637
L++L L +S P S+ L NL++L C +L++L + K+L L +
Sbjct: 583 ASLQHLACLSLSNTHPLIQFPRSMEDLHNLQILDASYCQNLKQLQPCIVLFKKLLVLDMT 642
Query: 638 DCDSLTSLPRQIGKLTSLNTLSKY-IVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTD 696
+C SL P+ IG L L L + G L E+ L +L + +L R + +
Sbjct: 643 NCGSLECFPKGIGSLVKLEVLLGFKPARSNNGCKLSEVKNLTNLRKLGL-SLTRGDQIEE 701
Query: 697 AKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQW 756
+ ++ L++L +S N +++ ++AL P +L+ + Y G P W
Sbjct: 702 EELDSLI--NLSKL-MSISINCYDSYGDDLITKIDALTP-PHQLHELSLQFYPGKSSPSW 757
Query: 757 ISIPSLNDLKSLEL 770
+S L L+ + +
Sbjct: 758 LSPHKLPMLRYMSI 771
Score = 35.0 bits (79), Expect = 1.1
Identities = 22/84 (26%), Positives = 40/84 (47%), Gaps = 3/84 (3%)
Query: 973 SDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEI 1032
+ Q+L CL ++ + + F +++ + L+ L S NL+ L CI L +
Sbjct: 583 ASLQHLACL---SLSNTHPLIQFPRSMEDLHNLQILDASYCQNLKQLQPCIVLFKKLLVL 639
Query: 1033 NIYSCPKLACLPTSIQQISGLEIL 1056
++ +C L C P I + LE+L
Sbjct: 640 DMTNCGSLECFPKGIGSLVKLEVL 663
>R133_ARATH (Q9STE7) Putative disease resistance RPP13-like protein
3
Length = 847
Score = 232 bits (591), Expect = 5e-60
Identities = 241/905 (26%), Positives = 408/905 (44%), Gaps = 114/905 (12%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M +AV VL + + E+ +G + L + LT I L+D E ++ D E+
Sbjct: 1 MVDAVTGFVLNKIGGYLINEVLALMGVKDDLEELKTELTCIHGYLKDVEARERED-EVS- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKRIGVW 120
K+W + D AY ++D++D T L++E ++ + GL + K+ KK +
Sbjct: 59 --KEWTKLVLDIAYDIEDVLD---TYFLKLEERSLRRGL----LRLTNKIGKKRDAYNI- 108
Query: 121 LDDIAAEKNKFHLTEIVRERSGV---------------VPDWRQTTSIVTQPLVYGRNED 165
++DI K + RE G+ V R+ + + LV G +D
Sbjct: 109 VEDIRTLKRRILDITRKRETFGIGSFNEPRGENITNVRVRQLRRAPPVDQEELVVGLEDD 168
Query: 166 KDKIVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDF 225
++ L+ D +E++ + I G+GGLGKT LA+ ++N + F+ + W VS+++
Sbjct: 169 VKILLVKLLSD-NEKDKSYIISIFGMGGLGKTALARKLYNSGDVKRRFDCRAWTYVSQEY 227
Query: 226 TLKRMTKAIIEGATKKSCEDLDL-------ELLQRKLQDLLRRKRYLLVLDDVWNDKQEN 278
+ + II S E+++ E L+ L LL K Y++V+DDVW+ +
Sbjct: 228 KTRDILIRIIRSLGIVSAEEMEKIKMFEEDEELEVYLYGLLEGKNYMVVVDDVWDP--DA 285
Query: 279 WQRLKSVLACGGKGASILVTTRLPKVAK-IMGTIPHHELSRLSDEDCWELFKQRAFGPNE 337
W+ LK L C +G+ +++TTR+ +A+ + GT+ H+L L+ E+ W LF+++AF E
Sbjct: 286 WESLKRALPCDHRGSKVIITTRIRAIAEGVEGTVYAHKLRFLTFEESWTLFERKAFSNIE 345
Query: 338 VQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWN--LQGEAYVM 395
++L GKE++KKCGG PLA + L LL KR EW V S LW ++
Sbjct: 346 KVDEDLQRTGKEMVKKCGGLPLAIVVLSGLLSRKR-TNEWHEVCAS-LWRRLKDNSIHIS 403
Query: 396 PALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEV 455
LS+ + +L+ CF + ++FP+D I + LI L A GFI ++ + +D+
Sbjct: 404 TVFDLSFKEMRHELKLCFLYFSVFPEDYEIKVEKLIHLLVAEGFIQEDEEMMMEDVARCY 463
Query: 456 WNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHL 515
+EL RS + E + G++ ++HDL+ DLA +++ + N + S+ R
Sbjct: 464 IDELVDRSLVK-AERIERGKVMSCRIHDLLRDLAIKKAKELNFVNVYNEKQHSSDICRRE 522
Query: 516 LIYN-RNSFAEANSIQLHHVKSLKTYME-FNFDVYEAGQLS---PQVLNCYSLRVLLSHR 570
++++ N + + ++S E F L +VLN L + +
Sbjct: 523 VVHHLMNDYYLCDRRVNKRMRSFLFIGERRGFGYVNTTNLKLKLLRVLNMEGLLFVSKNI 582
Query: 571 LNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKR 630
N L IG L +LRYL I++ LP S+ L L+ L G Q
Sbjct: 583 SNTLPDVIGELIHLRYLGIADTYVSILPASISNLRFLQTLDASGNDPFQ----------- 631
Query: 631 LQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLN------LKGQLH 684
D LTSL IGK + + ++GE G L+ L ++ L +L
Sbjct: 632 ----YTTDLSKLTSLRHVIGKF-----VGECLIGE--GVNLQTLRSISSYSWSKLNHEL- 679
Query: 685 IKNLERLKSVTDAKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSFG 744
++NL+ L+ +K + R LN ++S+ + +N+ + ++ + S
Sbjct: 680 LRNLQDLEIYDHSKWVDQRRVPLN--FVSFSK------PKNLRVLKLEMRNFKLSSESRT 731
Query: 745 VGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLN-LPELWKLPSLKYLKLSNMIHVIYLF 803
G FP L+SL LV N +P L KLP L+ L L +
Sbjct: 732 TIGLVDVNFP---------SLESLTLVGTTLEENSMPALQKLPRLEDLVLKDC------- 775
Query: 804 HESYDGEGLMALKTLFLEKLPNLIGLSREERVMFPRLKALEITECPNLLGLPCLPSLSDL 863
+Y G +M++ +L NL +S E R L L I E +PSL L
Sbjct: 776 --NYSGVKIMSISAQGFGRLKNL-EMSMERR--GHGLDELRIEE-------EAMPSLIKL 823
Query: 864 YIQGK 868
++G+
Sbjct: 824 TVKGR 828
>RPM1_ARATH (Q39214) Disease resistance protein RPM1 (Resistance to
Pseudomonas syringae protein 3)
Length = 926
Score = 231 bits (588), Expect = 1e-59
Identities = 234/950 (24%), Positives = 431/950 (44%), Gaps = 133/950 (14%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEK--QFSDSEI 58
MA A ++ +G + ++ E L G E +++ L +K+ LED + S +
Sbjct: 1 MASATVDFGIGRILSVLENETLLLSGVHGEIDKMKKELLIMKSFLEDTHKHGGNGSTTTT 60
Query: 59 GRDVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH--KHIAFRYKLAKKMKR 116
+ + ++ +D AY ++DI+DE A H +++ R+ +A+K+
Sbjct: 61 TQLFQTFVANTRDLAYQIEDILDEFGYHIHGYRSCAKIWRAFHFPRYMWARHSIAQKLGM 120
Query: 117 IGVWLDDIAAEKNKFHLTEIVRERSGVVP-----DWRQTTSIVTQPLVYGRNE------D 165
+ V + I+ +++ +E ++ ++P D + +I L + N
Sbjct: 121 VNVMIQSISDSMKRYYHSE--NYQAALLPPIDDGDAKWVNNISESSLFFSENSLVGIDAP 178
Query: 166 KDKIVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDF 225
K K++ L+ S + V +VG+GG GKTTL+ +F + HFE WV +S+ +
Sbjct: 179 KGKLIGRLL---SPEPQRIVVAVVGMGGSGKTTLSANIFKSQSVRRHFESYAWVTISKSY 235
Query: 226 TLKRMTKAIIEGATKKSCEDLDLEL-------LQRKLQDLLRRKRYLLVLDDVWNDKQEN 278
++ + + +I+ K++ + EL L KL + L+ KRY++VLDDVW
Sbjct: 236 VIEDVFRTMIKEFYKEADTQIPAELYSLGYRELVEKLVEYLQSKRYIVVLDDVWTTGL-- 293
Query: 279 WQRLKSVLACGGKGASILVTTRLPKVAKIMGTI--PHHELSRLSDEDCWELFKQRAFGPN 336
W+ + L G G+ +++TTR VA I HE+ L +++ W LF +AF P
Sbjct: 294 WREISIALPDGIYGSRVMMTTRDMNVASFPYGIGSTKHEIELLKEDEAWVLFSNKAF-PA 352
Query: 337 EVQQ---KELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGE-- 391
++Q + L + ++++++C G PLA +LGS++ K+ E EW V + W L
Sbjct: 353 SLEQCRTQNLEPIARKLVERCQGLPLAIASLGSMMSTKKFESEWKKVYSTLNWELNNNHE 412
Query: 392 -AYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADD 450
V + LS+ LP L++CF +C+LFP + + ++ LI +W A F+ + ++A++
Sbjct: 413 LKIVRSIMFLSFNDLPYPLKRCFLYCSLFPVNYRMKRKRLIRMWMAQRFVEPIRGVKAEE 472
Query: 451 IGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVT--QDVCCITDDNSMRTM 508
+ + NEL +R+ + FG+ FKMHD++ ++A SV+ + C + +D+S
Sbjct: 473 VADSYLNELVYRNMLQVILWNPFGRPKAFKMHDVIWEIALSVSKLERFCDVYNDDSDGDD 532
Query: 509 SEET------RHLLIYNRNSFAEANSIQLHHV---KSLKTYMEFNFDVYEAGQLSPQVLN 559
+ ET RHL I + + LH + S K ME L P LN
Sbjct: 533 AAETMENYGSRHLCIQKEMTPDSIRATNLHSLLVCSSAKHKME----------LLPS-LN 581
Query: 560 CYSLRVLLSHRLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQ 619
L ++ L + + L+YL++S+ + K LP + KL NLE L ++
Sbjct: 582 LLRALDLEDSSISKLPDCLVTMFNLKYLNLSKTQVKELPKNFHKLVNLETLNTKHS-KIE 640
Query: 620 KLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQL-- 677
+LP G+ +LK+L+ L + R G ++ N Y++G + +L L
Sbjct: 641 ELPLGMWKLKKLR--------YLITFRRNEGHDSNWN----YVLGTRVVPKIWQLKDLQV 688
Query: 678 ----NLKGQLHIKNLERLKSVTDAKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEAL 733
N + +L IKNL + +T + R+ L S N++ ++
Sbjct: 689 MDCFNAEDEL-IKNLGCMTQLTRISLVMVRREHGRDLCDS--LNKIKRI----------- 734
Query: 734 QPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLNLPELWKLPSLKYLK- 792
+++S+ S+++ + LE+ D + ++ +L+ L+ +
Sbjct: 735 ---------------------RFLSLTSIDEEEPLEIDDLIATASIEKLFLAGKLERVPS 773
Query: 793 -LSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREERVMFPR---------LKA 842
+ + ++ YL G L L ++ LP L+ LS M PR LK
Sbjct: 774 WFNTLQNLTYL---GLRGSQLQENAILSIQTLPRLVWLSFYNAYMGPRLRFAQGFQNLKI 830
Query: 843 LEITECPNLLGL----PCLPSLSDLYIQG-KYNQQLPSSIHKLGSLESLH 887
LEI + +L + + L LY++ + + +P I L +L+ LH
Sbjct: 831 LEIVQMKHLTEVVIEDGAMFELQKLYVRACRGLEYVPRGIENLINLQELH 880
>RP8H_ARATH (P59584) Disease resistance protein RPH8A (RPP8 homolog
A)
Length = 910
Score = 223 bits (569), Expect = 2e-57
Identities = 239/900 (26%), Positives = 409/900 (44%), Gaps = 125/900 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAE + L L +L+ +E G D++ + L L ++++ L+DA+ K+
Sbjct: 1 MAEGFVSFGLEKLWDLLSRESERLQGIDEQLDGLKRQLRSLQSLLKDADAKKHGSDR--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH--KHIAFRYKLAKKMKRIG 118
V+++L +KD + +DI++ L E K K + + + R+K+A ++ I
Sbjct: 58 -VRNFLEDVKDLVFDAEDIIESYVLNKLRGEGKGVKKHVRRLARFLTDRHKVASDIEGIT 116
Query: 119 VWLDDIAAEKNKFHLTEIV--------RERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIV 170
+ ++ E F + +I+ +ER V + RQT ++ + G + ++V
Sbjct: 117 KRISEVIGEMQSFGIQQIIDGGRSLSLQERQRVQREIRQTYPDSSESDLVGVEQSVTELV 176
Query: 171 DFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRM 230
LV E + V I G+GG+GKTTLA+ VF+HD + HF+ WVCVS+ FT K +
Sbjct: 177 CHLV----ENDVHQVVSIAGMGGIGKTTLARQVFHHDLVRRHFDGFAWVCVSQQFTQKHV 232
Query: 231 TKAIIEGATKKSCEDLDLE--LLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLAC 288
+ I++ E L ++ +Q KL LL RYL+VLDDVW K+E+W R+K+V
Sbjct: 233 WQRILQELQPHDGEILQMDEYTIQGKLFQLLETGRYLVVLDDVW--KKEDWDRIKAVFP- 289
Query: 289 GGKGASILVTTRLPKVA-KIMGTIPHHELSRLSDEDCWELFKQRAF---GPNEVQ-QKEL 343
+G +L+T+R V T S L+ E+ W+L ++ F EV+ +E+
Sbjct: 290 RKRGWKMLLTSRNEGVGIHADPTCLTFRASILNPEESWKLCERIVFPRRDETEVRLDEEM 349
Query: 344 VIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA--------YVM 395
+GKE++ CGG PLA ALG LL K EW V ++ + G + V
Sbjct: 350 EAMGKEMVTHCGGLPLAVKALGGLLANKHTVPEWKRVSDNIGSQIVGGSCLDDNSLNSVY 409
Query: 396 PALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEV 455
L LSY LP L+ CF A +P+D I Q L + W A G + + D G
Sbjct: 410 RILSLSYEDLPTHLKHCFLHLAHYPEDSKIYTQDLFNYWAAEGIYDGSTI---QDSGEYY 466
Query: 456 WNELYWRSFF--ENTENVGFGQITIFKMHDLVHDLAGS---------VTQDVCCITDDNS 504
EL R+ +N + +I +MHD++ ++ S + +D C + N+
Sbjct: 467 LEELVRRNLVIADNRYLISEFKIKNCQMHDMMREVCLSKAKEENFLQIIKDPTCTSTINA 526
Query: 505 --------MRTMSEETRHLLIYNRNS---------FAE---ANSIQLHHVKSLKTYMEFN 544
+ S + H+L + RN+ F E S + H +L ++ +
Sbjct: 527 QSPSRSRRLSIHSGKAFHILGHKRNAKVRSLIVSRFEEDFWIRSASVFHNLTLLRVLDLS 586
Query: 545 FDVYEAGQLSPQVLNCYSLRVLLSHR--LNNLSSSIGRLKYLRYLDIS--EGRFKNLPNS 600
+ +E G+L + LR L + +++L S++ LK L YL++S ++PN
Sbjct: 587 WVKFEGGKLPCSIGGLIHLRYLRLYGAVVSHLPSTMRNLKLLLYLNLSVHNEDLIHVPNV 646
Query: 601 LCKLCNLEV------------LKLDGCVSLQKLPG---------GLTRLKRLQNL--SLR 637
L ++ L L+L V+L+ L G L R+ +L+NL SL
Sbjct: 647 LKEMIELRYLSIPVKMDDKTKLELGDLVNLEYLYGFSTQHTSVTDLLRMTKLRNLTVSLS 706
Query: 638 DCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDA 697
+ + +L + +L +L TL Y++ + ++++ +G+ L +H+K L + ++
Sbjct: 707 ERYNFKTLSSSLRELRNLETL--YVLFSRKTYMVDHMGEFVLDHFIHLKELGLVVRMSKI 764
Query: 698 KKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQ--PYAQKLYSFGVG--------G 747
+ L ++L + ++E+ ILE L Q Y VG G
Sbjct: 765 PDQHQFPPHLVHIFLFY-----CGMEEDPMPILEKLHHLKSVQLRYKAFVGRRMVCSKDG 819
Query: 748 YT---------GAYFPQWI-SIPSLNDLKSLELVDCKSCLNLPE-LWKLPSLKYLKLSNM 796
+T + WI S+ L++L + DC+ LP+ L + SLK LK+ M
Sbjct: 820 FTQLCALDISKQSELEDWIVEEGSMPCLRTLTIHDCEKLKELPDGLKYITSLKELKIEGM 879
Score = 48.1 bits (113), Expect = 1e-04
Identities = 64/251 (25%), Positives = 114/251 (44%), Gaps = 37/251 (14%)
Query: 846 TECPNLLGLPCLPSLSDLYIQGKYN-QQLPSSIHKLGSLESLH--FSDNEELIYFPDGIL 902
T +LL + L +L+ + + +YN + L SS+ +L +LE+L+ FS ++ +
Sbjct: 687 TSVTDLLRMTKLRNLT-VSLSERYNFKTLSSSLRELRNLETLYVLFSRKTYMVDHMGEFV 745
Query: 903 RNLASPLKTLGFH-RHSKLK---MLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSL 958
+ LK LG R SK+ P ++HI ++ C +EE P ++++LH L
Sbjct: 746 LDHFIHLKELGLVVRMSKIPDQHQFPPHLVHI------FLFYC-GMEEDPMPILEKLHHL 798
Query: 959 KELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEY 1018
K + + +Y + + CS+ +GF T L +L +S LE
Sbjct: 799 KSVQL------------RYKAFVGRRMV--CSK-DGF-------TQLCALDISKQSELED 836
Query: 1019 LPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPK 1078
G++ L + I+ C KL LP ++ I+ L+ L I + K GED+ K
Sbjct: 837 WIVEEGSMPCLRTLTIHDCEKLKELPDGLKYITSLKELKIEGMKREWKEKLVPGGEDYYK 896
Query: 1079 IVHVQYIEIEN 1089
+ H+ ++ N
Sbjct: 897 VQHIPDVQFIN 907
>DR11_ARATH (Q94HW3) Probable disease resistance protein RDL6/RF9
Length = 1049
Score = 219 bits (558), Expect = 3e-56
Identities = 286/1146 (24%), Positives = 496/1146 (42%), Gaps = 159/1146 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MA ++ + +L L+ +E LF G + + L L + + L+DA+ K+ + +
Sbjct: 1 MAGELISFGIQNLWNLLSQECELFQGVEDQVTELKRDLNLLSSFLKDADAKKHTSAV--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEY-KASKCGLSHKHIAF----RYKLAKKMK 115
VK+ + ++K+ Y +D ++ T LE K S S + +A R + A +
Sbjct: 58 -VKNCVEEIKEIIYDGEDTIE---TFVLEQNLGKTSGIKKSIRRLACIIPDRRRYALGIG 113
Query: 116 RIGVWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKI-----V 170
+ + + + F + + + + P + + + + +++D D + V
Sbjct: 114 GLSNRISKVIRDMQSFGVQQAIVDGGYKQPQGDKQREMRPR---FSKDDDSDFVGLEANV 170
Query: 171 DFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRM 230
LVG ++ ++ V I G+GGLGKTTLA+ VFNH+ + + F+ WVCVS+DFT +
Sbjct: 171 KKLVGYLVDEANVQVVSITGMGGLGKTTLAKQVFNHEDVKHQFDGLSWVCVSQDFTRMNV 230
Query: 231 TKAIIEGATKKSCEDLDLELLQRKLQD----LLRRKRYLLVLDDVWNDKQENWQRLKSVL 286
+ I+ K E +E+ Q LQ LL + L+VLDD+W ++E+W+ +K +
Sbjct: 231 WQKILRDLKPKEEEKKIMEMTQDTLQGELIRLLETSKSLIVLDDIW--EKEDWELIKPIF 288
Query: 287 ACGGKGASILVTTRLPKVAKIMGT-IPHHELSRLSDEDCWELFKQRAFGPNEVQQ----K 341
KG +L+T+R VA T + + L+ ED W LF++ A + + +
Sbjct: 289 P-PTKGWKVLLTSRNESVAMRRNTSYINFKPECLTTEDSWTLFQRIALPMKDAAEFKIDE 347
Query: 342 ELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA--------- 392
E +GK +IK CGG PLA LG +L K +W + E+ +L G
Sbjct: 348 EKEELGKLMIKHCGGLPLAIRVLGGMLAEKYTSHDWRRLSENIGSHLVGGRTNFNDDNNN 407
Query: 393 ---YVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFIS----SNQM 445
YV L LS+ LP L+ CF + A FP D I+ + L W A G ++
Sbjct: 408 TCNYV---LSLSFEELPSYLKHCFLYLAHFPDDYEINVKNLSYYWAAEGIFQPRHYDGEI 464
Query: 446 LEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHD--LAGSVTQDVCCITDDN 503
+ D+G+ EL R+ + +V + +HD++ + L + ++ IT
Sbjct: 465 IR--DVGDVYIEELVRRNMVISERDVKTSRFETCHLHDMMREVCLLKAKEENFLQITSSR 522
Query: 504 SM--RTMSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAG----QLSPQV 557
+ ++S T L+Y + ++ K + N ++ G L
Sbjct: 523 TSTGNSLSIVTSRRLVYQYPITLDVEK-DINDPKLRSLVVVANTYMFWGGWSWMLLGSSF 581
Query: 558 LNCYSLRVLLSHRL----NNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLD 613
+ LRVL HR L+SSIG+L +LRYL++ ++P SL L L L L
Sbjct: 582 IRLELLRVLDIHRAKLKGGKLASSIGQLIHLRYLNLKHAEVTHIPYSLGNLKLLIYLNLV 641
Query: 614 GCVSLQKL-PGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLE 672
VS L P L +++L+ L +LP+ +G+ T L + + + F +
Sbjct: 642 ILVSGSTLVPNVLKEMQQLRYL---------ALPKDMGRKTKLELSNLVKLETLKNFSTK 692
Query: 673 ELGQLNLKGQLHIKNLE-RLKSVTDAKKANMS---RKKLNQLWLSWERNEVSQLQENVEQ 728
+L+G + ++ L L+ T + S K L L ++ +E+ + +
Sbjct: 693 NCSLEDLRGMVRLRTLTIELRKETSLETLAASIGGLKYLESLTITDLGSEMRTKEAGIVF 752
Query: 729 ILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLN-LPELWKLPS 787
L+ KLY + +FP + L +L L C+ + +P L KL
Sbjct: 753 DFVYLKTLTLKLYMPRLS--KEQHFP--------SHLTTLYLQHCRLEEDPMPILEKLHQ 802
Query: 788 LKYLKLSNMIHVIYLFHESYDGEGLMALKTLF--LEKLPNLIGLS-----REERVMFPRL 840
LK L+L +S+ G+ ++ F L+KL ++ GL + E P L
Sbjct: 803 LKELELR---------RKSFSGKEMVCSSGGFPQLQKL-SIKGLEEWEDWKVEESSMPVL 852
Query: 841 KALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPDG 900
L+I +C L LP ++ LPS + + SL F EE
Sbjct: 853 HTLDIRDCRKLKQLP--------------DEHLPSHLTSI----SLFFCCLEE------- 887
Query: 901 ILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKE 960
P+ TL ++H+ LQ L+ + I +LH LK
Sbjct: 888 ------DPMPTL------------ERLVHLKELQLLFRSFSGRIMVCAGSGFPQLHKLKL 929
Query: 961 LDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLP 1020
++ G ++ + + L TL I C +++ L++L L++L E
Sbjct: 930 SELDGLEEWIVEDG--SMPQLHTLEIRRCPKLKKLPNGFPQ---LQNLELNELEEWEEWI 984
Query: 1021 ECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIV 1080
G++ LLH + I++CPKL LP ++ I L+ L++ + +KR K GED+ K+
Sbjct: 985 VEDGSMPLLHTLRIWNCPKLKQLPDGLRFIYSLKNLTVP--KRWKKRLSKG-GEDYYKVQ 1041
Query: 1081 HVQYIE 1086
H+ +E
Sbjct: 1042 HIPSVE 1047
>RP13_ARATH (Q9M667) Disease resistance protein RPP13 (Resistance to
Peronospora parasitica protein 13)
Length = 835
Score = 219 bits (557), Expect = 4e-56
Identities = 199/748 (26%), Positives = 359/748 (47%), Gaps = 86/748 (11%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M +A+ E V+G + + +E S+F+ ++ L + LT I L+D E ++ D E+
Sbjct: 1 MVDAITEFVVGKIGNYLIEEASMFMAVKEDLEELKTELTCIHGYLKDVEARERED-EVS- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKRIGVW 120
K+W + D AY ++D++D T L++E ++ + GL K+ +KM +
Sbjct: 59 --KEWSKLVLDFAYDVEDVLD---TYHLKLEERSQRRGLRR----LTNKIGRKMDAYSI- 108
Query: 121 LDDIAAEKNKFHLTEIVRERSGV-----VPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVG 175
+DDI K + RE G+ T+S+ + L R+ D++++V L
Sbjct: 109 VDDIRILKRRILDITRKRETYGIGGLKEPQGGGNTSSLRVRQLRRARSVDQEEVVVGLED 168
Query: 176 DAS---------EQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFT 226
DA E+++ + I G+GGLGKT LA+ ++N + FE + W VS+++
Sbjct: 169 DAKILLEKLLDYEEKNRFIISIFGMGGLGKTALARKLYNSRDVKERFEYRAWTYVSQEYK 228
Query: 227 LKRMTKAIIEGATKKSCEDLDL------ELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQ 280
+ II S E+L+ E L+ L LL K+YL+V+DD+W ++E W
Sbjct: 229 TGDILMRIIRSLGMTSGEELEKIRKFAEEELEVYLYGLLEGKKYLVVVDDIW--EREAWD 286
Query: 281 RLKSVLACGGKGASILVTTRLPKVAK-IMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQ 339
LK L C +G+ +++TTR+ VA+ + G H+L L+ E+ WELF+QRAF + +
Sbjct: 287 SLKRALPCNHEGSRVIITTRIKAVAEGVDGRFYAHKLRFLTFEESWELFEQRAFRNIQRK 346
Query: 340 QKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA-YVMP-A 397
++L+ GKE+++KC G PL + L LL ++ EW V S L+ ++ +V P
Sbjct: 347 DEDLLKTGKEMVQKCRGLPLCIVVLAGLLS-RKTPSEWNDVCNSLWRRLKDDSIHVAPIV 405
Query: 398 LRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWN 457
LS+ L + + CF + ++FP+D I + LI L A GFI ++ + +D+
Sbjct: 406 FDLSFKELRHESKLCFLYLSIFPEDYEIDLEKLIHLLVAEGFIQGDEEMMMEDVARYYIE 465
Query: 458 ELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCIT--DDNSMRTMSEETRHL 515
EL RS E G++ ++HDL+ D+A ++++ + +D+ + S R
Sbjct: 466 ELIDRSLLEAVRRER-GKVMSCRIHDLLRDVAIKKSKELNFVNVYNDHVAQHSSTTCRRE 524
Query: 516 LIYNRNSFAEANSIQLHHVKSLKTYMEFNFDV---YEAGQLSPQVLNCYSLRVLLSHRLN 572
+++++ + + ++S + EF+ V +E +L +VL+ SL L ++N
Sbjct: 525 VVHHQFKRYSSEKRKNKRMRSFLYFGEFDHLVGLDFETLKLL-RVLDFGSL--WLPFKIN 581
Query: 573 NLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQ 632
G L +LRYL I + + +++L+ LQ
Sbjct: 582 ------GDLIHLRYLGIDGNSINDF----------------------DIAAIISKLRFLQ 613
Query: 633 NLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGE-ERGFLLEELGQLNLKGQLHIKNLERL 691
L + D + + KLTSL ++++G G L+ ++ L + + +L
Sbjct: 614 TLFVSD-NYFIEETIDLRKLTSL----RHVIGNFFGGLLIGDVANLQTLTSISFDSWNKL 668
Query: 692 K-----SVTDAKKANMSRKKLNQLWLSW 714
K ++ D + MSR K ++ +SW
Sbjct: 669 KPELLINLRDLGISEMSRSKERRVHVSW 696
>R8L4_ARATH (Q9FJK8) Probable disease resistance RPP8-like protein 4
Length = 908
Score = 214 bits (545), Expect = 1e-54
Identities = 267/1008 (26%), Positives = 441/1008 (43%), Gaps = 142/1008 (14%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAE + L L +L+ +E G D++ + L L ++++ L+DA+ K+
Sbjct: 1 MAEGFVSFGLEKLWDLLSRESERLQGIDEQLDGLKRQLRSLQSLLKDADAKKHGSDR--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH--KHIAFRYKLAKKMKRIG 118
V+++L +KD + +DI++ L E K K + + + R+K+A ++ I
Sbjct: 58 -VRNFLEDVKDLVFDAEDIIESYVLNKLRGEGKGVKKHVRRLARFLTDRHKVASDIEGIT 116
Query: 119 VWLDDIAAEKNKFHLTEIV--------RERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIV 170
+ D+ E F + +I+ +ER V + RQT ++ + G + +++V
Sbjct: 117 KRISDVIGEMQSFGIQQIIDGVRSLSLQERQRVQREIRQTYPDSSESDLVGVEQSVEELV 176
Query: 171 DFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRM 230
LV E + V I G+GG+GKTTLA+ VF+HD + HF+ WVCVS+ FTLK +
Sbjct: 177 GHLV----ENDIYQVVSIAGMGGIGKTTLARQVFHHDLVRRHFDGFAWVCVSQQFTLKHV 232
Query: 231 TKAIIEGATKK--SCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLAC 288
+ I++ + +D LQ KL LL RYLLVLDDVW K+E+W R+K+V
Sbjct: 233 WQRILQELQPHDGNILQMDESALQPKLFQLLETGRYLLVLDDVW--KKEDWDRIKAVFP- 289
Query: 289 GGKGASILVTTRLPKVA-KIMGTIPHHELSRLSDEDCWELFKQRAF---GPNEVQ-QKEL 343
+G +L+T+R V T S L+ E+ W+L ++ F EV+ +E+
Sbjct: 290 RKRGWKMLLTSRNEGVGIHADPTCLTFRASILNPEESWKLCERIVFPRRDETEVRLDEEM 349
Query: 344 VIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA--------YVM 395
+GKE++ CGG PLA ALG LL K EW V ++ + G + V
Sbjct: 350 EAMGKEMVTHCGGLPLAVKALGGLLANKHTVPEWKRVSDNIGSQIVGGSCLDDNSLNSVN 409
Query: 396 PALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEV 455
L LSY LP L+ F + A FP+D I Q L + W A G + + D G
Sbjct: 410 RILSLSYEDLPTHLKHRFLYLAHFPEDSKIYTQDLFNYWAAEGIYDGSTI---QDSGEYY 466
Query: 456 WNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHL 515
EL R+ + +MHD++ ++ S + ++N ++ + + T
Sbjct: 467 LEELVRRNLVIADNRYLSLEFNFCQMHDMMREVCLSKAK------EENFLQIIKDPTSTS 520
Query: 516 LIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLS 575
I N S + + +H K+ N +P+V + R + + +
Sbjct: 521 TI-NAQSPSRSRRFSIHSGKAFHILGHRN---------NPKVRSLIVSRFEEDFWIRS-A 569
Query: 576 SSIGRLKYLRYLDISEGRFK--NLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQN 633
S L LR LD+S +F+ LP+S+ L +L L L G V + LP + LK L
Sbjct: 570 SVFHNLTLLRVLDLSRVKFEGGKLPSSIGGLIHLRYLSLYGAV-VSHLPSTMRNLKLLLF 628
Query: 634 LSLR-DCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLK 692
L+LR D +P + ++ L LS +++ L ELG L NLE L
Sbjct: 629 LNLRVDNKEPIHVPNVLKEMLELRYLSLPQEMDDKTKL--ELGDL--------VNLEYL- 677
Query: 693 SVTDAKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAY 752
+ S + + V+ L KL + GV
Sbjct: 678 -----------------WYFSTQHSSVTDLLR------------MTKLRNLGVSLSERCN 708
Query: 753 FPQWISIPSLNDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGL 812
F S SL +L++LE++ + L PE+ + + L + IH+ L GL
Sbjct: 709 FETLSS--SLRELRNLEML---NVLFSPEIVMVDHMGEFVLDHFIHLKQL--------GL 755
Query: 813 MALKTLFLEKLPNLIGLSREERVMFPRLKALEITEC-------PNLLGLPCLPSLSDLYI 865
+ + K+P ++ P L + + C P L L L S++ Y
Sbjct: 756 ----AVRMSKIP-------DQHQFPPHLAHIHLVHCVMKEDPMPILEKLLHLKSVALSY- 803
Query: 866 QGKY-NQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASP-LKTLGFHRHSKLKML 923
G + +++ S L +L S EL + I+ + P L+TL H KLK L
Sbjct: 804 -GAFIGRRVVCSKGGFPQLCALGISGESEL---EEWIVEEGSMPCLRTLTIHDCEKLKEL 859
Query: 924 PTEMIHIHALQQLYINDCRN--IEEL--PNEVMQRLHSLKELDIVGCD 967
P + +I +L++L I + + E+L E ++ + ++ + CD
Sbjct: 860 PDGLKYITSLKELKIREMKREWKEKLVPGGEDYYKVQHIPDVQFINCD 907
Score = 40.0 bits (92), Expect = 0.035
Identities = 74/323 (22%), Positives = 135/323 (40%), Gaps = 75/323 (23%)
Query: 777 LNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREERVM 836
L+LP+ ++ L+L +++++ YL++ S + L L + KL NL G+S ER
Sbjct: 654 LSLPQ--EMDDKTKLELGDLVNLEYLWYFSTQHSSVTDL--LRMTKLRNL-GVSLSERCN 708
Query: 837 FPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELI- 895
F + L SS+ +L +LE L+ + E++
Sbjct: 709 F---------------------------------ETLSSSLRELRNLEMLNVLFSPEIVM 735
Query: 896 --YFPDGILRNLASPLKTLGFH-RHSKLK---MLPTEMIHIHALQQLYINDCRNIEELPN 949
+ + +L + LK LG R SK+ P + HIH + + ++E P
Sbjct: 736 VDHMGEFVLDHFIH-LKQLGLAVRMSKIPDQHQFPPHLAHIHLVHCV-------MKEDPM 787
Query: 950 EVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLT 1009
++++L LK + + Y + + CS+ GF + L +L
Sbjct: 788 PILEKLLHLKSVAL------------SYGAFIGRRVV--CSK-GGFPQ-------LCALG 825
Query: 1010 LSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQ 1069
+S LE G++ L + I+ C KL LP ++ I+ L+ L I + + K
Sbjct: 826 ISGESELEEWIVEEGSMPCLRTLTIHDCEKLKELPDGLKYITSLKELKIREMKREWKEKL 885
Query: 1070 KEIGEDWPKIVHVQYIEIENDNL 1092
GED+ K+ H+ ++ N +L
Sbjct: 886 VPGGEDYYKVQHIPDVQFINCDL 908
>RPP8_ARATH (Q8W4J9) Disease resistance protein RPP8 (Resistance to
Peronospora parasitica protein 8)
Length = 908
Score = 214 bits (544), Expect = 1e-54
Identities = 232/901 (25%), Positives = 406/901 (44%), Gaps = 129/901 (14%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA + L L +L+ +E G D + + L L ++++ L+DA+ K+
Sbjct: 1 MAEAFVSFGLEKLWDLLSRESERLQGIDGQLDGLKRQLRSLQSLLKDADAKKHGSDR--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHI-------AFRYKLAKK 113
V+++L +KD + +DI++ L + K K KH+ R+K+A
Sbjct: 58 -VRNFLEDVKDLVFDAEDIIESYVLNKLSGKGKGVK-----KHVRRLACFLTDRHKVASD 111
Query: 114 MKRIGVWLDDIAAEKNKFHLTEIV--------RERSGVVPDWRQTTSIVTQPLVYGRNED 165
++ I + ++ E F + +I+ +ER V + RQT ++ + G +
Sbjct: 112 IEGITKRISEVIGEMQSFGIQQIIDGGRSLSLQERQRVQREIRQTYPDSSESDLVGVEQS 171
Query: 166 KDKIVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDF 225
++V LV E + V I G+GG+GKTTLA+ VF+HD + HF+ WVCVS+ F
Sbjct: 172 VKELVGHLV----ENDVHQVVSIAGMGGIGKTTLARQVFHHDLVRRHFDGFAWVCVSQQF 227
Query: 226 TLKRMTKAIIEGATKKSCEDLDLE--LLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLK 283
T K + + I++ + L ++ LQRKL LL RYL+VLDDVW K+E+W +K
Sbjct: 228 TQKHVWQRILQELQPHDGDILQMDEYALQRKLFQLLEAGRYLVVLDDVW--KKEDWDVIK 285
Query: 284 SVLACGGKGASILVTTRLPKVA-KIMGTIPHHELSRLSDEDCWELFKQRAF---GPNEVQ 339
+V +G +L+T+R V T S L+ E+ W+L ++ F EV+
Sbjct: 286 AVFP-RKRGWKMLLTSRNEGVGIHADPTCLTFRASILNPEESWKLCERIVFPRRDETEVR 344
Query: 340 -QKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEAY----- 393
+E+ +GKE++ CGG PLA ALG LL K EW V ++ + G ++
Sbjct: 345 LDEEMEAMGKEMVTHCGGLPLAVKALGGLLANKHTVPEWKRVFDNIGSQIVGGSWLDDNS 404
Query: 394 ---VMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADD 450
V L LSY LP L+ CF A FP+D IS L W A G + + +D
Sbjct: 405 LNSVYRILSLSYEDLPTHLKHCFLNLAHFPEDSEISTYSLFYYWAAEGIYDGSTI---ED 461
Query: 451 IGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQD---VCCITDDNSMRT 507
G EL R+ +N Q +MHD++ ++ S ++ + I D T
Sbjct: 462 SGEYYLEELVRRNLVIADDNYLSWQSKYCQMHDMMREVCLSKAKEENFLQIIIDPTCTST 521
Query: 508 MSEE----TRHLLIYNRNSF----------------------AEANSIQLHHVKSLKTYM 541
++ + +R L I++ +F S + H +L +
Sbjct: 522 INAQSPSRSRRLSIHSGKAFHILGHKNKTKVRSLIVPRFEEDYWIRSASVFHNLTLLRVL 581
Query: 542 EFNFDVYEAGQLSPQVLNCYSLRVLLSH--RLNNLSSSIGRLKYLRYLDISEGRFK--NL 597
+ ++ +E G+L + LR L + ++++L S++ LK L YL++ + ++
Sbjct: 582 DLSWVKFEGGKLPCSIGGLIHLRYLSLYEAKVSHLPSTMRNLKLLLYLNLRVDTEEPIHV 641
Query: 598 PNSLCKLCNLEV------------LKLDGCVSLQKLPG---------GLTRLKRLQNLSL 636
PN L ++ L L+L V+L+ L G L R+ +L+ L++
Sbjct: 642 PNVLKEMIQLRYLSLPLKMDDKTKLELGDLVNLEYLYGFSTQHSSVTDLLRMTKLRYLAV 701
Query: 637 R-----DCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNL--- 688
+ ++L+S R++ L +LN L ++++ +G+ L +H+K L
Sbjct: 702 SLSERCNFETLSSSLRELRNLETLNFLFSL-----ETYMVDYMGEFVLDHFIHLKQLGLA 756
Query: 689 ERLKSVTDAKKANMSRKKLNQLWLSWERNEVSQLQE--NVEQILEALQPYAQKLYSFGVG 746
R+ + D + L ++ E + + L++ +++ + A + + G
Sbjct: 757 VRMSKIPDQHQFPPHLVHLFLIYCGMEEDPMPILEKLLHLKSVRLARKAFLGSRMVCSKG 816
Query: 747 GY---------TGAYFPQWI-SIPSLNDLKSLELVDCKSCLNLPE-LWKLPSLKYLKLSN 795
G+ + +WI S+ L++L + DCK LP+ L + SLK LK+
Sbjct: 817 GFPQLCVIEISKESELEEWIVEEGSMPCLRTLTIDDCKKLKELPDGLKYITSLKELKIEG 876
Query: 796 M 796
M
Sbjct: 877 M 877
Score = 39.3 bits (90), Expect = 0.060
Identities = 57/231 (24%), Positives = 97/231 (41%), Gaps = 47/231 (20%)
Query: 871 QQLPSSIHKLGSLESLHFSDNEELI---YFPDGILRNLASPLKTLGFH-RHSKLK---ML 923
+ L SS+ +L +LE+L+F + E Y + +L + LK LG R SK+
Sbjct: 710 ETLSSSLRELRNLETLNFLFSLETYMVDYMGEFVLDHFIH-LKQLGLAVRMSKIPDQHQF 768
Query: 924 PTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDI-----VGCDKLKLSSDFQYL 978
P ++H L++ C +EE P ++++L LK + + +G + F L
Sbjct: 769 PPHLVH------LFLIYC-GMEEDPMPILEKLLHLKSVRLARKAFLGSRMVCSKGGFPQL 821
Query: 979 TCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCP 1038
+E I SE+E + M L++LT+ D C
Sbjct: 822 CVIE---ISKESELEEWIVEEGSMPCLRTLTIDD------------------------CK 854
Query: 1039 KLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEIEN 1089
KL LP ++ I+ L+ L I + K GED+ K+ H+ ++ N
Sbjct: 855 KLKELPDGLKYITSLKELKIEGMKREWKEKLVPGGEDYYKVQHIPDVQFIN 905
>R8L2_ARATH (Q9MAG6) Probable disease resistance RPP8-like protein 2
Length = 1271
Score = 211 bits (537), Expect = 9e-54
Identities = 204/702 (29%), Positives = 332/702 (47%), Gaps = 65/702 (9%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEAV+ + L EL+ +E + G D++ + L L +++ L+DA+ K+ +E R
Sbjct: 1 MAEAVVSFGVEKLWELLSRESARLNGIDEQVDGLKRQLGRLQSLLKDADAKK---NETER 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHK--HIAFRYKLAKKMKRIG 118
V+++L +KD Y DDI++ L + K K + + R K A ++ I
Sbjct: 58 -VRNFLEDVKDIVYDADDIIESFLLNELRGKEKGIKKQVRTLACFLVDRRKFASDIEGIT 116
Query: 119 VWLDDIAAEKNKFHLTEI---------VRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKI 169
+ ++ + I ++ER + RQT S ++ + G ++ +++
Sbjct: 117 KRISEVIVGMQSLGIQHIADGGGRSLSLQERQREI---RQTFSRNSESDLVGLDQSVEEL 173
Query: 170 VDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKR 229
VD LV E + + V + G+GG+GKTTLA+ VF+HD + HF+ WVCVS+ FT K
Sbjct: 174 VDHLV----ENDSVQVVSVSGMGGIGKTTLARQVFHHDIVRRHFDGFSWVCVSQQFTRKD 229
Query: 230 MTKAIIEGAT--KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLA 287
+ + I++ + +D LQ +L +LL RYLLVLDDVW K+E+W R+K+V
Sbjct: 230 VWQRILQDLRPYDEGIIQMDEYTLQGELFELLESGRYLLVLDDVW--KEEDWDRIKAVFP 287
Query: 288 CGGKGASILVTTRLPKVAKIMGTIPHHELSR-LSDEDCWELFKQRAFG-PNEVQQKELVI 345
+G +L+T+R + R L+ E W+LF++ ++ + K
Sbjct: 288 -HKRGWKMLLTSRNEGLGLHADPTCFAFRPRILTPEQSWKLFERIVSSRRDKTEFKVDEA 346
Query: 346 VGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA--------YVMPA 397
+GKE++ CGG PLA LG LL K EW V + + ++ G++ V
Sbjct: 347 MGKEMVTYCGGLPLAVKVLGGLLAKKHTVLEWKRVHSNIVTHIVGKSGLSDDNSNSVYRV 406
Query: 398 LRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISS-NQMLEADDIGNEVW 456
L LSY LP++L+ CF + A FP+D I ++L + W A G I+ + D G
Sbjct: 407 LSLSYEDLPMQLKHCFFYLAHFPEDYKIDVKILFNYWVAEGIITPFHDGSTIQDTGESYL 466
Query: 457 NELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLL 516
EL R+ E+ +I +MHD++ ++ S + ++N +R + T
Sbjct: 467 EELVRRNMVVVEESYLTSRIEYCQMHDMMREVCLSKAK------EENFIRVVKVPTTTST 520
Query: 517 IYNRNSFAEANSIQLH---------HVKSLKTYMEFNFDVYEAGQLSPQVLNCYS-LRVL 566
N S + + LH H + K F V E P+ C LRVL
Sbjct: 521 TINAQSPCRSRRLVLHSGNALHMLGHKDNKKARSVLIFGV-EEKFWKPRGFQCLPLLRVL 579
Query: 567 ----LSHRLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVS--LQK 620
+ L SSIG L +LR+L + E +LP+SL L L L L G L
Sbjct: 580 DLSYVQFEGGKLPSSIGDLIHLRFLSLYEAGVSHLPSSLGNLKLLLCLNL-GVADRLLVH 638
Query: 621 LPGGLTRLKRLQNLSL-RDCDSLTSLPRQIGKLTSLNTLSKY 661
+P L ++ L+ L L R + T L ++G L +L +L+ +
Sbjct: 639 VPNVLKEMQELRYLRLPRSMPAKTKL--ELGDLVNLESLTNF 678
>R8L3_ARATH (Q9FJB5) Probable disease resistance RPP8-like protein 3
Length = 901
Score = 207 bits (528), Expect = 1e-52
Identities = 243/937 (25%), Positives = 413/937 (43%), Gaps = 125/937 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAE V+ + L L+ +E G D++ + L L +++ L+DA+ K+
Sbjct: 1 MAEGVVSFGVQKLWALLNRESERLNGIDEQVDGLKRQLRGLQSLLKDADAKKHGSDR--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASK-------CGLSHKHIAFRYKLAKK 113
V+++L +KD + +DI++ L E K K C L+ +H K+A
Sbjct: 58 -VRNFLEDVKDLVFDAEDIIESYVLNKLRGEGKGVKNHVRRLACFLTDRH-----KVASD 111
Query: 114 MKRIGVWLDDIAAEKNKFHLTEIVRERS------GVVPDWRQTTSIVTQPLVYGRNEDKD 167
++ I + + E + + + + + + RQT ++ + G +
Sbjct: 112 IEGITKRISKVIGEMQSLGIQQQIIDGGRSLSLQDIQREIRQTFPNSSESDLVGVEQS-- 169
Query: 168 KIVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTL 227
V+ LVG E +++ V I G+GG+GKTTLA+ +F+HD + HF+ WVCVS+ FT
Sbjct: 170 --VEELVGPMVEIDNIQVVSISGMGGIGKTTLARQIFHHDLVRRHFDGFAWVCVSQQFTQ 227
Query: 228 KRMTKAIIEGATKKSCEDLDLE--LLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSV 285
K + + I++ E L ++ +Q KL LL RYL+VLDDVW K+E+W R+K V
Sbjct: 228 KHVWQRILQELRPHDGEILQMDEYTIQGKLFQLLETGRYLVVLDDVW--KEEDWDRIKEV 285
Query: 286 LACGGKGASILVTTRLPKVA-KIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELV 344
+G +L+T+R V T L+ ++ W+LF++ NE + +E+
Sbjct: 286 FP-RKRGWKMLLTSRNEGVGLHADPTCLSFRARILNPKESWKLFERIVPRRNETEYEEME 344
Query: 345 IVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA--------YVMP 396
+GKE++ CGG PLA LG LL K EW V E+ + G++ V
Sbjct: 345 AIGKEMVTYCGGLPLAVKVLGGLLANKHTASEWKRVSENIGAQIVGKSCLDDNSLNSVYR 404
Query: 397 ALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVW 456
L LSY LP L+ CF + A FP+D I + L W A G +L D G +
Sbjct: 405 ILSLSYEDLPTDLKHCFLYLAHFPEDYKIKTRTLYSYWAAEGIYDGLTIL---DSGEDYL 461
Query: 457 NELYWRSFFENTENVGFGQITIFKMHDLVHDLAGS------VTQDVCCITDDNSMRTMS- 509
EL R+ ++ ++ + +MHD++ ++ S Q + T +++ S
Sbjct: 462 EELVRRNLVIAEKSNLSWRLKLCQMHDMMREVCISKAKVENFLQIIKVPTSTSTIIAQSP 521
Query: 510 EETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVL--- 566
+R L +++ +F L H K +++ + Q + + + LRVL
Sbjct: 522 SRSRRLTVHSGKAFH-----ILGHKKKVRSLLVLGLKEDLWIQSASRFQSLPLLRVLDLS 576
Query: 567 -LSHRLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQ-KLPGG 624
+ L SSIG L +LR+L + + +LP+++ L + L L + + +P
Sbjct: 577 SVKFEGGKLPSSIGGLIHLRFLSLHQAVVSHLPSTIRNLKLMLYLNLHVAIGVPVHVPNV 636
Query: 625 LTRLKRLQNLSL-RDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQL 683
L + L+ LSL D T L ++G L +L L +
Sbjct: 637 LKEMLELRYLSLPLDMHDKTKL--ELGDLVNLEYLWCFST-------------------- 674
Query: 684 HIKNLERLKSVTDAKKANMSRKKLNQLWLSW-ERNEVSQLQENVEQI--LEALQ-PYAQK 739
+ SVTD + KL +S+ ER L ++ Q LE L Y++K
Sbjct: 675 ------QHSSVTDL----LRMTKLRFFGVSFSERCTFENLSSSLRQFRKLETLSFIYSRK 724
Query: 740 LYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHV 799
Y + Y G + +I + L+ L +P+ +LP ++ H+
Sbjct: 725 TY---MVDYVGEFVLDFIHLKKLSLGVHLS--------KIPDQHQLP-------PHIAHI 766
Query: 800 IYLF-HESYDG----EGLMALKTLFLEKLPNLIGLSREERVMFPRLKALEITECPNL--- 851
LF H D E L+ LK++ L + + + FP+L+AL+I+E L
Sbjct: 767 YLLFCHMEEDPMPILEKLLHLKSVELRRKAFIGRRMVCSKGGFPQLRALQISEQSELEEW 826
Query: 852 -LGLPCLPSLSDLYIQG-KYNQQLPSSIHKLGSLESL 886
+ +P L DL I + ++LP + + SL+ L
Sbjct: 827 IVEEGSMPCLRDLIIHSCEKLEELPDGLKYVTSLKEL 863
Score = 43.9 bits (102), Expect = 0.002
Identities = 56/230 (24%), Positives = 102/230 (44%), Gaps = 48/230 (20%)
Query: 871 QQLPSSIHKLGSLESLHFSDNEELI---YFPDGILRNLASPLKTLGFHRHSKLK---MLP 924
+ L SS+ + LE+L F + + Y + +L + +LG H SK+ LP
Sbjct: 702 ENLSSSLRQFRKLETLSFIYSRKTYMVDYVGEFVLDFIHLKKLSLGVHL-SKIPDQHQLP 760
Query: 925 TEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDI-----VGCDKLKLSSDFQYLT 979
+ HI Y+ C ++EE P ++++L LK +++ +G + F L
Sbjct: 761 PHIAHI------YLLFC-HMEEDPMPILEKLLHLKSVELRRKAFIGRRMVCSKGGFPQLR 813
Query: 980 CLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPK 1039
L+ I SE+E E++ E G++ L ++ I+SC K
Sbjct: 814 ALQ---ISEQSELE-----------------------EWIVE-EGSMPCLRDLIIHSCEK 846
Query: 1040 LACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEIEN 1089
L LP ++ ++ L+ L I + K +K +GED+ K+ H+ ++ N
Sbjct: 847 LEELPDGLKYVTSLKELKIEGMKREWK--EKLVGEDYYKVQHIPDVQFFN 894
>DR13_ARATH (Q9XIF0) Putative disease resistance protein At1g59780
Length = 906
Score = 204 bits (518), Expect = 1e-51
Identities = 240/924 (25%), Positives = 414/924 (43%), Gaps = 129/924 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M ++++ + L +L+ +E F G +++ L L + A L DA+ K+ + +
Sbjct: 6 MVDSIVSFGVEKLWKLLSQEYERFQGVEEQITELRDDLKMLMAFLSDADAKKQTRAL--- 62
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATE-ALEMEYKASKCGLSHKHIAFRY--------KLA 111
++ L ++K+ Y +DI++ + ++ M A G + IA + K+
Sbjct: 63 -ARNCLEEIKEITYDAEDIIEIFLLKGSVNMRSLACFPG-GRREIALQITSISKRISKVI 120
Query: 112 KKMKRIGVWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVD 171
+ M+ +G+ D + + ++ R+R + R T S ++ + G ++ +K+V+
Sbjct: 121 QVMQNLGIKSDIMDGVDSH---AQLERKR-----ELRHTFSSESESNLVGLEKNVEKLVE 172
Query: 172 FLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMT 231
LVG+ S I GLGGLGKTTLA+ +F+HDK+ +HF+ WVCVS++FT K +
Sbjct: 173 ELVGNDSSHG----VSITGLGGLGKTTLARQIFDHDKVKSHFDGLAWVCVSQEFTRKDVW 228
Query: 232 KAIIEGATKK-SCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGG 290
K I+ + K DL + +Q+KL LL K+ L+V DD+W K+E+W R+ +
Sbjct: 229 KTILGNLSPKYKDSDLPEDDIQKKLFQLLETKKALIVFDDLW--KREDWYRIAPMFPERK 286
Query: 291 KGASILVTTRLPKVAKIMGTIPH---HELSRLSDEDCWELFKQRAFGPNE-----VQQKE 342
G +L+T+R + PH + L+ ++CW+L ++ AF + + KE
Sbjct: 287 AGWKVLLTSRNDAIH------PHCVTFKPELLTHDECWKLLQRIAFSKQKTITGYIIDKE 340
Query: 343 LVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKL---------WNLQGEAY 393
+V + KE+ K C PLA LG LL K ++W + E+ + N +
Sbjct: 341 MVKMAKEMTKHCKRLPLAVKLLGGLLDAKHTLRQWKLISENIISHIVVGGTSSNENDSSS 400
Query: 394 VMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANG--FISSNQMLEADDI 451
V L LS+ LP L+ C + A +P+D I + L +W A G + + + D+
Sbjct: 401 VNHVLSLSFEGLPGYLKHCLLYLASYPEDHEIEIERLSYVWAAEGITYPGNYEGATIRDV 460
Query: 452 GNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLA------GSVTQDVCCITDDNSM 505
+ EL R+ + + + ++HDL+ ++ + Q V T +S+
Sbjct: 461 ADLYIEELVKRNMVISERDALTSRFEKCQLHDLMREICLLKAKEENFLQIVTDPTSSSSV 520
Query: 506 RTM-SEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLR 564
++ S +R L++YN + F+ N ++ ++SL L
Sbjct: 521 HSLASSRSRRLVVYNTSIFSGENDMKNSKLRSL-------------------------LF 555
Query: 565 VLLSHRLNNLSSSIGRLKYLRYLDISEGRFK--NLPNSLCKLCNLEVLKLDGCVSLQKLP 622
+ + + ++ S+ L LR LD+ +FK LP+S+ KL +L+ L L S+ LP
Sbjct: 556 IPVGYSRFSMGSNFIELPLLRVLDLDGAKFKGGKLPSSIGKLIHLKYLSLYQ-ASVTYLP 614
Query: 623 GGLTRLKRLQNLSLR-DCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKG 681
L LK L L+LR + L ++P ++ L LS + E ELG L LK
Sbjct: 615 SSLRNLKSLLYLNLRINSGQLINVPNVFKEMLELRYLS--LPWERSSLTKLELGNL-LKL 671
Query: 682 QLHIKNLERLKSVTDAKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQPYAQKLY 741
+ I + SVTD + M++ + Q+ +S E + L + + + + L
Sbjct: 672 ETLINFSTKDSSVTDLHR--MTKLRTLQILISGEGLHMETLSSALSML-----GHLEDLT 724
Query: 742 SFGVGGYTGAYFPQWISIPSLND-------LKSLELVDCKSCLN-LPELWKLPSLKYLKL 793
P+ I P L D L ++ LV C + +P L KL LK
Sbjct: 725 VTPSENSVQFKHPKLIYRPMLPDVQHFPSHLTTISLVYCFLEEDPMPTLEKLLQLK---- 780
Query: 794 SNMIHVIYLFHESY-------DGEGLMALKTLFLEKLPNLIGLSREERVMFPRLKALEIT 846
V+ L++ +Y G G L L + L L EE M P L L I
Sbjct: 781 -----VVSLWYNAYVGRRMVCTGGGFPPLHRLEIWGLDALEEWIVEEGSM-PLLHTLHIV 834
Query: 847 ECPNLL----GLPCLPSLSDLYIQ 866
+C L GL + SL +L I+
Sbjct: 835 DCKKLKEIPDGLRFISSLKELAIR 858
Score = 47.8 bits (112), Expect = 2e-04
Identities = 85/353 (24%), Positives = 145/353 (40%), Gaps = 88/353 (24%)
Query: 761 SLNDLKSLELVDCK----SCLNLPELWK-LPSLKYLKLSNMIHVIYLFHESYDGEGLMAL 815
SL +LKSL ++ + +N+P ++K + L+YL L ++ L L
Sbjct: 616 SLRNLKSLLYLNLRINSGQLINVPNVFKEMLELRYLSLP------------WERSSLTKL 663
Query: 816 KTLFLEKLPNLIGLSREERVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPS 875
+ L KL LI S ++ +T+ + L L L + +G + + L S
Sbjct: 664 ELGNLLKLETLINFSTKDS---------SVTDLHRMTKLRTLQIL--ISGEGLHMETLSS 712
Query: 876 SIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKL---KMLPTEMIHIHA 932
++ LG LE L + +E + F +H KL MLP
Sbjct: 713 ALSMLGHLEDLTVTPSENSVQF------------------KHPKLIYRPMLPDVQHFPSH 754
Query: 933 LQQLYINDCRNIEELPNEVMQRLHSLKELDI-----VGCDKLKLSSDFQYLTCLETLAIG 987
L + + C +EE P +++L LK + + VG + F L LE +
Sbjct: 755 LTTISLVYCF-LEEDPMPTLEKLLQLKVVSLWYNAYVGRRMVCTGGGFPPLHRLEIWGL- 812
Query: 988 SCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSI 1047
+AL+ E++ E G++ LLH ++I C KL +P +
Sbjct: 813 ---------DALE----------------EWIVE-EGSMPLLHTLHIVDCKKLKEIPDGL 846
Query: 1048 QQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYI------EIENDNLIH 1094
+ IS L+ L+I K+ ++ + GED+ K+ HV I E EN+ +I+
Sbjct: 847 RFISSLKELAIRTNEKVFQKKVSKGGEDYYKMQHVPLIRYNWPQEPENNEVIY 899
>R8L1_ARATH (O04093) Putative disease resistance protein RPP8-like
protein 1
Length = 821
Score = 199 bits (505), Expect = 5e-50
Identities = 206/735 (28%), Positives = 337/735 (45%), Gaps = 83/735 (11%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAE V+ + L EL+ +E + G ++ + L L +++ L+DA+ K+
Sbjct: 1 MAEGVVLFGVHKLWELLNRESARLNGIGEQVDGLKRQLGRLQSLLKDADAKKHESER--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKR-IGV 119
V+++L ++D Y +DI++ L E++ + G+ K A+++ + +
Sbjct: 58 -VRNFLEDVRDIVYDAEDIIESF----LLNEFRTKEKGIK--------KHARRLACFLSL 104
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
+ +I + L E RE+ + RQT + ++ + G + V+ L G E
Sbjct: 105 GIQEIIDGASSMSLQERQREQKEI----RQTFANSSESDLVGVEQS----VEALAGHLVE 156
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+++ V I G+GG+GKTTLA+ VF+HD + HF+ WV VS+ FT K + + I +
Sbjct: 157 NDNIQVVSISGMGGIGKTTLARQVFHHDMVQRHFDGFAWVFVSQQFTQKHVWQRIWQELQ 216
Query: 240 KKS--CEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILV 297
++ +D +LQ KL LL RYL+VLDDVW K+E+W R+K+V +G +L+
Sbjct: 217 PQNGDISHMDEHILQGKLFKLLETGRYLVVLDDVW--KEEDWDRIKAVFP-RKRGWKMLL 273
Query: 298 TTRLPKVA-----KIMGTIPHHELSRLSDEDCWELFKQRAFGPNEV--QQKELVIVGKEI 350
T+R V K G + L+ E+ W+L ++ F + ++ +GKE+
Sbjct: 274 TSRNEGVGIHADPKSFG----FKTRILTPEESWKLCEKIVFHRRDETGTLSDMEAMGKEM 329
Query: 351 IKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA-------YVMPALRLSYL 403
+ CGG PLA LG LL K EW V ++ +L G + + L LSY
Sbjct: 330 VTCCGGLPLAVKVLGGLLATKHTVPEWKRVYDNIGPHLAGRSSLDDNLNSIYRVLSLSYE 389
Query: 404 HLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFI-SSNQMLEADDIGNEVWNELYWR 462
+LP+ L+ CF + A FP+ I + L + A G I SS+ D G + EL R
Sbjct: 390 NLPMCLKHCFLYLAHFPEYYEIHVKRLFNYLAAEGIITSSDDGTTIQDKGEDYLEELARR 449
Query: 463 SFFENTENVGFGQITIFKMHDLVHDLAGSVTQD--------VCCITDDNSMRTMSEETRH 514
+ +N F + +MHD++ ++ S ++ V T + R++S ++R
Sbjct: 450 NMITIDKNYMFLRKKHCQMHDMMREVCLSKAKEENFLEIFKVSTATSAINARSLS-KSRR 508
Query: 515 LLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRL--- 571
L ++ N+ V+SL Y F + +P + LRVL R+
Sbjct: 509 LSVHGGNALPSLGQTINKKVRSL-LYFAFEDEFCILESTTPCFRSLPLLRVLDLSRVKFE 567
Query: 572 -NNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKL--DGCVSLQKLPGGLTRL 628
L SSIG L +LR+L + +LP+SL L L L L +G V + + + L
Sbjct: 568 GGKLPSSIGDLIHLRFLSLHRAWISHLPSSLRNLKLLLYLNLGFNGMVHVPNVLKEMQEL 627
Query: 629 KRLQ---------NLSLRDCDSLTSLPRQIGK---------LTSLNTLSKYIVGEERGFL 670
+ LQ L L D +L SL K +T L LS +I L
Sbjct: 628 RYLQLPMSMHDKTKLELSDLVNLESLMNFSTKYASVMDLLHMTKLRELSLFITDGSSDTL 687
Query: 671 LEELGQLNLKGQLHI 685
LGQL LH+
Sbjct: 688 SSSLGQLRSLEVLHL 702
Score = 35.8 bits (81), Expect = 0.66
Identities = 52/184 (28%), Positives = 81/184 (43%), Gaps = 12/184 (6%)
Query: 782 LWKLPSLKYLKL--SNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREERVMFPR 839
L L L YL L + M+HV + E + L ++ + L L E +M
Sbjct: 598 LRNLKLLLYLNLGFNGMVHVPNVLKEMQELRYLQLPMSMHDKTKLELSDLVNLESLMNFS 657
Query: 840 LKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEE--LIYF 897
K + + LL + L LS L+I + L SS+ +L SLE LH D +E + Y
Sbjct: 658 TKYASVMD---LLHMTKLRELS-LFITDGSSDTLSSSLGQLRSLEVLHLYDRQEPRVAYH 713
Query: 898 PDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHS 957
I+ N LK L H + P + + L +Y+ C ++EE P +++RL
Sbjct: 714 GGEIVLNCIH-LKELELAIH--MPRFPDQYLFHPHLSHIYL-WCCSMEEDPIPILERLLH 769
Query: 958 LKEL 961
LK +
Sbjct: 770 LKSV 773
>R132_ARATH (Q9STE5) Putative disease resistance RPP13-like protein
2
Length = 847
Score = 194 bits (492), Expect = 1e-48
Identities = 184/715 (25%), Positives = 335/715 (46%), Gaps = 55/715 (7%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M +A+ E V+G + + +E + +G + L + LT I+ L++ E D E+
Sbjct: 1 MVDAITEFVVGKIDNYLIEEAPMLIGVKDDLEELKTELTCIQVYLKNVEVCDKED-EVS- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGL-------SHKHIAFRYKLAKK 113
K+W + D AY ++D++D T L++E + + GL S K A Y +
Sbjct: 59 --KEWTKLVLDIAYDVEDVLD---TYFLKLEKRLHRLGLMRLTNIISDKKDA--YNILDD 111
Query: 114 MKRIGVWLDDIAAEKNKFHLTEIVRER----SGVVPDWRQTTSIVTQPLVYGRNEDKDKI 169
+K + D+ + + + R + V + R+ S + V G +D +
Sbjct: 112 IKTLKRRTLDVTRKLEMYGIGNFNEHRVVASTSRVREVRRARSDDQEERVVGLTDDAKVL 171
Query: 170 VDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKR 229
+ L+ D + + + + I G+ GLGKT+LA+ +FN + FE ++W VS + +
Sbjct: 172 LTKLLDDDGDNK-IYMISIFGMEGLGKTSLARKLFNSSDVKESFEYRVWTNVSGECNTRD 230
Query: 230 MTKAII---EGATKKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVL 286
+ II E ++ E + + L+ L D+L+ KRYL+V+DD+W + E + LK L
Sbjct: 231 ILMRIISSLEETSEGELEKMAQQELEVYLHDILQEKRYLVVVDDIW--ESEALESLKRAL 288
Query: 287 ACGGKGASILVTTRLPKVAKIMGT-IPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVI 345
C +G+ +++TT + VA+ + H + L+ ++ W LF+++AF +EL
Sbjct: 289 PCSYQGSRVIITTSIRVVAEGRDKRVYTHNIRFLTFKESWNLFEKKAFRYILKVDQELQK 348
Query: 346 VGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEAYVMPALRLSYLHL 405
+GKE+++KCGG P + L L+ +++ EW V S L +V LS+ +
Sbjct: 349 IGKEMVQKCGGLPRTTVVLAGLMS-RKKPNEWNDV-WSSLRVKDDNIHVSSLFDLSFKDM 406
Query: 406 PVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFF 465
+L+ CF + ++FP+D + + LI L A GFI ++ + +D+ +L + S
Sbjct: 407 GHELKLCFLYLSVFPEDYEVDVEKLIQLLVAEGFIQEDEEMTMEDVARYYIEDLVYISLV 466
Query: 466 ENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCIT--DDNSMRTMS--EETRHLLIYNRN 521
E + G++ F++HDLV + ++++ + D+ T S E HL+ N
Sbjct: 467 EVVKRKK-GKLMSFRIHDLVREFTIKKSKELNFVNVYDEQHSSTTSRREVVHHLMDDNYL 525
Query: 522 SFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLN--------- 572
N+ +++++ F + + L LRVL L+
Sbjct: 526 CDRRVNT-------QMRSFLFFGKRRNDITYVETITLKLKLLRVLNLGGLHFICQGYSPW 578
Query: 573 NLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQ 632
+L IG L +LRYL I++ NLP+ + L L+ L G S +++ L+ L L+
Sbjct: 579 SLPDVIGGLVHLRYLGIADTVVNNLPDFISNLRFLQTLDASG-NSFERMT-DLSNLTSLR 636
Query: 633 NLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKN 687
+L+ R L L L +L ++S Y + + LL L L + + HI N
Sbjct: 637 HLTGRFIGEL--LIGDAVNLQTLRSISSYSWSKLKHELLINLRDLEIY-EFHILN 688
>DRL8_ARATH (Q8W3K3) Putative disease resistance protein At1g58400
Length = 910
Score = 192 bits (489), Expect = 3e-48
Identities = 180/720 (25%), Positives = 324/720 (45%), Gaps = 59/720 (8%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M EA++ + L + + +E F G + L S L +K+ L+DAE K+ + +
Sbjct: 1 MVEAIVSFGVEKLWDRLTQEYEQFQGVEDRIAELKSNLNLLKSFLKDAEAKKNTSQMVRH 60
Query: 61 DVKDWLLKLKDAAYTLDD-----IMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMK 115
V++ +K+ Y ++ I+ E A + + + +K H R++ A +
Sbjct: 61 CVEE----IKEIVYDTENMIETFILKEAARKRSGIIRRITKLTCIKVH---RWEFASDIG 113
Query: 116 RIGVWLDDIAAEKNKFHLTEIVRERSGVVP-------DWRQTTSIVTQPLVYGRNEDKDK 168
I + + + + F + +++ + S + RQT S + G + K
Sbjct: 114 GISKRISKVIQDMHSFGVQQMISDGSQSSHLLQEREREMRQTFSRGYESDFVGLEVNVKK 173
Query: 169 IVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLK 228
+V +LV E++D+ + + G+GGLGKTTLA+ VFNH+ + + F+ WVCVS++FT K
Sbjct: 174 LVGYLV----EEDDIQIVSVTGMGGLGKTTLARQVFNHEDVKHQFDRLAWVCVSQEFTRK 229
Query: 229 RMTKAIIEGATKKSCEDLDLELLQRKLQD----LLRRKRYLLVLDDVWNDKQENWQRLKS 284
+ + I++ T + +D L++ + +L D LL + L+V DD+W K+E+W +
Sbjct: 230 NVWQMILQNLTSRETKDEILQMEEAELHDELFQLLETSKSLIVFDDIW--KEEDWGLINP 287
Query: 285 VLACGGKGASILVTTRLPKVAKIMG-TIPHHELSRLSDEDCWELFKQRAFGPNEVQQ--- 340
+ KG +L+T+R +A + + L+ + W LF++ A + +
Sbjct: 288 IFP-PKKGWKVLITSRTETIAMHGNRRYVNFKPECLTILESWILFQRIAMPRVDESEFKV 346
Query: 341 -KELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQG--------E 391
KE+ ++GK++IK CGG PLA LG LL K +W + E+ ++ G
Sbjct: 347 DKEMEMMGKQMIKYCGGLPLAVKVLGGLLAAKYTFHDWKRLSENIGCHIVGRTDFSDGNN 406
Query: 392 AYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQM--LEAD 449
+ V L LS+ LP L+ CF + A FP+D I + L W A G +
Sbjct: 407 SSVYHVLSLSFEELPSYLKHCFLYLAHFPEDHNIKVEKLSYCWAAEGILEPRHYHGQTIR 466
Query: 450 DIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQD-----VCCITDDNS 504
D+G EL R+ +V + +HD++ ++ ++ + I +
Sbjct: 467 DVGESYIEELVRRNMVIAERDVTTLRFEACHLHDMMREVCLLKAKEENFVQIASILPPTA 526
Query: 505 MRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLR 564
+R + N + + I ++SL E ++ L + LR
Sbjct: 527 NSQYPGTSRRFVSQNPTTLHVSRDINNPKLQSLLIVWENRRKSWKL--LGSSFIRLELLR 584
Query: 565 VLLSHRL----NNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQK 620
VL ++ NL S IG+L +LRYL++ R LP+SL L L L ++ C
Sbjct: 585 VLDLYKAKFEGRNLPSGIGKLIHLRYLNLDLARVSRLPSSLGNLRLLIYLDINVCTKSLF 644
Query: 621 LPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLK 680
+P L + L+ L L ++ + + L +L TL + E L + G ++L+
Sbjct: 645 VPNCLMGMHELRYLRL-PFNTSKEIKLGLCNLVNLETLENF--STENSSLEDLRGMVSLR 701
Score = 51.6 bits (122), Expect = 1e-05
Identities = 101/423 (23%), Positives = 177/423 (40%), Gaps = 81/423 (19%)
Query: 723 QENVEQILEALQPYAQKLYS-----FGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCL 777
+EN QI L P A Y F T + + I+ P L L +
Sbjct: 512 EENFVQIASILPPTANSQYPGTSRRFVSQNPTTLHVSRDINNPKLQSLLIV-------WE 564
Query: 778 NLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMA-------LKTLFLE-----KLPN 825
N + WKL +++L ++ V+ L+ ++G L + L+ L L+ +LP+
Sbjct: 565 NRRKSWKLLGSSFIRLE-LLRVLDLYKAKFEGRNLPSGIGKLIHLRYLNLDLARVSRLPS 623
Query: 826 LIGLSREERVMFPRLKALEITECPNLLGLP-CLPSLSDL-YIQGKYN--QQLPSSIHKLG 881
+G R L L+I C L +P CL + +L Y++ +N +++ + L
Sbjct: 624 SLGNLR-------LLIYLDINVCTKSLFVPNCLMGMHELRYLRLPFNTSKEIKLGLCNLV 676
Query: 882 SLESL-HFSDNEELIYFPDGI--LRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYI 938
+LE+L +FS + G+ LR L T+G +H + L ++ + L+ L I
Sbjct: 677 NLETLENFSTENSSLEDLRGMVSLRTL-----TIGLFKHISKETLFASILGMRHLENLSI 731
Query: 939 ------NDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEV 992
+ + I E V+ +H LK+L++ KL + + + L ++++ C V
Sbjct: 732 RTPDGSSKFKRIME-DGIVLDAIH-LKQLNL-RLYMPKLPDEQHFPSHLTSISLDGCCLV 788
Query: 993 EGFHEALQHMTTLKSLTLS--------------DLPNLEYL--------PECI---GNLT 1027
E L+ + LK + L P L L E I G++
Sbjct: 789 EDPLPILEKLLELKEVRLDFRAFCGKRMVSSDGGFPQLHRLYIWGLAEWEEWIVEEGSMP 848
Query: 1028 LLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEI 1087
LH + I++C KL LP ++ I ++ L D K K E GE++ K+ H+ ++
Sbjct: 849 RLHTLTIWNCQKLKQLPDGLRFIYSIKDL---DMDKKWKEILSEGGEEYYKVQHIPSVKF 905
Query: 1088 END 1090
E D
Sbjct: 906 EKD 908
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.138 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 131,491,123
Number of Sequences: 164201
Number of extensions: 5734705
Number of successful extensions: 17097
Number of sequences better than 10.0: 184
Number of HSP's better than 10.0 without gapping: 86
Number of HSP's successfully gapped in prelim test: 98
Number of HSP's that attempted gapping in prelim test: 15678
Number of HSP's gapped (non-prelim): 706
length of query: 1113
length of database: 59,974,054
effective HSP length: 121
effective length of query: 992
effective length of database: 40,105,733
effective search space: 39784887136
effective search space used: 39784887136
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 71 (32.0 bits)
Medicago: description of AC134049.3


