

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133780.8 - phase: 0 /pseudo
(287 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
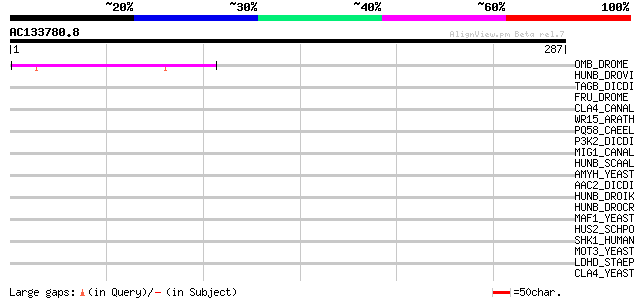
Score E
Sequences producing significant alignments: (bits) Value
OMB_DROME (Q24432) Optomotor-blind protein (Lethal(1)optomotor-b... 53 8e-07
HUNB_DROVI (P13361) Hunchback protein 42 0.002
TAGB_DICDI (P54683) Prestalk-specific protein tagB precursor (EC... 41 0.003
FRU_DROME (Q8IN81) Sex determination protein fruitless 37 0.043
CLA4_CANAL (O14427) Serine/threonine-protein kinase CLA4 (EC 2.7... 37 0.043
WR15_ARATH (O22176) Probable WRKY transcription factor 15 (WRKY ... 37 0.056
PQ58_CAEEL (P34552) Protein pqn-58 (Protein YNK1) 37 0.073
P3K2_DICDI (P54674) Phosphatidylinositol 3-kinase 2 (EC 2.7.1.13... 36 0.095
MIG1_CANAL (Q9Y7G2) Regulatory protein MIG1 36 0.095
HUNB_SCAAL (O46254) Hunchback protein (Fragments) 36 0.095
AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3) (G... 36 0.095
AAC2_DICDI (P14196) AAC-rich mRNA clone AAC11 protein (Fragment) 36 0.095
HUNB_DROIK (O46242) Hunchback protein (Fragments) 36 0.12
HUNB_DROCR (O46236) Hunchback protein (Fragments) 36 0.12
MAF1_YEAST (P41910) Repressor of RNA polymerase III transcriptio... 35 0.16
HUS2_SCHPO (Q09811) ATP-dependent DNA helicase hus2/rqh1 35 0.16
SHK1_HUMAN (Q9Y566) SH3 and multiple ankyrin repeat domains prot... 35 0.21
MOT3_YEAST (P54785) Zinc finger protein MOT3/HMS1 35 0.21
LDHD_STAEP (Q8CN22) D-lactate dehydrogenase (EC 1.1.1.28) (D-LDH... 35 0.21
CLA4_YEAST (P48562) Serine/threonine-protein kinase CLA4 (EC 2.7... 35 0.21
>OMB_DROME (Q24432) Optomotor-blind protein
(Lethal(1)optomotor-blind) (L(1)omb) (Bifid protein)
Length = 972
Score = 53.1 bits (126), Expect = 8e-07
Identities = 35/119 (29%), Positives = 51/119 (42%), Gaps = 13/119 (10%)
Query: 2 NNNNNNNNQNQ------SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPA 55
NNNNNNNN +Q S+ +E P + TPP + P+PP N+ ++ + + A A
Sbjct: 133 NNNNNNNNTSQKQGHHLSTTEEPPSPAGTPPPTIVGLPPIPPPNNNSSSSSSNNSASAAA 192
Query: 56 VPSSHPDNKKAALQTASADGQGQP-------QVHYYQQDHPYVQHSPVDKPSSSPMESI 107
PS HP + T +A P H+ QQ QH P P ++
Sbjct: 193 HPSHHPTAAHHSPSTGAAAPPAGPTGLPPPTPPHHLQQQQQQQQHPAPPPPPYFPAAAL 251
>HUNB_DROVI (P13361) Hunchback protein
Length = 816
Score = 41.6 bits (96), Expect = 0.002
Identities = 28/93 (30%), Positives = 42/93 (45%), Gaps = 1/93 (1%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHP 61
++N++N+N SSP ++P SP PPS+ Q + + A A+ S
Sbjct: 45 HDNHSNSNSGASSPRQSPLPSPIPPSTQLEQYLKQQQQQQQQQQQPMDTLCAAAMTPSPS 104
Query: 62 DNKKAALQTASADGQGQ-PQVHYYQQDHPYVQH 93
+N + +LQ A Q Q Q YQQ QH
Sbjct: 105 NNDQNSLQHFDATLQQQLLQQQQYQQHFQAAQH 137
>TAGB_DICDI (P54683) Prestalk-specific protein tagB precursor (EC
3.4.21.-)
Length = 1905
Score = 41.2 bits (95), Expect = 0.003
Identities = 28/119 (23%), Positives = 39/119 (32%), Gaps = 13/119 (10%)
Query: 1 MNNNNNNNNQNQSSPDENPFASPTPPS-------------SSSSSVPLPPQNHATTENWG 47
+ NNNNN N N ++ + P +S TPP+ + P PPQ +
Sbjct: 1763 LQNNNNNKNNNNNNNNNEPSSSSTPPNDQPTPPPQEQQEQKNDQPPPPPPQEQQEQQEQQ 1822
Query: 48 THMMGAPAVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMES 106
++ Q Q Q Q Q D P + V P P ES
Sbjct: 1823 QQQQQEQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQNDQPPNDYDQVPPPPPLPSES 1881
>FRU_DROME (Q8IN81) Sex determination protein fruitless
Length = 955
Score = 37.4 bits (85), Expect = 0.043
Identities = 27/105 (25%), Positives = 39/105 (36%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHP 61
NNNNNNNN N SS + N +S ++S + + + SS
Sbjct: 400 NNNNNNNNNNNSSSNNNNSSSNRERNNSGERERERERERERDRDRELSTTPVEQLSSSKR 459
Query: 62 DNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMES 106
K ++ ++ HY Q + SPV K S ES
Sbjct: 460 RRKNSSSNCDNSLSSSHQDRHYPQDSQANFKSSPVPKTGGSTSES 504
>CLA4_CANAL (O14427) Serine/threonine-protein kinase CLA4 (EC
2.7.1.37)
Length = 971
Score = 37.4 bits (85), Expect = 0.043
Identities = 30/122 (24%), Positives = 47/122 (37%), Gaps = 30/122 (24%)
Query: 2 NNNNNNNNQNQSSPDENPF-------------ASPTPPSSSSSSVPLPPQNHATTENWGT 48
NNNNNN+ N ++ + +P +P PP+S +SS +NH +
Sbjct: 384 NNNNNNSTNNNNTKNVSPLNNLMNKSELIPARRAPPPPTSGTSSDTYSNKNHQDRSGY-- 441
Query: 49 HMMGAPAVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMESIL 108
++ +T S+ Q Q + H YQQ Q P +P SS
Sbjct: 442 --------------EQQRQQRTDSSQQQQQQKQHQYQQKSQQQQQQP-QQPLSSHQGGTS 486
Query: 109 HM 110
H+
Sbjct: 487 HI 488
>WR15_ARATH (O22176) Probable WRKY transcription factor 15 (WRKY
DNA-binding protein 15)
Length = 317
Score = 37.0 bits (84), Expect = 0.056
Identities = 32/137 (23%), Positives = 56/137 (40%), Gaps = 29/137 (21%)
Query: 12 QSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTA 71
Q P PF SP PP PPQ M+ + SS ++L +
Sbjct: 110 QEEPKTTPFQSPLPP---------PPQ-----------MIRKGSFSSSMKTIDFSSLSSV 149
Query: 72 SADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMESILHMFDSWSKKAEATANNIWHNLKT 131
+ + Q ++H++Q+ P +S +S+ S+SK + N+ NL T
Sbjct: 150 TTESDNQKKIHHHQRPSE-------TAPFASQTQSLSTTVSSFSKSTKRKCNS--ENLLT 200
Query: 132 GPSVSSAAMGKMNLTVK 148
G S+++ G+ + + K
Sbjct: 201 GKCASASSSGRCHCSKK 217
>PQ58_CAEEL (P34552) Protein pqn-58 (Protein YNK1)
Length = 861
Score = 36.6 bits (83), Expect = 0.073
Identities = 27/98 (27%), Positives = 40/98 (40%), Gaps = 9/98 (9%)
Query: 6 NNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKK 65
+ N QS + P P P++ S P+PP T+ GAP + + ++
Sbjct: 721 SGTNAAQSPANAPPRPPPPRPAAPSVESPIPPPR---TQQSMQATPGAPPQYNPYQQQQQ 777
Query: 66 AALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSP 103
+Q Q Q YYQQ PY Q P+ +P P
Sbjct: 778 PQMQ------QFQQHPGYYQQPMPYGQPQPMFQPQYQP 809
>P3K2_DICDI (P54674) Phosphatidylinositol 3-kinase 2 (EC 2.7.1.137)
(PI3-kinase) (PtdIns-3-kinase) (PI3K)
Length = 1858
Score = 36.2 bits (82), Expect = 0.095
Identities = 26/84 (30%), Positives = 38/84 (44%), Gaps = 12/84 (14%)
Query: 2 NNNNNNNNQNQSSPDENPFASPT------------PPSSSSSSVPLPPQNHATTENWGTH 49
NNNNNNNN N ++ + N S T PPSSSSSS Q + N ++
Sbjct: 209 NNNNNNNNNNNNNNNNNNTTSTTTTTTSILISSSPPPSSSSSSSSNDEQFNNNNNNNNSN 268
Query: 50 MMGAPAVPSSHPDNKKAALQTASA 73
G+ + +S K + + +A
Sbjct: 269 SGGSSRMITSKSQIKPLIVTSNTA 292
>MIG1_CANAL (Q9Y7G2) Regulatory protein MIG1
Length = 574
Score = 36.2 bits (82), Expect = 0.095
Identities = 21/75 (28%), Positives = 36/75 (48%), Gaps = 4/75 (5%)
Query: 2 NNNNNNNNQNQSSPDEN----PFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVP 57
+ NN NN S+ + N P P PSS+S ++ N+ +N+ TH P P
Sbjct: 377 STTNNTNNTTTSNTNNNNMTKPSIIPKQPSSTSLNLEFYNGNNQQQQNYHTHKKSRPNSP 436
Query: 58 SSHPDNKKAALQTAS 72
S P + ++ ++A+
Sbjct: 437 SQTPIHLSSSRKSAN 451
Score = 32.3 bits (72), Expect = 1.4
Identities = 25/113 (22%), Positives = 46/113 (40%), Gaps = 17/113 (15%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPP----------QNHATTENWGTHMM 51
+N+NN+ N SS + + SSS++ L N+ TT N + M
Sbjct: 336 SNSNNSRLFNASSSSLSSLSGKIRSSSSTNLAGLQRLTPLTSTTNNTNNTTTSNTNNNNM 395
Query: 52 GAPAVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPM 104
P++ P + L+ + + Q Q H +++ P + PS +P+
Sbjct: 396 TKPSIIPKQPSSTSLNLEFYNGNNQQQQNYHTHKKSRP-------NSPSQTPI 441
>HUNB_SCAAL (O46254) Hunchback protein (Fragments)
Length = 171
Score = 36.2 bits (82), Expect = 0.095
Identities = 14/28 (50%), Positives = 22/28 (78%)
Query: 3 NNNNNNNQNQSSPDENPFASPTPPSSSS 30
++++N+N N SSP ++P SP PPSSS+
Sbjct: 28 HHHSNSNSNASSPRQSPLPSPNPPSSSN 55
>AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3)
(Glucan 1,4-alpha-glucosidase) (1,4-alpha-D-glucan
glucohydrolase)
Length = 1367
Score = 36.2 bits (82), Expect = 0.095
Identities = 39/185 (21%), Positives = 72/185 (38%), Gaps = 27/185 (14%)
Query: 5 NNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNK 64
++++N S+P PF+S S+ SSSVP+P + +TTE+ + V SS ++
Sbjct: 813 SSSSNITSSAPSSTPFSS----STESSSVPVPTPSSSTTES------SSAPVSSSTTESS 862
Query: 65 KAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMESILHMFDSWSKKAEATANN 124
A + T S+ + PSS P S F + + +++
Sbjct: 863 VAPVPTPSSS-----------------SNITSSAPSSIPFSSTTESFSTGTTVTPSSSKY 905
Query: 125 IWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTG 184
+T S ++ T +++ ++ + +T N + T C T T
Sbjct: 906 PGSQTETSVSSTTETTIVPTKTTTSVTTPSTTTITTTVCSTGTNSAGETTSGCSPKTVTT 965
Query: 185 PVAGT 189
V T
Sbjct: 966 TVPTT 970
Score = 35.0 bits (79), Expect = 0.21
Identities = 31/112 (27%), Positives = 51/112 (44%), Gaps = 18/112 (16%)
Query: 6 NNNNQNQSSPDENPFAS---------PTPPSSS--SSSVPLPPQNHATTENWGTHMMGAP 54
++ ++ S+P P +S PTP SS+ SSS P+P + +TTE+ +
Sbjct: 628 SSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTES------SSA 681
Query: 55 AVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMES 106
V SS ++ A + +++ + P V +PV PSSS ES
Sbjct: 682 PVTSSTTESSSAPVTSSTTESSSAP-VPTPSSSTTESSSAPVPTPSSSTTES 732
Score = 34.3 bits (77), Expect = 0.36
Identities = 29/101 (28%), Positives = 45/101 (43%), Gaps = 12/101 (11%)
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWG-------THMMGAPAVPSSHPDNKK 65
SS E+ A T ++ SSS P+P + +TTE+ T AP V SS ++
Sbjct: 373 SSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAP-VTSSTTESSS 431
Query: 66 AALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMES 106
A + +++ + P + +PV PSSS ES
Sbjct: 432 APVTSSTTESSSAPVTSSTTES----SSAPVPTPSSSTTES 468
Score = 33.9 bits (76), Expect = 0.47
Identities = 29/110 (26%), Positives = 47/110 (42%), Gaps = 27/110 (24%)
Query: 5 NNNNNQNQSSPDENPFASPTPPSSS--------SSSVPLPPQNHATTENWGTHMMGAPAV 56
+++ ++ S+P P +S T SS+ SSS P+P + +TTE+ T + +
Sbjct: 516 SSSTTESSSAPAPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSTPVTSSTTE 575
Query: 57 PSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMES 106
SS P ++ T S+ +PV PSSS ES
Sbjct: 576 SSSAPVPTPSSSTTESSS-------------------APVPTPSSSTTES 606
Score = 33.9 bits (76), Expect = 0.47
Identities = 30/102 (29%), Positives = 40/102 (38%), Gaps = 8/102 (7%)
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWG-------THMMGAPA-VPSSHPDNK 64
SS E+ A T ++ SSS P+P + +TTE+ T AP PSS
Sbjct: 436 SSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTES 495
Query: 65 KAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMES 106
+A T+S V +P PSSS ES
Sbjct: 496 SSAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTES 537
Score = 33.5 bits (75), Expect = 0.62
Identities = 33/129 (25%), Positives = 55/129 (42%), Gaps = 15/129 (11%)
Query: 6 NNNNQNQSSPDENPFAS---------PTPPSSS--SSSVPLPPQNHATTENWGTHMMGAP 54
++ ++ S+P P +S PTP SS+ SSS P+P + +TTE+ + +
Sbjct: 697 SSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSST 756
Query: 55 AVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMESILHMFDSW 114
SS P ++ T S+ V +PV PSSS ES + +
Sbjct: 757 TESSSAPVPTPSSSTTESSSA----PVPTPSSSTTESSSAPVPTPSSSTTESSVAPVPTP 812
Query: 115 SKKAEATAN 123
S + T++
Sbjct: 813 SSSSNITSS 821
Score = 32.3 bits (72), Expect = 1.4
Identities = 31/112 (27%), Positives = 47/112 (41%), Gaps = 15/112 (13%)
Query: 6 NNNNQNQSSPDENPFAS---------PTPPSSS--SSSVPLPPQNHATTENWGTHMMGAP 54
++ ++ S+P P +S PTP SS+ SSS P P + +TTE+ + +
Sbjct: 571 SSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTSST 630
Query: 55 AVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMES 106
SS P ++ T S+ V +PV PSSS ES
Sbjct: 631 TESSSAPVPTPSSSTTESSSA----PVPTPSSSTTESSSAPVPTPSSSTTES 678
Score = 32.3 bits (72), Expect = 1.4
Identities = 26/95 (27%), Positives = 42/95 (43%), Gaps = 15/95 (15%)
Query: 23 PTPPSSS--SSSVPLPPQNHATTENWGTHMMGA---------PAVPSSHPDNKKAALQTA 71
PTP SS+ SSS P+P + +TTE+ + + P SS ++ A + ++
Sbjct: 315 PTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSS 374
Query: 72 SADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMES 106
+ + P + +PV PSSS ES
Sbjct: 375 TTESSSAPVTSSTTES----SSAPVPTPSSSTTES 405
Score = 31.2 bits (69), Expect = 3.1
Identities = 29/102 (28%), Positives = 42/102 (40%), Gaps = 8/102 (7%)
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTAS 72
SS E+ A T ++ SSS P+P + +TTE+ + + SS P ++ T S
Sbjct: 463 SSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTES 522
Query: 73 ADGQG-QPQVHYYQQDHPYVQHS-------PVDKPSSSPMES 106
+ P + V S PV PSSS ES
Sbjct: 523 SSAPAPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTES 564
>AAC2_DICDI (P14196) AAC-rich mRNA clone AAC11 protein (Fragment)
Length = 448
Score = 36.2 bits (82), Expect = 0.095
Identities = 22/80 (27%), Positives = 35/80 (43%), Gaps = 8/80 (10%)
Query: 1 MNNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSH 60
+NNNNNN+N N ++ + N + +P SS+ N A + G P+ P
Sbjct: 127 INNNNNNSNNNNNNNNSNLGINSSPTQSSA--------NSADKRSRGRPRKNPPSEPKDT 178
Query: 61 PDNKKAALQTASADGQGQPQ 80
K+ + D +G PQ
Sbjct: 179 SGPKRKRGRPPKMDEEGNPQ 198
>HUNB_DROIK (O46242) Hunchback protein (Fragments)
Length = 193
Score = 35.8 bits (81), Expect = 0.12
Identities = 13/29 (44%), Positives = 23/29 (78%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSSS 30
+++++N+N N SSP ++P SP PPSS++
Sbjct: 31 HSHDSNSNSNASSPHQSPLPSPNPPSSNN 59
>HUNB_DROCR (O46236) Hunchback protein (Fragments)
Length = 190
Score = 35.8 bits (81), Expect = 0.12
Identities = 14/26 (53%), Positives = 21/26 (79%)
Query: 5 NNNNNQNQSSPDENPFASPTPPSSSS 30
++N+N+N SSP ++P SP PPSSS+
Sbjct: 33 HSNSNRNASSPRQSPLPSPNPPSSSN 58
>MAF1_YEAST (P41910) Repressor of RNA polymerase III transcription
MAF1
Length = 395
Score = 35.4 bits (80), Expect = 0.16
Identities = 20/75 (26%), Positives = 41/75 (54%), Gaps = 3/75 (4%)
Query: 2 NNNNNNNNQNQSSPDENPFASP--TPPSSSSSSVPLPPQN-HATTENWGTHMMGAPAVPS 58
N+NNN NN N +S + N ++ P + P++ S L QN N+ + M + ++ S
Sbjct: 106 NSNNNTNNSNGNSSNNNNYSGPNGSSPATFPKSAKLNDQNLKELVSNYDSGSMSSSSLDS 165
Query: 59 SHPDNKKAALQTASA 73
S ++++ +++S+
Sbjct: 166 SSKNDERIRRRSSSS 180
>HUS2_SCHPO (Q09811) ATP-dependent DNA helicase hus2/rqh1
Length = 1328
Score = 35.4 bits (80), Expect = 0.16
Identities = 15/30 (50%), Positives = 21/30 (70%)
Query: 1 MNNNNNNNNQNQSSPDENPFASPTPPSSSS 30
+NNNN NNN + ++ ++ ASPTP S SS
Sbjct: 224 LNNNNTNNNNDNNAIEKRDSASPTPSSVSS 253
Score = 35.0 bits (79), Expect = 0.21
Identities = 16/35 (45%), Positives = 19/35 (53%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLP 36
NNNN+NNN + N TPPSS + VP P
Sbjct: 321 NNNNSNNNNGNNGTVPNAKTFFTPPSSITQQVPFP 355
>SHK1_HUMAN (Q9Y566) SH3 and multiple ankyrin repeat domains protein 1
(Shank1) (Somatostatin receptor interacting protein)
(SSTR interacting protein) (SSTRIP)
Length = 2161
Score = 35.0 bits (79), Expect = 0.21
Identities = 23/87 (26%), Positives = 37/87 (42%), Gaps = 3/87 (3%)
Query: 15 PDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAV--PSSHPDNKKAALQTAS 72
P E P+++PT PSSS P P + G +PA S PD + + + +
Sbjct: 928 PPEPPYSTPTVPSSSGRLTPSPRGGPFNPGSGGPLPASSPATFDGPSPPDTRVGSREKSL 987
Query: 73 ADGQGQPQVHYYQQDHPYVQHSPVDKP 99
P H++ H + H+P +P
Sbjct: 988 YHSGPLPPAHHHPPHHHH-HHAPPPQP 1013
Score = 32.3 bits (72), Expect = 1.4
Identities = 18/44 (40%), Positives = 21/44 (46%), Gaps = 3/44 (6%)
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHAT---TENWGTHMMGA 53
SSP P SP PPS S P P AT T +G ++GA
Sbjct: 1196 SSPSPAPAMSPVPPSPSPVPTPASPSGPATLDFTSQFGAALVGA 1239
>MOT3_YEAST (P54785) Zinc finger protein MOT3/HMS1
Length = 490
Score = 35.0 bits (79), Expect = 0.21
Identities = 24/103 (23%), Positives = 46/103 (44%), Gaps = 17/103 (16%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSS----------SSSVPLPPQNHATTENWGTHMM 51
NNNNNNNN N ++ N F + +S+ ++++P+P Q+ N +++
Sbjct: 145 NNNNNNNNNNNNNIHPNQFTAAANMNSNAAAAAYYSFPTANMPIPQQDQQYMFNPASYIS 204
Query: 52 GAPAVPSSHPDNKKAA-------LQTASADGQGQPQVHYYQQD 87
+ +S+ + AA +A A G P H++ +
Sbjct: 205 HYYSAVNSNNNGNNAANNGSNNSSHSAPAPAPGPPHHHHHHSN 247
Score = 31.6 bits (70), Expect = 2.3
Identities = 33/132 (25%), Positives = 50/132 (37%), Gaps = 31/132 (23%)
Query: 1 MNNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAV---- 56
+N+NNN NN + + + ++P P P PP +H N ++ AV
Sbjct: 210 VNSNNNGNNAANNGSNNSSHSAPAP-------APGPPHHHHHHSNTHNNLNNGGAVNTNN 262
Query: 57 -PSSHP----DNKKAALQTASAD------GQGQP-------QVHYYQQDHPYV--QHSPV 96
P HP D + LQ + QP +++ Q P H P
Sbjct: 263 APQHHPTIITDQFQFQLQQNPSPNLNLNINPAQPLHLPPGWKINTMPQPRPTTAPNHPPA 322
Query: 97 DKPSSSPMESIL 108
PSS+P+ S L
Sbjct: 323 PVPSSNPVASNL 334
>LDHD_STAEP (Q8CN22) D-lactate dehydrogenase (EC 1.1.1.28) (D-LDH)
(D-specific D-2-hydroxyacid dehydrogenase)
Length = 330
Score = 35.0 bits (79), Expect = 0.21
Identities = 35/125 (28%), Positives = 47/125 (37%), Gaps = 22/125 (17%)
Query: 93 HSPVDKPSSSPMESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISE 152
H P +K S H+FD NN++ N+K G + +AA G + T I
Sbjct: 205 HVPANKDS-------FHLFD----------NNMFKNVKKGAVLVNAARGAVINTPDLIEA 247
Query: 153 GGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTL-----YLSDIHLAFCSDRPLSF 207
+L TY NE TF C T P+ L L H+AF SD +
Sbjct: 248 VNNGTLSGAAIDTYENEANYFTFDCSNQTIDDPILLDLIRNENILVTPHIAFFSDEAVQN 307
Query: 208 TAPSG 212
G
Sbjct: 308 LVEGG 312
>CLA4_YEAST (P48562) Serine/threonine-protein kinase CLA4 (EC
2.7.1.37)
Length = 842
Score = 35.0 bits (79), Expect = 0.21
Identities = 24/81 (29%), Positives = 34/81 (41%), Gaps = 8/81 (9%)
Query: 28 SSSSSVP-----LPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADGQGQPQVH 82
+S S++P +P Q + N + G HP+N + +LQ Q Q Q H
Sbjct: 326 TSQSNIPRHLQNVPNQQYPKMRNGHSPTNGQFPRGPMHPNNSQRSLQQQQQQQQQQKQQH 385
Query: 83 YYQQDHPYVQHSPVDKPSSSP 103
Q +PY P PS SP
Sbjct: 386 ---QQYPYHHQGPSPSPSPSP 403
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.127 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,173,633
Number of Sequences: 164201
Number of extensions: 1640795
Number of successful extensions: 9874
Number of sequences better than 10.0: 188
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 121
Number of HSP's that attempted gapping in prelim test: 7987
Number of HSP's gapped (non-prelim): 966
length of query: 287
length of database: 59,974,054
effective HSP length: 109
effective length of query: 178
effective length of database: 42,076,145
effective search space: 7489553810
effective search space used: 7489553810
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 65 (29.6 bits)
Medicago: description of AC133780.8


