

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133779.5 + phase: 1 /pseudo
(547 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
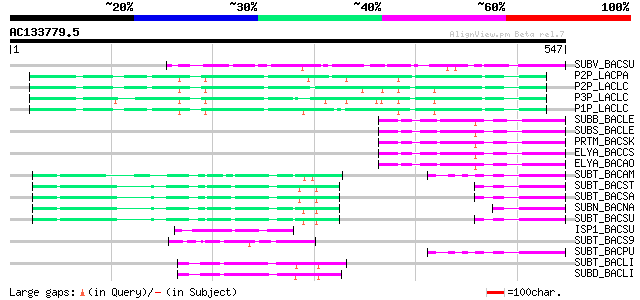
Score E
Sequences producing significant alignments: (bits) Value
SUBV_BACSU (P29141) Minor extracellular protease vpr precursor (... 106 1e-22
P2P_LACPA (Q02470) PII-type proteinase precursor (EC 3.4.21.96) ... 78 5e-14
P2P_LACLC (P15293) PII-type proteinase precursor (EC 3.4.21.96) ... 75 3e-13
P3P_LACLC (P15292) PIII-type proteinase precursor (EC 3.4.21.96)... 74 7e-13
P1P_LACLC (P16271) PI-type proteinase precursor (EC 3.4.21.-) (W... 73 2e-12
SUBB_BACLE (P29599) Subtilisin BL (EC 3.4.21.62) (Alkaline prote... 62 5e-09
SUBS_BACLE (P29600) Subtilisin Savinase (EC 3.4.21.62) (Alkaline... 59 4e-08
PRTM_BACSK (Q99405) M-protease (EC 3.4.21.-) 59 4e-08
ELYA_BACCS (P41362) Alkaline protease precursor (EC 3.4.21.-) 59 4e-08
ELYA_BACAO (P27693) Alkaline protease precursor (EC 3.4.21.-) 59 4e-08
SUBT_BACAM (P00782) Subtilisin BPN' precursor (EC 3.4.21.62) (Su... 55 6e-07
SUBT_BACST (P29142) Subtilisin J precursor (EC 3.4.21.62) 54 1e-06
SUBT_BACSA (P00783) Subtilisin amylosacchariticus precursor (EC ... 54 1e-06
SUBN_BACNA (P35835) Subtilisin NAT precursor (EC 3.4.21.62) 53 2e-06
SUBT_BACSU (P04189) Subtilisin E precursor (EC 3.4.21.62) 53 2e-06
ISP1_BACSU (P11018) Major intracellular serine protease precurso... 53 2e-06
SUBT_BACS9 (P28842) Subtilisin precursor (EC 3.4.21.62) 49 3e-05
SUBT_BACPU (P07518) Subtilisin (EC 3.4.21.62) (Alkaline mesenter... 49 3e-05
SUBT_BACLI (P00780) Subtilisin Carlsberg precursor (EC 3.4.21.62) 49 3e-05
SUBD_BACLI (P00781) Subtilisin DY (EC 3.4.21.62) 49 4e-05
>SUBV_BACSU (P29141) Minor extracellular protease vpr precursor (EC
3.4.21.-)
Length = 806
Score = 106 bits (265), Expect = 1e-22
Identities = 107/401 (26%), Positives = 183/401 (44%), Gaps = 68/401 (16%)
Query: 155 PRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDA 214
PR + T HGTH T N GTIKG +P A + Y+V
Sbjct: 226 PRGEAT-----DHGTHVAGTVAAN------------GTIKGVAPDATLLAYRVLGPGGSG 268
Query: 215 TSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVAS 274
T+ +V++ +++A+ DG D++++S G +S N+ + T S A++ ++ V S
Sbjct: 269 TT---ENVIAGVERAVQDGADVMNLSLG--NSLNNPDWAT---STALDWAMSEGVVAVTS 320
Query: 275 AGNEGPTPGSVVN--VAPWVFTVAASTLDRDFSSVMTIGNKTLTGASLFVNLPPNQDFTI 332
GN GP +V + + +V A+ L + +V T G+ + ++ + D
Sbjct: 321 NGNSGPNGWTVGSPGTSREAISVGATQLPLNEYAV-TFGSYS---SAKVMGYNKEDDVKA 376
Query: 333 VTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVI 392
+ + + +L A +A+ + + GK+ R G I V + A AGA G++
Sbjct: 377 LNNKEVELVEAGIGEAK-----DFEGKDLTGKVAVVKR-GSIAFVDKADNAKKAGAIGMV 430
Query: 393 LRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDI---IPSDIKSG---TKLRMSPA 446
+ N ++G+ + PG T SL+ + S +K+G T +++ +
Sbjct: 431 VYNN--LSGEI---------EANVPGMSVPTIKLSLEDGEKLVSALKAGETKTTFKLTVS 479
Query: 447 KTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGF 506
K L + +A +SSRGP + ++KPD++APGVNI++ D +
Sbjct: 480 KALGEQ-----VADFSSRGP-VMDTWMIKPDISAPGVNIVSTIPTH--------DPDHPY 525
Query: 507 PFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
+ QGTSM+ PH+AG +IK P WS IK+AI T
Sbjct: 526 GYGSKQGTSMASPHIAGAVAVIKQAKPKWSVEQIKAAIMNT 566
>P2P_LACPA (Q02470) PII-type proteinase precursor (EC 3.4.21.96)
(Lactocepin) (Cell wall-associated serine proteinase)
(LP151)
Length = 1902
Score = 78.2 bits (191), Expect = 5e-14
Identities = 116/537 (21%), Positives = 205/537 (37%), Gaps = 78/537 (14%)
Query: 20 SYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQK 79
SY +NGF+ + + ++ + V +V L+K + ++ ++ + W
Sbjct: 149 SYGYVVNGFSTKVRVVDIPKLKQIAGVKTVTLAKVYYPTDAKANSMA-----NVQAVWSN 203
Query: 80 GRF-GENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLI 138
++ GE T++ IDTG+ P K + +L K + K +
Sbjct: 204 YKYKGEGTVVSVIDTGIDPTHKDM----------RLSDDKDVKLTKYDVEKFTDTAKH-- 251
Query: 139 GARFFNKAYQKRNGKLPRSQQTARDFVG--HGTHTLSTAGGNFVPGASIFNIGNG----- 191
R+F + D V HG H G N G G
Sbjct: 252 -GRYFTSKVPYGFNYADNNDTITDDTVDEQHGMHVAGIIGAN----------GTGDDPTK 300
Query: 192 TIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEE 251
++ G +P A++ KV + + + A ++SAI+ + G D++++S G S + E
Sbjct: 301 SVVGVAPEAQLLAMKVFTNSDTSATTGSATLVSAIEDSAKIGADVLNMSLGSDSGNQTLE 360
Query: 252 IFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVF------TVAASTLDRDFS 305
D +A V SAGN G + + V + V R +
Sbjct: 361 ---DPEIAAVQNANESGTAAVISAGNSGTSGSATQGVNKDYYGLQDNEMVGTPGTSRGAT 417
Query: 306 SVMTIGNKTLTGASLFVNLPPNQDF-----TIVTSTDAKLANATNRDARFCRPRTLDPSK 360
+V + N + S V + +D TI S++ + + + + D SK
Sbjct: 418 TVASAENTDV--ISQAVTITDGKDLQLGPETIQLSSNDFTGSFDQKKFYVVKDASGDLSK 475
Query: 361 VNGKIVACDREGKIKSVAEGQ--------EALSAGAKGVILRNQPEINGKTLLSEPHVLS 412
D +GKI V G+ A +AGA G+I+ N T L+ + +
Sbjct: 476 GAAADYTADAKGKIAIVKRGELNFADKQKYAQAAGAAGLIIVNND--GTATPLTSIRLTT 533
Query: 413 TISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPS 472
T G S+T + +D + + ++++ N++ M+ ++S GP V
Sbjct: 534 TFPTFGLSSKTGQKLVDWVTAHPDDSLGVKIALTLLPNQKYTEDKMSDFTSYGP--VSNL 591
Query: 473 ILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIK 529
KPD+TAPG NI + T+ + M GTSM+ P +AG+ L+K
Sbjct: 592 SFKPDITAPGGNIWS--------------TQNNNGYTNMSGTSMASPFIAGSQALLK 634
>P2P_LACLC (P15293) PII-type proteinase precursor (EC 3.4.21.96)
(Lactocepin) (Cell wall-associated serine proteinase)
(LP151)
Length = 1902
Score = 75.5 bits (184), Expect = 3e-13
Identities = 117/541 (21%), Positives = 202/541 (36%), Gaps = 86/541 (15%)
Query: 20 SYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQK 79
SY +NGF+ + + ++ + V +V L+K + ++ ++ + W
Sbjct: 149 SYGYVVNGFSTKVRVVDIPKLKQIAGVKTVTLAKVYYPTDAKANSMA-----NVQAVWSN 203
Query: 80 GRF-GENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLI 138
++ GE T++ ID+G+ P K + +L K + K +
Sbjct: 204 YKYKGEGTVVSVIDSGIDPTHKDM----------RLSDDKDVKLTKSDVEKFTDTAKH-- 251
Query: 139 GARFFNKAYQKRNGKLPRSQQTARDFVG--HGTHTLSTAGGNFVPGASIFNIGNG----- 191
R+FN + D V HG H G N G G
Sbjct: 252 -GRYFNSKVPYGFNYADNNDTITDDTVDEQHGMHVAGIIGAN----------GTGDDPAK 300
Query: 192 TIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEE 251
++ G +P A++ KV + + + A ++SAI+ + G D++++S G S + E
Sbjct: 301 SVVGVAPEAQLLAMKVFTNSDTSATTGSATLVSAIEDSAKIGADVLNMSLGSDSGNQTLE 360
Query: 252 IFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIG 311
D +A V SAGN G + + V + + D +
Sbjct: 361 ---DPELAAVQNANESGTAAVISAGNSGTSGSATEGVNKDYYGLQ----DNEMVGTPGTS 413
Query: 312 NKTLTGASLFVNLPPNQDFTIVTSTDAKLANATNR-----------DARFCRPRTLDPSK 360
T AS Q TI T +L T + +F + +
Sbjct: 414 RGATTVASAENTDVITQAVTITDGTGLQLGPETIQLSSNDFTGSFDQKKFYVVKDASGNL 473
Query: 361 VNGKIV--ACDREGKIKSVAEGQ--------EALSAGAKGVILRNQPEINGKTLLSEPHV 410
GK+ D +GKI V G+ A +AGA G+I+ N N T +
Sbjct: 474 SKGKVADYTADAKGKIAIVKRGELTFADKQKYAQAAGAAGLIIVN----NDGTATPVTSM 529
Query: 411 LSTISYP--GNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNK 468
T ++P G S T + +D + + ++++ N++ M+ ++S GP
Sbjct: 530 ALTTTFPTFGLSSVTGQKLVDWVAAHPDDSLGVKIALTLVPNQKYTEDKMSDFTSYGP-- 587
Query: 469 VQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLI 528
V KPD+TAPG NI + T+ + M GTSM+ P +AG+ L+
Sbjct: 588 VSNLSFKPDITAPGGNIWS--------------TQNNNGYTNMSGTSMASPFIAGSQALL 633
Query: 529 K 529
K
Sbjct: 634 K 634
>P3P_LACLC (P15292) PIII-type proteinase precursor (EC 3.4.21.96)
(Lactocepin) (Cell wall-associated serine proteinase)
Length = 1902
Score = 74.3 bits (181), Expect = 7e-13
Identities = 122/552 (22%), Positives = 208/552 (37%), Gaps = 108/552 (19%)
Query: 20 SYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQK 79
SY +NGF+ + + ++ + V +V L+K + ++ ++ + W
Sbjct: 149 SYGYVVNGFSTKVRVVDIPKLKQIAGVKTVTLAKVYYPTDAKANSMA-----NVQAVWSN 203
Query: 80 GRF-GENTIIGNIDTGVWPESKSF---SDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNR 135
++ GE T++ ID+G+ P K D+ + KL S
Sbjct: 204 YKYKGEGTVVSVIDSGIDPTHKDMRLSDDKDV----------------KLTKSDVEKFTD 247
Query: 136 KLIGARFFNKAYQKRNGKLPRSQQTARDFVG--HGTHTLSTAGGNFVPGASIFNIGNG-- 191
+ R+FN + D V HG H G N G G
Sbjct: 248 TVKHGRYFNSKVPYGFNYADNNDTITDDKVDEQHGMHVAGIIGAN----------GTGDD 297
Query: 192 ---TIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTN 248
++ G +P A++ KV + + A V+SAI+ + G D++++S G S
Sbjct: 298 PAKSVVGVAPEAQLLAMKVFSNSDTSAKTGSATVVSAIEDSAKIGADVLNMSLGSNSGNQ 357
Query: 249 SEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVM 308
+ E D +A V SAGN G T GS A +++D+ +
Sbjct: 358 TLE---DPELAAVQNANESGTAAVISAGNSG-TSGS-----------ATEGVNKDYYGLQ 402
Query: 309 T---IGNKTLTGASLFVNLPPNQDF-----TIVTSTDAKLANATNRDARFCRPRTLDPSK 360
+G+ + + V N D TI T +L T + + + D K
Sbjct: 403 DNEMVGSPGTSRGATTVASAENTDVITQAVTITDGTGLQLGPETIQLSSHDFTGSFDQKK 462
Query: 361 ------VNGKI-------VACDREGKIKSVAEGQ--------EALSAGAKGVILRNQPEI 399
+G + D +GKI V G+ A +AGA G+I+ N
Sbjct: 463 FYIVKDASGNLSKGALADYTADAKGKIAIVKRGEFSFDDKQKYAQAAGAAGLIIVN---- 518
Query: 400 NGKTLLSEPHVLSTISYP--GNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPV 457
T + T ++P G S T + +D + + ++++ A N++
Sbjct: 519 TDGTATPMTSIALTTTFPTFGLSSVTGQKLVDWVTAHPDDSLGVKITLAMLPNQKYTEDK 578
Query: 458 MASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMS 517
M+ ++S GP V KPD+TAPG NI + T+ + M GTSM+
Sbjct: 579 MSDFTSYGP--VSNLSFKPDITAPGGNIWS--------------TQNNNGYTNMSGTSMA 622
Query: 518 CPHVAGTAGLIK 529
P +AG+ L+K
Sbjct: 623 SPFIAGSQALLK 634
>P1P_LACLC (P16271) PI-type proteinase precursor (EC 3.4.21.-)
(Wall-associated serine proteinase)
Length = 1902
Score = 72.8 bits (177), Expect = 2e-12
Identities = 116/542 (21%), Positives = 204/542 (37%), Gaps = 88/542 (16%)
Query: 20 SYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQK 79
SY +NGF+ + + ++ + V +V L+K + ++ ++ + W
Sbjct: 149 SYGYVVNGFSTKVRVVDIPKLKQIAGVKTVTLAKVYYPTDAKANSMA-----NVQAVWSN 203
Query: 80 GRF-GENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLI 138
++ GE T++ ID+G+ P K + +L K + K +
Sbjct: 204 YKYKGEGTVVSVIDSGIDPTHKDM----------RLSDDKDVKLTKSDVEKFTDTAKH-- 251
Query: 139 GARFFNKAYQKRNGKLPRSQQTARDFVG--HGTHTLSTAGGNFVPGASIFNIGNG----- 191
R+FN + D V HG H G N G G
Sbjct: 252 -GRYFNSKVPYGFNYADNNDTITDDTVDEQHGMHVAGIIGAN----------GTGDDPAK 300
Query: 192 TIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEE 251
++ G +P A++ KV + + + + ++SAI+ + G D++++S G S + E
Sbjct: 301 SVVGVAPEAQLLAMKVFTNSDTSATTGSSTLVSAIEDSAKIGADVLNMSLGSDSGNQTLE 360
Query: 252 IFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNV--------------APWVFTVAA 297
D +A V SAGN G + + V P A
Sbjct: 361 ---DPELAAVQNANESGTAAVISAGNSGTSGSATEGVNKDYYGLQDNEMVGTPGTSRGAT 417
Query: 298 STLDRDFSSVMTIGNKTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLD 357
+ + + V+T G L L P TI S++ + + + + +
Sbjct: 418 TVASAENTDVITQAVTITDGTGL--QLGPG---TIQLSSNDFTGSFDQKKFYVVKDASGN 472
Query: 358 PSKVNGKIVACDREGKIKSVAEGQ--------EALSAGAKGVILRNQPEINGKTLLSEPH 409
SK D +GKI V G+ A +AGA G+I+ N N T
Sbjct: 473 LSKGALADYTADAKGKIAIVKRGELSFDDKQKYAQAAGAAGLIIVN----NDGTATPVTS 528
Query: 410 VLSTISYP--GNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPN 467
+ T ++P G S T + +D + + ++++ N++ M+ ++S GP
Sbjct: 529 MALTTTFPTFGLSSVTGQKLVDWVTAHPDDSLGVKIALTLVPNQKYTEDKMSDFTSYGP- 587
Query: 468 KVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGL 527
V KPD+TAPG NI + T+ + M GTSM+ P +AG+ L
Sbjct: 588 -VSNLSFKPDITAPGGNIWS--------------TQNNNGYTNMSGTSMASPFIAGSQAL 632
Query: 528 IK 529
+K
Sbjct: 633 LK 634
>SUBB_BACLE (P29599) Subtilisin BL (EC 3.4.21.62) (Alkaline
protease)
Length = 269
Score = 61.6 bits (148), Expect = 5e-09
Identities = 52/189 (27%), Positives = 80/189 (41%), Gaps = 38/189 (20%)
Query: 364 KIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRT 423
K++ D G I S+A+G E AG G+ + N L P +T+ N +
Sbjct: 92 KVLGADGRGAISSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLEQAVNSA-- 140
Query: 424 TGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPV-----MASYSSRGPNKVQPSILKPDV 478
T R + ++ + SG PA+ N AS+S G D+
Sbjct: 141 TSRGVLVVAASGNSGASSISYPARYANAMAVGATDQNNNRASFSQYGAGL--------DI 192
Query: 479 TAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPA 538
APGVN+ + Y G + + GTSM+ PHVAG A L+K +P+WS
Sbjct: 193 VAPGVNVQSTYP--------------GSTYASLNGTSMATPHVAGAAALVKQKNPSWSNV 238
Query: 539 AIKSAIRTT 547
I++ ++ T
Sbjct: 239 QIRNHLKNT 247
>SUBS_BACLE (P29600) Subtilisin Savinase (EC 3.4.21.62) (Alkaline
protease)
Length = 269
Score = 58.5 bits (140), Expect = 4e-08
Identities = 50/189 (26%), Positives = 79/189 (41%), Gaps = 38/189 (20%)
Query: 364 KIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRT 423
K++ G + S+A+G E AG G+ + N L P +T+ N +
Sbjct: 92 KVLGASGSGSVSSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLEQAVNSA-- 140
Query: 424 TGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPV-----MASYSSRGPNKVQPSILKPDV 478
T R + ++ + SG PA+ N AS+S G D+
Sbjct: 141 TSRGVLVVAASGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAGL--------DI 192
Query: 479 TAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPA 538
APGVN+ + Y G + + GTSM+ PHVAG A L+K +P+WS
Sbjct: 193 VAPGVNVQSTYP--------------GSTYASLNGTSMATPHVAGAAALVKQKNPSWSNV 238
Query: 539 AIKSAIRTT 547
I++ ++ T
Sbjct: 239 QIRNHLKNT 247
>PRTM_BACSK (Q99405) M-protease (EC 3.4.21.-)
Length = 269
Score = 58.5 bits (140), Expect = 4e-08
Identities = 50/189 (26%), Positives = 79/189 (41%), Gaps = 38/189 (20%)
Query: 364 KIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRT 423
K++ G + S+A+G E AG G+ + N L P +T+ N +
Sbjct: 92 KVLGASGSGSVSSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLEQAVNSA-- 140
Query: 424 TGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPV-----MASYSSRGPNKVQPSILKPDV 478
T R + ++ + SG PA+ N AS+S G D+
Sbjct: 141 TSRGVLVVAASGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAGL--------DI 192
Query: 479 TAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPA 538
APGVN+ + Y G + + GTSM+ PHVAG A L+K +P+WS
Sbjct: 193 VAPGVNVQSTYP--------------GSTYASLNGTSMATPHVAGVAALVKQKNPSWSNV 238
Query: 539 AIKSAIRTT 547
I++ ++ T
Sbjct: 239 QIRNHLKNT 247
>ELYA_BACCS (P41362) Alkaline protease precursor (EC 3.4.21.-)
Length = 380
Score = 58.5 bits (140), Expect = 4e-08
Identities = 50/189 (26%), Positives = 79/189 (41%), Gaps = 38/189 (20%)
Query: 364 KIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRT 423
K++ G + S+A+G E AG G+ + N L P +T+ N +
Sbjct: 203 KVLGASGSGSVSSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLEQAVNSA-- 251
Query: 424 TGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPV-----MASYSSRGPNKVQPSILKPDV 478
T R + ++ + SG PA+ N AS+S G D+
Sbjct: 252 TSRGVLVVAASGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAGL--------DI 303
Query: 479 TAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPA 538
APGVN+ + Y G + + GTSM+ PHVAG A L+K +P+WS
Sbjct: 304 VAPGVNVQSTYP--------------GSTYASLNGTSMATPHVAGAAALVKQKNPSWSNV 349
Query: 539 AIKSAIRTT 547
I++ ++ T
Sbjct: 350 QIRNHLKNT 358
>ELYA_BACAO (P27693) Alkaline protease precursor (EC 3.4.21.-)
Length = 380
Score = 58.5 bits (140), Expect = 4e-08
Identities = 50/189 (26%), Positives = 79/189 (41%), Gaps = 38/189 (20%)
Query: 364 KIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRT 423
K++ G + S+A+G E AG G+ + N L P +T+ N +
Sbjct: 203 KVLGASGSGSVSSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLEQAVNSA-- 251
Query: 424 TGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPV-----MASYSSRGPNKVQPSILKPDV 478
T R + ++ + SG PA+ N AS+S G D+
Sbjct: 252 TSRGVLVVAASGNSGAGSISYPARYANAMAVGATDQNNNRASFSQYGAGL--------DI 303
Query: 479 TAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPA 538
APGVN+ + Y G + + GTSM+ PHVAG A L+K +P+WS
Sbjct: 304 VAPGVNVQSTYP--------------GSTYASLNGTSMATPHVAGAAALVKQKNPSWSNV 349
Query: 539 AIKSAIRTT 547
I++ ++ T
Sbjct: 350 QIRNHLKNT 358
>SUBT_BACAM (P00782) Subtilisin BPN' precursor (EC 3.4.21.62)
(Subtilisin Novo) (Subtilisin DFE) (Alkaline protease)
Length = 382
Score = 54.7 bits (130), Expect = 6e-07
Identities = 44/136 (32%), Positives = 59/136 (43%), Gaps = 45/136 (33%)
Query: 412 STISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQP 471
ST+ YPG + PS I G + N+R AS+SS GP
Sbjct: 270 STVGYPGKY-----------PSVIAVGA------VDSSNQR------ASFSSVGPEL--- 303
Query: 472 SILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTL 531
DV APGV+I + T G + GTSM+ PHVAG A LI +
Sbjct: 304 -----DVMAPGVSIQS--------------TLPGNKYGAYNGTSMASPHVAGAAALILSK 344
Query: 532 HPNWSPAAIKSAIRTT 547
HPNW+ ++S++ T
Sbjct: 345 HPNWTNTQVRSSLENT 360
Score = 50.1 bits (118), Expect = 2e-05
Identities = 72/314 (22%), Positives = 120/314 (37%), Gaps = 73/314 (23%)
Query: 23 KQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQKGRF 82
K ++ +A L E+ ++ K+P V ++ ++H H G+ + +G
Sbjct: 72 KYVDAASATLNEKAVKELKKDPSVA--YVEEDHVAHAYAQSVPYGVSQIKAPALHSQGYT 129
Query: 83 GENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARF 142
G N + ID+G+ D + KV ++
Sbjct: 130 GSNVKVAVIDSGI---------------------------DSSHPDLKVAGGASMV---- 158
Query: 143 FNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARV 202
P +D HGTH T N G + G +P A +
Sbjct: 159 ------------PSETNPFQDNNSHGTHVAGTVAA--------LNNSIGVL-GVAPSASL 197
Query: 203 ATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAF 262
KV + D + + +++ I+ AI + +D+I++S GGPS + + + D+
Sbjct: 198 YAVKVLGA--DGSGQYSW-IINGIEWAIANNMDVINMSLGGPSGSAALKAAVDK------ 248
Query: 263 HALARNILLVASAGNEGPTPGSVVNVA-----PWVFTVAA---STLDRDFSSVMTIGNKT 314
A+A +++VA+AGNEG T GS V P V V A S FSSV +
Sbjct: 249 -AVASGVVVVAAAGNEG-TSGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVM 306
Query: 315 LTGASLFVNLPPNQ 328
G S+ LP N+
Sbjct: 307 APGVSIQSTLPGNK 320
>SUBT_BACST (P29142) Subtilisin J precursor (EC 3.4.21.62)
Length = 381
Score = 53.9 bits (128), Expect = 1e-06
Identities = 70/310 (22%), Positives = 118/310 (37%), Gaps = 71/310 (22%)
Query: 23 KQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQKGRF 82
K +N AA L+E+ ++ K+P V ++ ++H H G+ + +G
Sbjct: 71 KYVNAAAATLDEKAVKELKKDPSVA--YVEEDHIAHEYAQSVPYGISQIKAPALHSQGYT 128
Query: 83 GENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARF 142
G N + ID+G+ + RG GA F
Sbjct: 129 GSNVKVAVIDSGIDSSHPDLNVRG--------------------------------GASF 156
Query: 143 FNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARV 202
+P +D HGTH T N G + G SP A +
Sbjct: 157 -----------VPSETNPYQDGSSHGTHVAGTIAA--------LNNSIGVL-GVSPSASL 196
Query: 203 ATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAF 262
KV S + +++ I+ AI + +D+I++S GGPS + + + D+
Sbjct: 197 YAVKVLDSTGSGQYSW---IINGIEWAISNNMDVINMSLGGPSGSTALKTVVDK------ 247
Query: 263 HALARNILLVASAGNEGPTPGS--VVNVAPWVFTVAASTLD-----RDFSSVMTIGNKTL 315
A++ I++ A+AGNEG + S V A + T+A ++ FSS + +
Sbjct: 248 -AVSSGIVVAAAAGNEGSSGSSSTVGYPAKYPSTIAVGAVNSSNQRASFSSAGSELDVMA 306
Query: 316 TGASLFVNLP 325
G S+ LP
Sbjct: 307 PGVSIQSTLP 316
Score = 48.5 bits (114), Expect = 4e-05
Identities = 31/89 (34%), Positives = 42/89 (46%), Gaps = 22/89 (24%)
Query: 459 ASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSC 518
AS+SS G DV APGV+I + T G + GTSM+
Sbjct: 293 ASFSSAGSEL--------DVMAPGVSIQS--------------TLPGGTYGAYNGTSMAT 330
Query: 519 PHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
PHVAG A LI + HP W+ A ++ + +T
Sbjct: 331 PHVAGAAALILSKHPTWTNAQVRDRLEST 359
>SUBT_BACSA (P00783) Subtilisin amylosacchariticus precursor (EC
3.4.21.62)
Length = 381
Score = 53.9 bits (128), Expect = 1e-06
Identities = 70/310 (22%), Positives = 118/310 (37%), Gaps = 71/310 (22%)
Query: 23 KQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQKGRF 82
K +N AA L+E+ ++ K+P V ++ ++H H G+ + +G
Sbjct: 71 KYVNAAAATLDEKAVKELKKDPSVA--YVEEDHIAHEYAQSVPYGISQIKAPALHSQGYT 128
Query: 83 GENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARF 142
G N + ID+G+ + RG GA F
Sbjct: 129 GSNVKVAVIDSGIDSSHPDLNVRG--------------------------------GASF 156
Query: 143 FNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARV 202
+P +D HGTH T N G + G SP A +
Sbjct: 157 -----------VPSETNPYQDGSSHGTHVAGTIAA--------LNNSIGVL-GVSPSASL 196
Query: 203 ATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAF 262
KV S + +++ I+ AI + +D+I++S GGPS + + + D+
Sbjct: 197 YAVKVLDSTGSGQYSW---IINGIEWAISNNMDVINMSLGGPSGSTALKTVVDK------ 247
Query: 263 HALARNILLVASAGNEGPTPGS--VVNVAPWVFTVAASTLD-----RDFSSVMTIGNKTL 315
A++ I++ A+AGNEG + S V A + T+A ++ FSS + +
Sbjct: 248 -AVSSGIVVAAAAGNEGSSGSSSTVGYPAKYPSTIAVGAVNSSNQRASFSSAGSELDVMA 306
Query: 316 TGASLFVNLP 325
G S+ LP
Sbjct: 307 PGVSIQSTLP 316
Score = 48.5 bits (114), Expect = 4e-05
Identities = 31/89 (34%), Positives = 42/89 (46%), Gaps = 22/89 (24%)
Query: 459 ASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSC 518
AS+SS G DV APGV+I + T G + GTSM+
Sbjct: 293 ASFSSAGSEL--------DVMAPGVSIQS--------------TLPGGTYGAYNGTSMAT 330
Query: 519 PHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
PHVAG A LI + HP W+ A ++ + +T
Sbjct: 331 PHVAGAAALILSKHPTWTNAQVRDRLEST 359
>SUBN_BACNA (P35835) Subtilisin NAT precursor (EC 3.4.21.62)
Length = 381
Score = 53.1 bits (126), Expect = 2e-06
Identities = 70/311 (22%), Positives = 120/311 (38%), Gaps = 73/311 (23%)
Query: 23 KQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQKGRF 82
K +N AA L+E+ ++ K+P V ++ ++H H G+ + +G
Sbjct: 71 KYVNAAAATLDEKAVKELKKDPSVA--YVEEDHIAHEYAQSVPYGISQIKAPALHSQGYT 128
Query: 83 GENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARF 142
G N + ID+G+ + RG GA F
Sbjct: 129 GSNVKVAVIDSGIDSSHPDLNVRG--------------------------------GASF 156
Query: 143 FNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARV 202
+P +D HGTH T N G + G +P A +
Sbjct: 157 -----------VPSETNPYQDGSSHGTHVAGTIAA--------LNNSIGVL-GVAPSASL 196
Query: 203 ATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAF 262
KV S + +++ I+ AI + +D+I++S GGP+ + + + D+
Sbjct: 197 YAVKVLDSTGSGQYSW---IINGIEWAISNNMDVINMSLGGPTGSTALKTVVDK------ 247
Query: 263 HALARNILLVASAGNEGPTPGSVVNV---APWVFTVAASTLD-----RDFSSVMTIGNKT 314
A++ I++ A+AGNEG + GS V A + T+A ++ FSSV + +
Sbjct: 248 -AVSSGIVVAAAAGNEG-SSGSTSTVGYPAKYPSTIAVGAVNSSNQRASFSSVGSELDVM 305
Query: 315 LTGASLFVNLP 325
G S+ LP
Sbjct: 306 APGVSIQSTLP 316
Score = 47.8 bits (112), Expect = 7e-05
Identities = 26/71 (36%), Positives = 36/71 (50%), Gaps = 14/71 (19%)
Query: 477 DVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWS 536
DV APGV+I + T G + GTSM+ PHVAG A LI + HP W+
Sbjct: 303 DVMAPGVSIQS--------------TLPGGTYGAYNGTSMATPHVAGAAALILSKHPTWT 348
Query: 537 PAAIKSAIRTT 547
A ++ + +T
Sbjct: 349 NAQVRDRLEST 359
>SUBT_BACSU (P04189) Subtilisin E precursor (EC 3.4.21.62)
Length = 381
Score = 52.8 bits (125), Expect = 2e-06
Identities = 70/311 (22%), Positives = 119/311 (37%), Gaps = 73/311 (23%)
Query: 23 KQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQKGRF 82
K +N AA L+E+ ++ K+P V ++ ++H H G+ + +G
Sbjct: 71 KYVNAAAATLDEKAVKELKKDPSVA--YVEEDHIAHEYAQSVPYGISQIKAPALHSQGYT 128
Query: 83 GENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARF 142
G N + ID+G+ + RG GA F
Sbjct: 129 GSNVKVAVIDSGIDSSHPDLNVRG--------------------------------GASF 156
Query: 143 FNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARV 202
+P +D HGTH T N G + G SP A +
Sbjct: 157 -----------VPSETNPYQDGSSHGTHVAGTIAA--------LNNSIGVL-GVSPSASL 196
Query: 203 ATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAF 262
KV S + +++ I+ AI + +D+I++S GGP+ + + + D+
Sbjct: 197 YAVKVLDSTGSGQYSW---IINGIEWAISNNMDVINMSLGGPTGSTALKTVVDK------ 247
Query: 263 HALARNILLVASAGNEGPTPGSVVNV---APWVFTVAASTLD-----RDFSSVMTIGNKT 314
A++ I++ A+AGNEG + GS V A + T+A ++ FSS + +
Sbjct: 248 -AVSSGIVVAAAAGNEG-SSGSTSTVGYPAKYPSTIAVGAVNSSNQRASFSSAGSELDVM 305
Query: 315 LTGASLFVNLP 325
G S+ LP
Sbjct: 306 APGVSIQSTLP 316
Score = 48.5 bits (114), Expect = 4e-05
Identities = 31/89 (34%), Positives = 42/89 (46%), Gaps = 22/89 (24%)
Query: 459 ASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSC 518
AS+SS G DV APGV+I + T G + GTSM+
Sbjct: 293 ASFSSAGSEL--------DVMAPGVSIQS--------------TLPGGTYGAYNGTSMAT 330
Query: 519 PHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
PHVAG A LI + HP W+ A ++ + +T
Sbjct: 331 PHVAGAAALILSKHPTWTNAQVRDRLEST 359
>ISP1_BACSU (P11018) Major intracellular serine protease precursor
(EC 3.4.21.-) (ISP-1)
Length = 319
Score = 52.8 bits (125), Expect = 2e-06
Identities = 38/117 (32%), Positives = 55/117 (46%), Gaps = 18/117 (15%)
Query: 163 DFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADV 222
D+ GHGTH V G N NG I G +P A + KV + +
Sbjct: 83 DYNGHGTH---------VAGTIAANDSNGGIAGVAPEASLLIVKVLGGENGSGQYEW--I 131
Query: 223 LSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEG 279
++ I+ A++ VDIIS+S GGPS E+ +A+ +L+V +AGNEG
Sbjct: 132 INGINYAVEQKVDIISMSLGGPSD-------VPELKEAVKNAVKNGVLVVCAAGNEG 181
Score = 32.0 bits (71), Expect = 4.3
Identities = 19/54 (35%), Positives = 26/54 (47%), Gaps = 14/54 (25%)
Query: 477 DVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKT 530
D+ APG NIL+ T + + GTSM+ PHV+G LIK+
Sbjct: 222 DLVAPGENILS--------------TLPNKKYGKLTGTSMAAPHVSGALALIKS 261
>SUBT_BACS9 (P28842) Subtilisin precursor (EC 3.4.21.62)
Length = 420
Score = 49.3 bits (116), Expect = 3e-05
Identities = 44/150 (29%), Positives = 71/150 (47%), Gaps = 23/150 (15%)
Query: 157 SQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATS 216
+ + D GHGTH AG G + GNG + G +P A + YKV L D S
Sbjct: 172 TNNSCTDRQGHGTHV---AGSALADGGT----GNG-VYGVAPDADLWAYKV---LGDDGS 220
Query: 217 CFGADVLSAIDQAIDDGVD-----IISVSAGGPSSTNSEEIFTDEISIGAFHALARNILL 271
+ D+ +AI A D +I++S G S+ + T+ ++ ++ + +L+
Sbjct: 221 GYADDIAAAIRHAGDQATALNTKVVINMSLG---SSGESSLITNAVN----YSYNKGVLI 273
Query: 272 VASAGNEGPTPGSVVNVAPWVFTVAASTLD 301
+A+AGN GP GS+ V VA + L+
Sbjct: 274 IAAAGNSGPYQGSIGYPGALVNAVAVAALE 303
>SUBT_BACPU (P07518) Subtilisin (EC 3.4.21.62) (Alkaline
mesentericopeptidase)
Length = 275
Score = 48.9 bits (115), Expect = 3e-05
Identities = 41/136 (30%), Positives = 56/136 (41%), Gaps = 45/136 (33%)
Query: 412 STISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQP 471
ST+ YP + PS I G + N+R AS+SS G
Sbjct: 163 STVGYPAKY-----------PSTIAVGA------VNSANQR------ASFSSAGSEL--- 196
Query: 472 SILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTL 531
DV APGV+I + T G + GTSM+ PHVAG A LI +
Sbjct: 197 -----DVMAPGVSIQS--------------TLPGGTYGAYNGTSMATPHVAGAAALILSK 237
Query: 532 HPNWSPAAIKSAIRTT 547
HP W+ A ++ + +T
Sbjct: 238 HPTWTNAQVRDRLEST 253
Score = 42.7 bits (99), Expect = 0.002
Identities = 45/180 (25%), Positives = 79/180 (43%), Gaps = 28/180 (15%)
Query: 154 LPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTD 213
+P +D HGTH T N G + G +P + + KV S
Sbjct: 51 VPSETNPYQDGSSHGTHVAGTIAA--------LNNSIGVL-GVAPSSALYAVKVLDSTGS 101
Query: 214 ATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVA 273
+ +++ I+ AI + +D+I++S GGP+ + + + D+ A++ I++ A
Sbjct: 102 GQYSW---IINGIEWAISNNMDVINMSLGGPTGSTALKTVVDK-------AVSSGIVVAA 151
Query: 274 SAGNEGPTPGSVVNV---APWVFTVAASTLD-----RDFSSVMTIGNKTLTGASLFVNLP 325
+AGNEG + GS V A + T+A ++ FSS + + G S+ LP
Sbjct: 152 AAGNEG-SSGSTSTVGYPAKYPSTIAVGAVNSANQRASFSSAGSELDVMAPGVSIQSTLP 210
>SUBT_BACLI (P00780) Subtilisin Carlsberg precursor (EC 3.4.21.62)
Length = 379
Score = 48.9 bits (115), Expect = 3e-05
Identities = 46/175 (26%), Positives = 79/175 (44%), Gaps = 28/175 (16%)
Query: 166 GHGTHTLSTAGGNFVPGASIFNIGNGT-IKGGSPRARVATYKVCWSLTDATSCFGADVLS 224
GHGTH T + N T + G +P + KV L + S + ++S
Sbjct: 167 GHGTHVAGTVAA----------LDNTTGVLGVAPSVSLYAVKV---LNSSGSGTYSGIVS 213
Query: 225 AIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPT--P 282
I+ A +G+D+I++S GGPS + + + D +A AR +++VA+AGN G +
Sbjct: 214 GIEWATTNGMDVINMSLGGPSGSTAMKQAVD-------NAYARGVVVVAAAGNSGSSGNT 266
Query: 283 GSVVNVAPWVFTVAASTLDRD-----FSSVMTIGNKTLTGASLFVNLPPNQDFTI 332
++ A + +A +D + FSSV GA ++ P + T+
Sbjct: 267 NTIGYPAKYDSVIAVGAVDSNSNRASFSSVGAELEVMAPGAGVYSTYPTSTYATL 321
Score = 43.1 bits (100), Expect = 0.002
Identities = 23/71 (32%), Positives = 36/71 (50%), Gaps = 14/71 (19%)
Query: 477 DVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWS 536
+V APG + + Y A+ + GTSM+ PHVAG A LI + HPN S
Sbjct: 301 EVMAPGAGVYSTYPTSTYAT--------------LNGTSMASPHVAGAAALILSKHPNLS 346
Query: 537 PAAIKSAIRTT 547
+ +++ + +T
Sbjct: 347 ASQVRNRLSST 357
>SUBD_BACLI (P00781) Subtilisin DY (EC 3.4.21.62)
Length = 274
Score = 48.5 bits (114), Expect = 4e-05
Identities = 46/170 (27%), Positives = 76/170 (44%), Gaps = 28/170 (16%)
Query: 166 GHGTHTLSTAGGNFVPGASIFNIGNGT-IKGGSPRARVATYKVCWSLTDATSCFGADVLS 224
GHGTH T + N T + G +P + KV L + S + ++S
Sbjct: 62 GHGTHVAGTVAA----------LDNTTGVLGVAPNVSLYAIKV---LNSSGSGTYSAIVS 108
Query: 225 AIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGP--TP 282
I+ A +G+D+I++S GGPS + + + D+ A A I++VA+AGN G +
Sbjct: 109 GIEWATQNGLDVINMSLGGPSGSTALKQAVDK-------AYASGIVVVAAAGNSGSSGSQ 161
Query: 283 GSVVNVAPWVFTVAASTLDRD-----FSSVMTIGNKTLTGASLFVNLPPN 327
++ A + +A +D + FSSV G S++ P N
Sbjct: 162 NTIGYPAKYDSVIAVGAVDSNKNRASFSSVGAELEVMAPGVSVYSTYPSN 211
Score = 41.2 bits (95), Expect = 0.007
Identities = 24/71 (33%), Positives = 38/71 (52%), Gaps = 14/71 (19%)
Query: 477 DVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWS 536
+V APGV++ + Y SN T + GTSM+ PHVAG A LI + +P S
Sbjct: 196 EVMAPGVSVYSTYP-----SNTYTS---------LNGTSMASPHVAGAAALILSKYPTLS 241
Query: 537 PAAIKSAIRTT 547
+ +++ + +T
Sbjct: 242 ASQVRNRLSST 252
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.132 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,616,453
Number of Sequences: 164201
Number of extensions: 2810353
Number of successful extensions: 6664
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 45
Number of HSP's that attempted gapping in prelim test: 6502
Number of HSP's gapped (non-prelim): 148
length of query: 547
length of database: 59,974,054
effective HSP length: 115
effective length of query: 432
effective length of database: 41,090,939
effective search space: 17751285648
effective search space used: 17751285648
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC133779.5


