

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
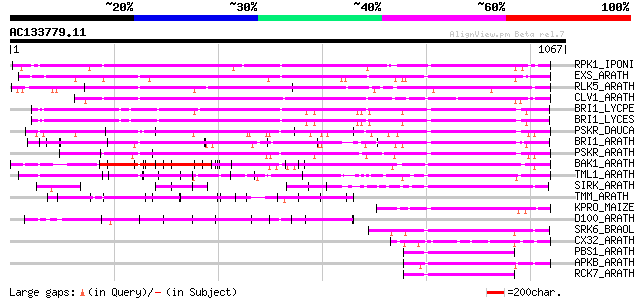
Query= AC133779.11 - phase: 0
(1067 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 545 e-154
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 479 e-134
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 478 e-134
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 433 e-120
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 409 e-113
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 407 e-113
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 399 e-110
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 389 e-107
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 385 e-106
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 222 4e-57
TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor 207 9e-53
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 170 2e-41
TMM_ARATH (Q9SSD1) TOO MANY MOUTHS protein precursor (TMM) 157 1e-37
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 157 2e-37
D100_ARATH (Q00874) DNA-damage-repair/toleration protein DRT100 ... 153 3e-36
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 146 3e-34
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 144 1e-33
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 144 1e-33
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 144 2e-33
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 143 3e-33
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 545 bits (1403), Expect = e-154
Identities = 379/1120 (33%), Positives = 556/1120 (48%), Gaps = 120/1120 (10%)
Query: 6 TLIMILCVLPTLSVA----EDSEAKLALLK-WKDSFDDQSQTLLSTWK-NNTNPCKPKWR 59
T ++ LC ++ A D A L+L + W D +Q+ W +++ PC W
Sbjct: 7 TFLLFLCSTSSIYAAFALNSDGAALLSLTRHWTSIPSDITQS----WNASDSTPCS--WL 60
Query: 60 GIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNIS 119
G++CD+ F+ T+ L++ G+ G S +L + + N F+G+IP+Q+GN S +
Sbjct: 61 GVECDRRQFVDTLNLSSYGISGEFGP-EISHLKHLKKVVLSGNGFFGSIPSQLGNCSLLE 119
Query: 120 ILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKL------------------------NGA 155
+ +N F G+IP + L L+ L + F L NG+
Sbjct: 120 HIDLSSNSFTGNIPDTLGALQNLRNLSLFFNSLIGPFPESLLSIPHLETVYFTGNGLNGS 179
Query: 156 IPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNL 215
IP +IGN++ L+ L L N +SG P+P +G + L L + +NLVG++P + L NL
Sbjct: 180 IPSNIGNMSELTTLWLDDNQFSG-PVPSSLGNITTLQELYLNDNNLVGTLPVTLNNLENL 238
Query: 216 AYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLS 275
Y+D+ NSL G IP + ++DT+ LSNN + +G +P L N +SL + LS
Sbjct: 239 VYLDVRNNSLVGAIPLDFVSCKQIDTISLSNN-QFTGGLPPGLGNCTSLREFGAFSCALS 297
Query: 276 GSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLIN 335
G IP L L L L NH SG IP +G K++I L L N L G IP +G L
Sbjct: 298 GPIPSCFGQLTKLDTLYLAGNHFSGRIPPELGKCKSMIDLQLQQNQLEGEIPGELGMLSQ 357
Query: 336 LQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFV 395
LQ L + NNL+G +P SI ++ L ++ N L G +P + + +S + EN F
Sbjct: 358 LQYLHLYTNNLSGEVPLSIWKIQSLQSLQLYQNNLSGELPVDMTELKQLVSLALYENHFT 417
Query: 396 GHLPSQICSGGSLRLLNADHNRFTGPIPTSLKT------------------------CSS 431
G +P + + SL +L+ N FTG IP +L + CS+
Sbjct: 418 GVIPQDLGANSSLEVLDLTRNMFTGHIPPNLCSQKKLKRLLLGYNYLEGSVPSDLGGCST 477
Query: 432 IERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNIS 491
+ER+ LE N + G + DF L + DLS N F G I P+ G N+ +S+N +S
Sbjct: 478 LERLILEENNLRGGLP-DFVEKQNLLFFDLSGNNFTGPIPPSLGNLKNVTAIYLSSNQLS 536
Query: 492 GVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQ 551
G IP + L KL L+LS N L G LP E L L +L S+N + +IPS +G L
Sbjct: 537 GSIPPELGSLVKLEHLNLSHNILKGILPSE-LSNCHKLSELDASHNLLNGSIPSTLGSLT 595
Query: 552 RLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIP-IKFDSGLESLDLSGNFLK 610
L +L LG N SG IP L + L L L N + G IP + L SL+LS N L
Sbjct: 596 ELTKLSLGENSFSGGIPTSLFQSNKLLNLQLGGNLLAGDIPPVGALQALRSLNLSSNKLN 655
Query: 611 GNIPTGLADLVRLSKLNLSHNMLSGTIPQ-NFGRNLVFVNISDNQLEGPLP-KIPAFLSA 668
G +P L L L +L++SHN LSGT+ + ++L F+NIS N GP+P + FL++
Sbjct: 656 GQLPIDLGKLKMLEELDVSHNNLSGTLRVLSTIQSLTFINISHNLFSGPVPPSLTKFLNS 715
Query: 669 SFESLKNNNHLCGNIRG----------LDPC--ATSHSRKRKNVLRPVFIALGAVILVLC 716
S S N+ LC N L PC ++ + + L I LGA++ ++C
Sbjct: 716 SPTSFSGNSDLCINCPADGLACPESSILRPCNMQSNTGKGGLSTLGIAMIVLGALLFIIC 775
Query: 717 VV--GALMYIMCGRKKPNEESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVG 774
+ A +++ C KK +E + S DG ++ ++EAT N +DKY++G
Sbjct: 776 LFLFSAFLFLHC--KKSVQE---------IAISAQEGDGSLL-NKVLEATENLNDKYVIG 823
Query: 775 VGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGF 834
G+ G +YKA LS V AVKKL + S S + EIET+ ++HRN+IKL F
Sbjct: 824 KGAHGTIYKATLSPDKVYAVKKLVFTGIKN----GSVSMVREIETIGKVRHRNLIKLEEF 879
Query: 835 CSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPII 894
++ ++Y ++E GSL IL+ DW R N+ G A+ L+YLH DC P I+
Sbjct: 880 WLRKEYGLILYTYMENGSLHDILHETNPPKPLDWSTRHNIAVGTAHGLAYLHFDCDPAIV 939
Query: 895 HRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHS--WTQFAGTFGYAAPELAQTMEVNEK 952
HRDI N+LL+ D E H+SDFG AK L S GT GY APE A T + +
Sbjct: 940 HRDIKPMNILLDSDLEPHISDFGIAKLLDQSATSIPSNTVQGTIGYMAPENAFTTVKSRE 999
Query: 953 CDVYSFGVLALETIMGKH--------PGDLISLFLSPST-----RPMANNMLLTDVLDQR 999
DVYS+GV+ LE I K D++ S T + + + LL +++D
Sbjct: 1000 SDVYSYGVVLLELITRKKALDPSFNGETDIVGWVRSVWTQTGEIQKIVDPSLLDELID-- 1057
Query: 1000 PQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
VME + E + LA C + RP+M V K L
Sbjct: 1058 -SSVMEQVTEAL----SLALRCAEKEVDKRPTMRDVVKQL 1092
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 479 bits (1234), Expect = e-134
Identities = 376/1192 (31%), Positives = 541/1192 (44%), Gaps = 191/1192 (16%)
Query: 18 SVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSNFISTIGLANL 77
++ + S +L+ +K S ++ S LLS+W +++ W G+ C ++++ L +L
Sbjct: 19 AIVDLSSETTSLISFKRSLENPS--LLSSWNVSSSASHCDWVGVTCLLGR-VNSLSLPSL 75
Query: 78 GLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFK------------- 124
L+G + SS NL + + N F G IP +I NL ++ L
Sbjct: 76 SLRGQIPK-EISSLKNLRELCLAGNQFSGKIPPEIWNLKHLQTLDLSGNSLTGLLPRLLS 134
Query: 125 -----------NNYFDGSIPQEM-CTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILG 172
+N+F GS+P +L L LD+S L+G IP IG L+NLS L +G
Sbjct: 135 ELPQLLYLDLSDNHFSGSLPPSFFISLPALSSLDVSNNSLSGEIPPEIGKLSNLSNLYMG 194
Query: 173 GNNWSG-----------------------GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEI 209
N++SG GP+P EI KL +L L + + L SIP+
Sbjct: 195 LNSFSGQIPSEIGNISLLKNFAAPSCFFNGPLPKEISKLKHLAKLDLSYNPLKCSIPKSF 254
Query: 210 GFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTV--- 266
G L NL+ ++L L G IP +GN L +L+LS N+ +SGP+P L + LT
Sbjct: 255 GELHNLSILNLVSAELIGLIPPELGNCKSLKSLMLSFNS-LSGPLPLELSEIPLLTFSAE 313
Query: 267 --------------------LYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTI 306
L N SG IP I++ LK L+L N LSGSIP +
Sbjct: 314 RNQLSGSLPSWMGKWKVLDSLLLANNRFSGEIPHEIEDCPMLKHLSLASNLLSGSIPREL 373
Query: 307 GDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVA 366
+L + L N LSG I +L L + N + G+IP + L L ++
Sbjct: 374 CGSGSLEAIDLSGNLLSGTIEEVFDGCSSLGELLLTNNQINGSIPEDLWKLP-LMALDLD 432
Query: 367 TNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSL 426
+N G IP L+ TN + F S N G+LP++I + SL+ L N+ TG IP +
Sbjct: 433 SNNFTGEIPKSLWKSTNLMEFTASYNRLEGYLPAEIGNAASLKRLVLSDNQLTGEIPREI 492
Query: 427 KTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIIS 486
+S+ + L N +G I + G L LDL N GQI LQ ++S
Sbjct: 493 GKLTSLSVLNLNANMFQGKIPVELGDCTSLTTLDLGSNNLQGQIPDKITALAQLQCLVLS 552
Query: 487 NNNISGVIPL------------DFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKI 534
NN+SG IP D L G+ LS N+L+G +P E LG L ++ +
Sbjct: 553 YNNLSGSIPSKPSAYFHQIEMPDLSFLQHHGIFDLSYNRLSGPIPEE-LGECLVLVEISL 611
Query: 535 SNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIK 594
SNNH S IP+ + L L LDL GN L+G IPKE+ L+ LNL+ N++ G IP
Sbjct: 612 SNNHLSGEIPASLSRLTNLTILDLSGNALTGSIPKEMGNSLKLQGLNLANNQLNGHIPES 671
Query: 595 FD--SGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGT---------------- 636
F L L+L+ N L G +P L +L L+ ++LS N LSG
Sbjct: 672 FGLLGSLVKLNLTKNKLDGPVPASLGNLKELTHMDLSFNNLSGELSSELSTMEKLVGLYI 731
Query: 637 --------IPQNFGR--------------------------NLVFVNISDNQLEGPLPKI 662
IP G NL F+N++ N L G +P
Sbjct: 732 EQNKFTGEIPSELGNLTQLEYLDVSENLLSGEIPTKICGLPNLEFLNLAKNNLRGEVPSD 791
Query: 663 PAFLSASFESLKNNNHLCGNIRGLDPCATSHSRKRKNVLRPVFIALGAVILVLCVVGALM 722
S L N LCG + G D K ++ + LG I+V V +L
Sbjct: 792 GVCQDPSKALLSGNKELCGRVVGSD--CKIEGTKLRSAWGIAGLMLGFTIIVFVFVFSLR 849
Query: 723 -YIMCGRKKPNEESQTEE-------VQRGVLFSIWSHDGK------MMFE---------N 759
+ M R K ++ + E V + + F S + MFE +
Sbjct: 850 RWAMTKRVKQRDDPERMEESRLKGFVDQNLYFLSGSRSREPLSINIAMFEQPLLKVRLGD 909
Query: 760 IIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIET 819
I+EAT +F K ++G G G VYKA L VAVKKL E ++ FM+E+ET
Sbjct: 910 IVEATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKL-----SEAKTQGNREFMAEMET 964
Query: 820 LTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAV-AFDWEKRVNVVKGV 878
L +KH N++ L G+CS S+ LVY+++ GSLD L N T + DW KR+ + G
Sbjct: 965 LGKVKHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIAVGA 1024
Query: 879 ANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKP-GLHSWTQFAGTFG 937
A L++LHH P IIHRDI + N+LL+ D+E V+DFG A+ + H T AGTFG
Sbjct: 1025 ARGLAFLHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTVIAGTFG 1084
Query: 938 YAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP----------GDLISLFLSPSTRPMA 987
Y PE Q+ K DVYSFGV+ LE + GK P G+L+ + + A
Sbjct: 1085 YIPPEYGQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGGNLVGWAIQKINQGKA 1144
Query: 988 NNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
DV+D P V + + + ++A CL++ P RP+M V K L
Sbjct: 1145 -----VDVID--PLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKAL 1189
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 478 bits (1231), Expect = e-134
Identities = 336/919 (36%), Positives = 494/919 (53%), Gaps = 48/919 (5%)
Query: 145 LDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGS 204
+D+S L G P + +L +L L L N+ +G + +NL+ L + ++ LVGS
Sbjct: 70 VDLSSFMLVGPFPSILCHLPSLHSLSLYNNSINGSLSADDFDTCHNLISLDLSENLLVGS 129
Query: 205 IPQEIGF-LTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSS 263
IP+ + F L NL ++++S N+LS IP + G KL++L L+ N +SG IP SL N+++
Sbjct: 130 IPKSLPFNLPNLKFLEISGNNLSDTIPSSFGEFRKLESLNLAGNF-LSGTIPASLGNVTT 188
Query: 264 LTVLYFD-NIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNL 322
L L N+ IP + NL L+ L L +L G IP ++ L +L+ L L N L
Sbjct: 189 LKELKLAYNLFSPSQIPSQLGNLTELQVLWLAGCNLVGPIPPSLSRLTSLVNLDLTFNQL 248
Query: 323 SGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNIT 382
+G IP+ I L ++ + + N+ +G +P S+GN+ L F+ + NKL G+IP+ L N+
Sbjct: 249 TGSIPSWITQLKTVEQIELFNNSFSGELPESMGNMTTLKRFDASMNKLTGKIPDNL-NLL 307
Query: 383 NWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQI 442
N S + EN G LP I +L L +NR TG +P+ L S ++ + L N+
Sbjct: 308 NLESLNLFENMLEGPLPESITRSKTLSELKLFNNRLTGVLPSQLGANSPLQYVDLSYNRF 367
Query: 443 EGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLT 502
G+I + KL+YL L DN F G+IS N GK +L +SNN +SG IP F GL
Sbjct: 368 SGEIPANVCGEGKLEYLILIDNSFSGEISNNLGKCKSLTRVRLSNNKLSGQIPHGFWGLP 427
Query: 503 KLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNE 562
+L +L LS N TG +P ++G K+L +L+IS N FS +IP+EIG L + E+ N+
Sbjct: 428 RLSLLELSDNSFTGSIPKTIIGA-KNLSNLRISKNRFSGSIPNEIGSLNGIIEISGAEND 486
Query: 563 LSGKIPKELVELPNLRMLNLSRNKIEGIIP--IKFDSGLESLDLSGNFLKGNIPTGLADL 620
SG+IP+ LV+L L L+LS+N++ G IP ++ L L+L+ N L G IP + L
Sbjct: 487 FSGEIPESLVKLKQLSRLDLSKNQLSGEIPRELRGWKNLNELNLANNHLSGEIPKEVGIL 546
Query: 621 VRLSKLNLSHNMLSGTIP---QNFGRNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNN 677
L+ L+LS N SG IP QN N++ N+S N L G +P + A + + + N
Sbjct: 547 PVLNYLDLSSNQFSGEIPLELQNLKLNVL--NLSYNHLSGKIPPLYANKIYAHDFIGNPG 604
Query: 678 HLCGNIRGLDPCATSHSRKRKNV-----LRPVFIALGAVILVLCVVGALMYIMCGRKKPN 732
LC ++ GL T + KN+ L +F+ G V VVG +M+I RK
Sbjct: 605 -LCVDLDGLCRKIT----RSKNIGYVWILLTIFLLAGLVF----VVGIVMFIAKCRKLRA 655
Query: 733 EESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVV 792
+S T + S W K+ F E D+K ++G GS G VYK EL G VV
Sbjct: 656 LKSST------LAASKWRSFHKLHFSEH-EIADCLDEKNVIGFGSSGKVYKVELRGGEVV 708
Query: 793 AVKKLHLVTDEEMSCFSSKS-----FMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKF 847
AVKKL+ +SS S F +E+ETL I+H++I++L CS LVY++
Sbjct: 709 AVKKLNKSVKGGDDEYSSDSLNRDVFAAEVETLGTIRHKSIVRLWCCCSSGDCKLLVYEY 768
Query: 848 LEGGSLDQILNNDTQA-VAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLN 906
+ GSL +L+ D + V W +R+ + A LSYLHHDC PPI+HRD+ S N+LL+
Sbjct: 769 MPNGSLADVLHGDRKGGVVLGWPERLRIALDAAEGLSYLHHDCVPPIVHRDVKSSNILLD 828
Query: 907 LDYEAHVSDFGTAKFLKPG----LHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLA 962
DY A V+DFG AK + + + AG+ GY APE T+ VNEK D+YSFGV+
Sbjct: 829 SDYGAKVADFGIAKVGQMSGSKTPEAMSGIAGSCGYIAPEYVYTLRVNEKSDIYSFGVVL 888
Query: 963 LETIMGKHPGD--LISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFA 1020
LE + GK P D L ++ + L V+D + + EE+ + +
Sbjct: 889 LELVTGKQPTDSELGDKDMAKWVCTALDKCGLEPVIDPKLDLKFK---EEISKVIHIGLL 945
Query: 1021 CLSQNPRLRPSMGQVCKML 1039
C S P RPSM +V ML
Sbjct: 946 CTSPLPLNRPSMRKVVIML 964
Score = 268 bits (684), Expect = 8e-71
Identities = 195/599 (32%), Positives = 300/599 (49%), Gaps = 69/599 (11%)
Query: 3 VLPTLIMILCV----LPTLSVAEDS----EAKLALLKWKDSFDDQSQTLLSTWKNNTNPC 54
+L LI++LC+ LP+LS+ +D+ +AKL L D +Q+L S+W +N +
Sbjct: 1 MLYCLILLLCLSSTYLPSLSLNQDATILRQAKLGL-------SDPAQSL-SSWSDNNDVT 52
Query: 55 KPKWRGIKCDKSNFISTIGLANLGLKG----------TLHSLT--------------FSS 90
KW G+ CD ++ + ++ L++ L G +LHSL+ F +
Sbjct: 53 PCKWLGVSCDATSNVVSVDLSSFMLVGPFPSILCHLPSLHSLSLYNNSINGSLSADDFDT 112
Query: 91 FPNLLMIDIRNNSFYGTIPAQIG-NLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISF 149
NL+ +D+ N G+IP + NL N+ L N +IP L+ L+++
Sbjct: 113 CHNLISLDLSENLLVGSIPKSLPFNLPNLKFLEISGNNLSDTIPSSFGEFRKLESLNLAG 172
Query: 150 CKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEI 209
L+G IP S+GN+T L L L N +S IP ++G L L L + NLVG IP +
Sbjct: 173 NFLSGTIPASLGNVTTLKELKLAYNLFSPSQIPSQLGNLTELQVLWLAGCNLVGPIPPSL 232
Query: 210 GFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYF 269
LT+L +DL+ N L+G IP I L ++ + L NN+ SG +P S+ NM++L
Sbjct: 233 SRLTSLVNLDLTFNQLTGSIPSWITQLKTVEQIELFNNS-FSGELPESMGNMTTLKRFDA 291
Query: 270 DNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPAS 329
L+G IPD++ NL+NL+ L L N L G +P +I K L +L L +N L+G +P+
Sbjct: 292 SMNKLTGKIPDNL-NLLNLESLNLFENMLEGPLPESITRSKTLSELKLFNNRLTGVLPSQ 350
Query: 330 IGNLINLQVLSVQENNLTGTIPASI------------------------GNLKWLTVFEV 365
+G LQ + + N +G IPA++ G K LT +
Sbjct: 351 LGANSPLQYVDLSYNRFSGEIPANVCGEGKLEYLILIDNSFSGEISNNLGKCKSLTRVRL 410
Query: 366 ATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTS 425
+ NKL G+IP+G + + +S+N F G +P I +L L NRF+G IP
Sbjct: 411 SNNKLSGQIPHGFWGLPRLSLLELSDNSFTGSIPKTIIGAKNLSNLRISKNRFSGSIPNE 470
Query: 426 LKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFII 485
+ + + I I+ N G+I + +L LDLS N+ G+I NL +
Sbjct: 471 IGSLNGIIEISGAENDFSGEIPESLVKLKQLSRLDLSKNQLSGEIPRELRGWKNLNELNL 530
Query: 486 SNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIP 544
+NN++SG IP + L L L LSSNQ +G++P+E L +K L L +S NH S IP
Sbjct: 531 ANNHLSGEIPKEVGILPVLNYLDLSSNQFSGEIPLE-LQNLK-LNVLNLSYNHLSGKIP 587
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 433 bits (1114), Expect = e-120
Identities = 311/944 (32%), Positives = 483/944 (50%), Gaps = 52/944 (5%)
Query: 125 NNYFDGSIPQEMCTLTGLQF--------LDISFCKLNGAIPKSIGNLTNLSYLILGGNNW 176
+++ S P C+ +G+ L++SF L G I IG LT+L L L NN+
Sbjct: 47 HDWIHSSSPDAHCSFSGVSCDDDARVISLNVSFTPLFGTISPEIGMLTHLVNLTLAANNF 106
Query: 177 SGGPIPPEIGKLNNLLHLAIQKS-NLVGSIPQEI-GFLTNLAYIDLSKNSLSGGIPETIG 234
+G +P E+ L +L L I + NL G+ P EI + +L +D N+ +G +P +
Sbjct: 107 TG-ELPLEMKSLTSLKVLNISNNGNLTGTFPGEILKAMVDLEVLDTYNNNFNGKLPPEMS 165
Query: 235 NLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALD 294
L KL L N SG IP S ++ SL L + GLSG P + L NL+E+ +
Sbjct: 166 ELKKLKYLSFGGNF-FSGEIPESYGDIQSLEYLGLNGAGLSGKSPAFLSRLKNLREMYIG 224
Query: 295 I-NHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPAS 353
N +G +P G L L L + S L+G IP S+ NL +L L + NNLTG IP
Sbjct: 225 YYNSYTGGVPPEFGGLTKLEILDMASCTLTGEIPTSLSNLKHLHTLFLHINNLTGHIPPE 284
Query: 354 IGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNA 413
+ L L +++ N+L G IP N+ N + N+ G +P I L +
Sbjct: 285 LSGLVSLKSLDLSINQLTGEIPQSFINLGNITLINLFRNNLYGQIPEAIGELPKLEVFEV 344
Query: 414 DHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPN 473
N FT +P +L ++ ++ + N + G I +D KL+ L LS+N F G I
Sbjct: 345 WENNFTLQLPANLGRNGNLIKLDVSDNHLTGLIPKDLCRGEKLEMLILSNNFFFGPIPEE 404
Query: 474 WGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLK 533
GK +L I N ++G +P L + ++ L+ N +G+LP+ + G + L +
Sbjct: 405 LGKCKSLTKIRIVKNLLNGTVPAGLFNLPLVTIIELTDNFFSGELPVTMSGDV--LDQIY 462
Query: 534 ISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIP- 592
+SNN FS IP IG LQ L L N G IP+E+ EL +L +N S N I G IP
Sbjct: 463 LSNNWFSGEIPPAIGNFPNLQTLFLDRNRFRGNIPREIFELKHLSRINTSANNITGGIPD 522
Query: 593 -IKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGR--NLVFVN 649
I S L S+DLS N + G IP G+ ++ L LN+S N L+G+IP G +L ++
Sbjct: 523 SISRCSTLISVDLSRNRINGEIPKGINNVKNLGTLNISGNQLTGSIPTGIGNMTSLTTLD 582
Query: 650 ISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHSRKRKNVLRPVFIALG 709
+S N L G +P FL + S N +LC R C T + + +F
Sbjct: 583 LSFNDLSGRVPLGGQFLVFNETSFAGNTYLCLPHRV--SCPTRPGQTSDHNHTALFSPSR 640
Query: 710 AVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFSIWSHDGKMMF--ENIIEATANF 767
VI V+ + L+ I ++ N++ Q+ + + + + K+ F E+++E
Sbjct: 641 IVITVIAAITGLILISVAIRQMNKKKN----QKSLAWKLTAFQ-KLDFKSEDVLEC---L 692
Query: 768 DDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRN 827
++ ++G G G VY+ + + VA+K+L + S F +EI+TL I+HR+
Sbjct: 693 KEENIIGKGGAGIVYRGSMPNNVDVAIKRLV----GRGTGRSDHGFTAEIQTLGRIRHRH 748
Query: 828 IIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHH 887
I++L G+ ++ + L+Y+++ GSL ++L+ ++ WE R V A L YLHH
Sbjct: 749 IVRLLGYVANKDTNLLLYEYMPNGSLGELLHG-SKGGHLQWETRHRVAVEAAKGLCYLHH 807
Query: 888 DCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHS--WTQFAGTFGYAAPELAQ 945
DCSP I+HRD+ S N+LL+ D+EAHV+DFG AKFL G S + AG++GY APE A
Sbjct: 808 DCSPLILHRDVKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAPEYAY 867
Query: 946 TMEVNEKCDVYSFGVLALETIMGKHP----GDLISLFL------SPSTRPMANNMLLTDV 995
T++V+EK DVYSFGV+ LE I GK P G+ + + T+P ++ ++ +
Sbjct: 868 TLKVDEKSDVYSFGVVLLELIAGKKPVGEFGEGVDIVRWVRNTEEEITQP-SDAAIVVAI 926
Query: 996 LDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
+D P+ P+ VI + ++A C+ + RP+M +V ML
Sbjct: 927 VD--PRLTGYPL-TSVIHVFKIAMMCVEEEAAARPTMREVVHML 967
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 409 bits (1051), Expect = e-113
Identities = 345/1128 (30%), Positives = 535/1128 (46%), Gaps = 158/1128 (14%)
Query: 42 TLLSTWKNNTNPCKPKWRGIKCDKSNFISTIGLANLGLKGTLHSLTFSSFP--NLLMIDI 99
TLL W ++T+PC + G+ C K++ +S+I L+N L +T P NL + +
Sbjct: 59 TLLQNWLSSTDPCS--FTGVSC-KNSRVSSIDLSNTFLSVDFSLVTSYLLPLSNLESLVL 115
Query: 100 RNNSFYGTIPAQIGNLSNISI--LTFKNNYFDGSIPQ--EMCTLTGLQFLDISFCKLNGA 155
+N + G++ + + +++ + N G I + L+ L++S L+
Sbjct: 116 KNANLSGSLTSAAKSQCGVTLDSIDLAENTISGPISDISSFGVCSNLKSLNLSKNFLDPP 175
Query: 156 IPKSIGNLT-NLSYLILGGNNWSGGPIPPEIGKLN--NLLHLAIQKSNLVGSIPQEIGFL 212
+ + T +L L L NN SG + P + + L +I+ + L GSIP E+ F
Sbjct: 176 GKEMLKGATFSLQVLDLSYNNISGFNLFPWVSSMGFVELEFFSIKGNKLAGSIP-ELDF- 233
Query: 213 TNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNI 272
NL+Y+DLS N+ S P + + S L L LS+N K G I SL + L+ L N
Sbjct: 234 KNLSYLDLSANNFSTVFP-SFKDCSNLQHLDLSSN-KFYGDIGSSLSSCGKLSFLNLTNN 291
Query: 273 GLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDL-KNLIKLYLGSNNLSGPIPASIG 331
G +P +L+ L L N G P+ + DL K +++L L NN SG +P S+G
Sbjct: 292 QFVGLVPKLPSE--SLQYLYLRGNDFQGVYPNQLADLCKTVVELDLSYNNFSGMVPESLG 349
Query: 332 NLINLQVLSVQENNLTGTIPAS----IGNLKWLTVFEVATNKLHGRIPNGLYNITNWISF 387
+L+++ + NN +G +P + N+K + + + NK G +P+ N+ +
Sbjct: 350 ECSSLELVDISNNNFSGKLPVDTLLKLSNIKTMVL---SFNKFVGGLPDSFSNLPKLETL 406
Query: 388 VVSENDFVGHLPSQICSG--GSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGD 445
+S N+ G +PS IC +L++L +N F GPIP SL CS + + L N + G
Sbjct: 407 DMSSNNLTGIIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSNCSQLVSLDLSFNYLTGS 466
Query: 446 IAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLG 505
I G KL+ L L N+ G+I L+ I+ N+++G IP TKL
Sbjct: 467 IPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPIPASLSNCTKLN 526
Query: 506 VLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSG 565
+ LS+NQL+G++P LG + +L LK+ NN S NIP+E+G Q L LDL N L+G
Sbjct: 527 WISLSNNQLSGEIPAS-LGRLSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFLNG 585
Query: 566 KIPK----------------------------------ELVELPNLRMLNLSR------- 584
IP L+E +R L R
Sbjct: 586 SIPPPLFKQSGNIAVALLTGKRYVYIKNDGSKECHGAGNLLEFGGIRQEQLDRISTRHPC 645
Query: 585 ---NKIEGIIPIKFD--SGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQ 639
GI F+ + LDLS N L+G+IP L + LS LNL HN LSG IPQ
Sbjct: 646 NFTRVYRGITQPTFNHNGSMIFLDLSYNKLEGSIPKELGAMYYLSILNLGHNDLSGMIPQ 705
Query: 640 NFG--RNLVFVNISDNQLEGPLPKIPAFL-------------------SASFESLKN--- 675
G +N+ +++S N+ G +P L SA F++ +
Sbjct: 706 QLGGLKNVAILDLSYNRFNGTIPNSLTSLTLLGEIDLSNNNLSGMIPESAPFDTFPDYRF 765
Query: 676 -NNHLCGNIRGLDPC-------ATSHSRK-RKNVLRPVFIALGAVILVLCVVGALMY-IM 725
NN LCG L PC A H + R+ +A+G + + C+ G ++ I
Sbjct: 766 ANNSLCGYPLPL-PCSSGPKSDANQHQKSHRRQASLAGSVAMGLLFSLFCIFGLIIVAIE 824
Query: 726 CGRKKPNEESQTEEVQRGVLFSIWSHDG----------------------KMMFENIIEA 763
+++ +E+ E G S ++ K+ F +++EA
Sbjct: 825 TKKRRRKKEAALEAYMDGHSHSATANSAWKFTSAREALSINLAAFEKPLRKLTFADLLEA 884
Query: 764 TANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGI 823
T F + LVG G G+VYKA+L +G VVA+KKL V+ + + F +E+ET+ I
Sbjct: 885 TNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQ-----GDREFTAEMETIGKI 939
Query: 824 KHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQA-VAFDWEKRVNVVKGVANAL 882
KHRN++ L G+C + LVY++++ GSL+ +L++ + + +W R + G A L
Sbjct: 940 KHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKIAIGAARGL 999
Query: 883 SYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKP-GLH-SWTQFAGTFGYAA 940
++LHH+C P IIHRD+ S NVLL+ + EA VSDFG A+ + H S + AGT GY
Sbjct: 1000 AFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGYVP 1059
Query: 941 PELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNML--------- 991
PE Q+ + K DVYS+GV+ LE + GK P D S NN++
Sbjct: 1060 PEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTD--------SADFGDNNLVGWVKLHAKG 1111
Query: 992 -LTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKM 1038
+TDV D+ + I+ E++ ++A ACL RP+M QV M
Sbjct: 1112 KITDVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAM 1159
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 407 bits (1046), Expect = e-113
Identities = 342/1125 (30%), Positives = 533/1125 (46%), Gaps = 152/1125 (13%)
Query: 42 TLLSTWKNNTNPCKPKWRGIKCDKSNFISTIGLANLGLKGTLHSLTFSSFP--NLLMIDI 99
TLL W ++T PC + G+ C K++ +S+I L+N L +T P NL + +
Sbjct: 59 TLLQNWLSSTGPCS--FTGVSC-KNSRVSSIDLSNTFLSVDFSLVTSYLLPLSNLESLVL 115
Query: 100 RNNSFYGTIPAQIGNLSNISI--LTFKNNYFDGSIPQ--EMCTLTGLQFLDISFCKLNGA 155
+N + G++ + + +++ + N G I + L+ L++S L+
Sbjct: 116 KNANLSGSLTSAAKSQCGVTLDSIDLAENTISGPISDISSFGVCSNLKSLNLSKNFLDPP 175
Query: 156 IPKSIGNLT-NLSYLILGGNNWSGGPIPPEIGKLN--NLLHLAIQKSNLVGSIPQEIGFL 212
+ + T +L L L NN SG + P + + L +++ + L GSIP E+ F
Sbjct: 176 GKEMLKAATFSLQVLDLSYNNISGFNLFPWVSSMGFVELEFFSLKGNKLAGSIP-ELDF- 233
Query: 213 TNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNI 272
NL+Y+DLS N+ S P + + S L L LS+N K G I SL + L+ L N
Sbjct: 234 KNLSYLDLSANNFSTVFP-SFKDCSNLQHLDLSSN-KFYGDIGSSLSSCGKLSFLNLTNN 291
Query: 273 GLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDL-KNLIKLYLGSNNLSGPIPASIG 331
G +P +L+ L L N G P+ + DL K +++L L NN SG +P S+G
Sbjct: 292 QFVGLVPKLPSE--SLQYLYLRGNDFQGVYPNQLADLCKTVVELDLSYNNFSGMVPESLG 349
Query: 332 NLINLQVLSVQENNLTGTIPA-SIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVS 390
+L+++ + NN +G +P ++ L + ++ NK G +P+ N+ + +S
Sbjct: 350 ECSSLELVDISYNNFSGKLPVDTLSKLSNIKTMVLSFNKFVGGLPDSFSNLLKLETLDMS 409
Query: 391 ENDFVGHLPSQICSG--GSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQ 448
N+ G +PS IC +L++L +N F GPIP SL CS + + L N + G I
Sbjct: 410 SNNLTGVIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSNCSQLVSLDLSFNYLTGSIPS 469
Query: 449 DFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLH 508
G KL+ L L N+ G+I L+ I+ N+++G IP TKL +
Sbjct: 470 SLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPIPASLSNCTKLNWIS 529
Query: 509 LSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIP 568
LS+NQL+G++P LG + +L LK+ NN S NIP+E+G Q L LDL N L+G IP
Sbjct: 530 LSNNQLSGEIPAS-LGRLSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFLNGSIP 588
Query: 569 K----------------------------------ELVELPNLRMLNLSR---------- 584
L+E +R L R
Sbjct: 589 PPLFKQSGNIAVALLTGKRYVYIKNDGSKECHGAGNLLEFGGIRQEQLDRISTRHPCNFT 648
Query: 585 NKIEGIIPIKFD--SGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFG 642
GI F+ + LDLS N L+G+IP L + LS LNL HN LSG IPQ G
Sbjct: 649 RVYRGITQPTFNHNGSMIFLDLSYNKLEGSIPKELGAMYYLSILNLGHNDLSGMIPQQLG 708
Query: 643 --RNLVFVNISDNQLEGPLPKIPAFL-------------------SASFESLKN----NN 677
+N+ +++S N+ G +P L SA F++ + NN
Sbjct: 709 GLKNVAILDLSYNRFNGTIPNSLTSLTLLGEIDLSNNNLSGMIPESAPFDTFPDYRFANN 768
Query: 678 HLCGNIRGLDPC-------ATSHSRK-RKNVLRPVFIALGAVILVLCVVGALMY-IMCGR 728
LCG + PC A H + R+ +A+G + + C+ G ++ I +
Sbjct: 769 SLCGYPLPI-PCSSGPKSDANQHQKSHRRQASLAGSVAMGLLFSLFCIFGLIIVAIETKK 827
Query: 729 KKPNEESQTEEVQRGVLFSIWSHDG----------------------KMMFENIIEATAN 766
++ +E+ E G S ++ K+ F +++EAT
Sbjct: 828 RRRKKEAALEAYMDGHSHSATANSAWKFTSAREALSINLAAFEKPLRKLTFADLLEATNG 887
Query: 767 FDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHR 826
F + LVG G G+VYKA+L +G VVA+KKL V+ + + F +E+ET+ IKHR
Sbjct: 888 FHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQ-----GDREFTAEMETIGKIKHR 942
Query: 827 NIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQ-AVAFDWEKRVNVVKGVANALSYL 885
N++ L G+C + LVY++++ GSL+ +L++ + + +W R + G A L++L
Sbjct: 943 NLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKIAIGAARGLAFL 1002
Query: 886 HHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKP-GLH-SWTQFAGTFGYAAPEL 943
HH+C P IIHRD+ S NVLL+ + EA VSDFG A+ + H S + AGT GY PE
Sbjct: 1003 HHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGYVPPEY 1062
Query: 944 AQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNML----------LT 993
Q+ + K DVYS+GV+ LE + GK P D S NN++ +T
Sbjct: 1063 YQSFRCSTKGDVYSYGVVLLELLTGKQPTD--------SADFGDNNLVGWVKLHAKGKIT 1114
Query: 994 DVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKM 1038
DV D+ + I+ E++ ++A ACL RP+M QV M
Sbjct: 1115 DVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAM 1159
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 399 bits (1026), Expect = e-110
Identities = 285/952 (29%), Positives = 464/952 (47%), Gaps = 114/952 (11%)
Query: 184 EIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLV 243
++ + ++ L + + L G + + + L L ++L+ NSLSG I ++ NLS L+ L
Sbjct: 81 DVNESGRVVELELGRRKLSGKLSESVAKLDQLKVLNLTHNSLSGSIAASLLNLSNLEVLD 140
Query: 244 LSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSI-QNLVNLKELALDINHLSGSI 302
LS+N SG P SL N+ SL VL G IP S+ NL ++E+ L +N+ GSI
Sbjct: 141 LSSND-FSGLFP-SLINLPSLRVLNVYENSFHGLIPASLCNNLPRIREIDLAMNYFDGSI 198
Query: 303 PSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTV 362
P IG+ ++ L L SNNLSG IP + L NL VL++Q N L+G + + +G L L
Sbjct: 199 PVGIGNCSSVEYLGLASNNLSGSIPQELFQLSNLSVLALQNNRLSGALSSKLGKLSNLGR 258
Query: 363 FEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPI 422
++++NK G+IP+ + F N F G +P + + S+ LL+ +N +G I
Sbjct: 259 LDISSNKFSGKIPDVFLELNKLWYFSAQSNLFNGEMPRSLSNSRSISLLSLRNNTLSGQI 318
Query: 423 PTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQT 482
+ +++ + L N G I + +L+ ++ + KF QI ++ +L +
Sbjct: 319 YLNCSAMTNLTSLDLASNSFSGSIPSNLPNCLRLKTINFAKIKFIAQIPESFKNFQSLTS 378
Query: 483 FIISNNNISG-----------------VIPLDF----------IGLTKLGVLHLSSNQLT 515
SN++I V+ L+F + L VL ++S QL
Sbjct: 379 LSFSNSSIQNISSALEILQHCQNLKTLVLTLNFQKEELPSVPSLQFKNLKVLIIASCQLR 438
Query: 516 GKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELP 575
G +P + L SL L +S N S IP +G L L LDL N G+IP L L
Sbjct: 439 GTVP-QWLSNSPSLQLLDLSWNQLSGTIPPWLGSLNSLFYLDLSNNTFIGEIPHSLTSLQ 497
Query: 576 NL------------------------------------RMLNLSRNKIEGIIPIKFDS-- 597
+L M++LS N + G I +F
Sbjct: 498 SLVSKENAVEEPSPDFPFFKKKNTNAGGLQYNQPSSFPPMIDLSYNSLNGSIWPEFGDLR 557
Query: 598 GLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRN--LVFVNISDNQL 655
L L+L N L GNIP L+ + L L+LSHN LSG IP + + L +++ N+L
Sbjct: 558 QLHVLNLKNNNLSGNIPANLSGMTSLEVLDLSHNNLSGNIPPSLVKLSFLSTFSVAYNKL 617
Query: 656 EGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHS-------RKRKNVLRPVFIAL 708
GP+P F + S + N LCG PC + + +KN+ + V +A+
Sbjct: 618 SGPIPTGVQFQTFPNSSFEGNQGLCGE--HASPCHITDQSPHGSAVKSKKNIRKIVAVAV 675
Query: 709 GAVILVLCVVGALMYIMC-----GRKKPNEESQTEEVQRG----VLFSIWSHDGKMMFEN 759
G + + ++ + I+ G P +++ +E++ G VLF + ++ ++
Sbjct: 676 GTGLGTVFLLTVTLLIILRTTSRGEVDPEKKADADEIELGSRSVVLFHNKDSNNELSLDD 735
Query: 760 IIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIET 819
I+++T++F+ ++G G G VYKA L +G VA+K+L T + + F +E+ET
Sbjct: 736 ILKSTSSFNQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQ-----MDREFQAEVET 790
Query: 820 LTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAV-AFDWEKRVNVVKGV 878
L+ +H N++ L G+C++ L+Y +++ GSLD L+ + DW+ R+ + +G
Sbjct: 791 LSRAQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDWKTRLRIARGA 850
Query: 879 ANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKP-GLHSWTQFAGTFG 937
A L+YLH C P I+HRDI S N+LL+ + AH++DFG A+ + P H T GT G
Sbjct: 851 AEGLAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYDTHVTTDLVGTLG 910
Query: 938 YAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNMLLTDVL- 996
Y PE Q K DVYSFGV+ LE + G+ P D+ +P + L++ VL
Sbjct: 911 YIPPEYGQASVATYKGDVYSFGVVLLELLTGRRPMDV--------CKPRGSRDLISWVLQ 962
Query: 997 ---DQRPQQVMEPI------DEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
++R ++ +P EE++L+ +A CL +NP+ RP+ Q+ L
Sbjct: 963 MKTEKRESEIFDPFIYDKDHAEEMLLVLEIACRCLGENPKTRPTTQQLVSWL 1014
Score = 197 bits (500), Expect = 2e-49
Identities = 174/602 (28%), Positives = 274/602 (44%), Gaps = 94/602 (15%)
Query: 30 LKWKDSFDDQSQTLLSTWKNN------TNPCKPKWRGIKC-----------DKSNFISTI 72
LK + F ++ + WK N +N C W GI C ++S + +
Sbjct: 34 LKALEGFMRGLESSIDGWKWNESSSFSSNCCD--WVGISCKSSVSLGLDDVNESGRVVEL 91
Query: 73 GLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKN------- 125
L L G L S + + L ++++ +NS G+I A + NLSN+ +L +
Sbjct: 92 ELGRRKLSGKL-SESVAKLDQLKVLNLTHNSLSGSIAASLLNLSNLEVLDLSSNDFSGLF 150
Query: 126 ----------------NYFDGSIPQEMC-TLTGLQFLDISFCKLNGAIPKSIGNLTNLSY 168
N F G IP +C L ++ +D++ +G+IP IGN +++ Y
Sbjct: 151 PSLINLPSLRVLNVYENSFHGLIPASLCNNLPRIREIDLAMNYFDGSIPVGIGNCSSVEY 210
Query: 169 LILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGG 228
L L NN SG IP E+ +L+NL LA+Q + L G++ ++G L+NL +D+S N SG
Sbjct: 211 LGLASNNLSGS-IPQELFQLSNLSVLALQNNRLSGALSSKLGKLSNLGRLDISSNKFSGK 269
Query: 229 IPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNL 288
IP+ L+KL +N +G +P SL N S+++L N LSG I + + NL
Sbjct: 270 IPDVFLELNKLWYFSAQSNL-FNGEMPRSLSNSRSISLLSLRNNTLSGQIYLNCSAMTNL 328
Query: 289 KELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTG 348
L L N SGSIPS + + L + IP S N +L LS +++
Sbjct: 329 TSLDLASNSFSGSIPSNLPNCLRLKTINFAKIKFIAQIPESFKNFQSLTSLSFSNSSIQN 388
Query: 349 TIPA-------------------------SIGNLKW--LTVFEVATNKLHGRIPNGLYNI 381
A S+ +L++ L V +A+ +L G +P L N
Sbjct: 389 ISSALEILQHCQNLKTLVLTLNFQKEELPSVPSLQFKNLKVLIIASCQLRGTVPQWLSNS 448
Query: 382 TNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQ 441
+ +S N G +P + S SL L+ +N F G IP SL +S++ + + N
Sbjct: 449 PSLQLLDLSWNQLSGTIPPWLGSLNSLFYLDLSNNTFIGEIPHSL---TSLQSLVSKENA 505
Query: 442 IEGDIAQDFGVYPK-------LQY---------LDLSDNKFHGQISPNWGKSLNLQTFII 485
+E + + DF + K LQY +DLS N +G I P +G L +
Sbjct: 506 VE-EPSPDFPFFKKKNTNAGGLQYNQPSSFPPMIDLSYNSLNGSIWPEFGDLRQLHVLNL 564
Query: 486 SNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPS 545
NNN+SG IP + G+T L VL LS N L+G +P ++ + L ++ N S IP+
Sbjct: 565 KNNNLSGNIPANLSGMTSLEVLDLSHNNLSGNIPPSLV-KLSFLSTFSVAYNKLSGPIPT 623
Query: 546 EI 547
+
Sbjct: 624 GV 625
Score = 157 bits (396), Expect = 2e-37
Identities = 131/436 (30%), Positives = 183/436 (41%), Gaps = 80/436 (18%)
Query: 280 DSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVL 339
D + + EL L LSG + ++ L L L L N+LSG I AS+ NL NL+VL
Sbjct: 80 DDVNESGRVVELELGRRKLSGKLSESVAKLDQLKVLNLTHNSLSGSIAASLLNLSNLEVL 139
Query: 340 SVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLP 399
+ N+ +G P+ I NL L V V N HG IP L N +LP
Sbjct: 140 DLSSNDFSGLFPSLI-NLPSLRVLNVYENSFHGLIPASLCN----------------NLP 182
Query: 400 SQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYL 459
+R ++ N F G IP + CSS+E + L N + G I Q+ L L
Sbjct: 183 -------RIREIDLAMNYFDGSIPVGIGNCSSVEYLGLASNNLSGSIPQELFQLSNLSVL 235
Query: 460 DLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLP 519
L +N+ G +S GK NL IS+N SG IP F+ L KL SN G++P
Sbjct: 236 ALQNNRLSGALSSKLGKLSNLGRLDISSNKFSGKIPDVFLELNKLWYFSAQSNLFNGEMP 295
Query: 520 MEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRM 579
L +S+ L + NN S I + L LDL N SG IP L L+
Sbjct: 296 RS-LSNSRSISLLSLRNNTLSGQIYLNCSAMTNLTSLDLASNSFSGSIPSNLPNCLRLKT 354
Query: 580 LNLSRNKIEGIIPIKFDS----------------------------GLESLDLSGNF--- 608
+N ++ K IP F + L++L L+ NF
Sbjct: 355 INFAKIKFIAQIPESFKNFQSLTSLSFSNSSIQNISSALEILQHCQNLKTLVLTLNFQKE 414
Query: 609 ----------------------LKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFG--RN 644
L+G +P L++ L L+LS N LSGTIP G +
Sbjct: 415 ELPSVPSLQFKNLKVLIIASCQLRGTVPQWLSNSPSLQLLDLSWNQLSGTIPPWLGSLNS 474
Query: 645 LVFVNISDNQLEGPLP 660
L ++++S+N G +P
Sbjct: 475 LFYLDLSNNTFIGEIP 490
Score = 38.9 bits (89), Expect = 0.074
Identities = 30/111 (27%), Positives = 50/111 (45%), Gaps = 7/111 (6%)
Query: 589 GIIPIKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGR--NLV 646
G+ + + L+L L G + +A L +L LNL+HN LSG+I + NL
Sbjct: 78 GLDDVNESGRVVELELGRRKLSGKLSESVAKLDQLKVLNLTHNSLSGSIAASLLNLSNLE 137
Query: 647 FVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHSRKR 697
+++S N G P + + SL+ N + GL P + ++ R
Sbjct: 138 VLDLSSNDFSGLFPSL-----INLPSLRVLNVYENSFHGLIPASLCNNLPR 183
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 389 bits (998), Expect = e-107
Identities = 317/998 (31%), Positives = 483/998 (47%), Gaps = 108/998 (10%)
Query: 99 IRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPK 158
I N G + + N+ L +N F IP + + LQ LDIS KL+G +
Sbjct: 207 ISGNKISGDV--DVSRCVNLEFLDVSSNNFSTGIPF-LGDCSALQHLDISGNKLSGDFSR 263
Query: 159 SIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEI-GFLTNLAY 217
+I T L L + N + G PIPP L +L +L++ ++ G IP + G L
Sbjct: 264 AISTCTELKLLNISSNQFVG-PIPPL--PLKSLQYLSLAENKFTGEIPDFLSGACDTLTG 320
Query: 218 IDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIP-HSLWNMSSLTVLYFDNIGLSG 276
+DLS N G +P G+ S L++L LS+N SG +P +L M L VL SG
Sbjct: 321 LDLSGNHFYGAVPPFFGSCSLLESLALSSNN-FSGELPMDTLLKMRGLKVLDLSFNEFSG 379
Query: 277 SIPDSIQNL-VNLKELALDINHLSGSI-PSTIGDLKNLIK-LYLGSNNLSGPIPASIGNL 333
+P+S+ NL +L L L N+ SG I P+ + KN ++ LYL +N +G IP ++ N
Sbjct: 380 ELPESLTNLSASLLTLDLSSNNFSGPILPNLCQNPKNTLQELYLQNNGFTGKIPPTLSNC 439
Query: 334 INLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSEND 393
L L + N L+GTIP+S+G+L L ++ N L G IP L + + ++ ND
Sbjct: 440 SELVSLHLSFNYLSGTIPSSLGSLSKLRDLKLWLNMLEGEIPQELMYVKTLETLILDFND 499
Query: 394 FVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVY 453
G +PS + + +L ++ +NR TG IP + ++ + L N G+I + G
Sbjct: 500 LTGEIPSGLSNCTNLNWISLSNNRLTGEIPKWIGRLENLAILKLSNNSFSGNIPAELGDC 559
Query: 454 PKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISG--VIPLDFIGLTKLGVLHLSS 511
L +LDL+ N F+G I K Q+ I+ N I+G + + G+ K H +
Sbjct: 560 RSLIWLDLNTNLFNGTIPAAMFK----QSGKIAANFIAGKRYVYIKNDGMKK--ECHGAG 613
Query: 512 NQLTGK-LPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKE 570
N L + + E L + + I++ + + + LD+ N LSG IPKE
Sbjct: 614 NLLEFQGIRSEQLNRLSTRNPCNITSRVYGGHTSPTFDNNGSMMFLDMSYNMLSGYIPKE 673
Query: 571 LVELPNLRMLNLSRNKIEGIIPIKFDS--GLESLDLSGNFLKGNIPTGLADLVRLSKLNL 628
+ +P L +LNL N I G IP + GL LDLS N L G IP ++ L L++++L
Sbjct: 674 IGSMPYLFILNLGHNDISGSIPDEVGDLRGLNILDLSSNKLDGRIPQAMSALTMLTEIDL 733
Query: 629 SHNMLSGTIPQNFGRNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCG------- 681
S+N LSG IP+ + + F A F NN LCG
Sbjct: 734 SNNNLSGPIPE-------------------MGQFETFPPAKF---LNNPGLCGYPLPRCD 771
Query: 682 --NIRGLDPCATSHSRKRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEESQTEE 739
N G SH R+ ++ V A+G + +C+ G ++ RK+ ++ E
Sbjct: 772 PSNADGYAHHQRSHGRRPASLAGSV--AMGLLFSFVCIFGLILVGREMRKRRRKKEAELE 829
Query: 740 V---------QRGVLFSIWSHDG-----------------KMMFENIIEATANFDDKYLV 773
+ R + W G K+ F ++++AT F + L+
Sbjct: 830 MYAEGHGNSGDRTANNTNWKLTGVKEALSINLAAFEKPLRKLTFADLLQATNGFHNDSLI 889
Query: 774 GVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHG 833
G G G+VYKA L +G VA+KKL V+ + + FM+E+ET+ IKHRN++ L G
Sbjct: 890 GSGGFGDVYKAILKDGSAVAIKKLIHVSGQ-----GDREFMAEMETIGKIKHRNLVPLLG 944
Query: 834 FCSHSKFSFLVYKFLEGGSLDQILNNDTQA-VAFDWEKRVNVVKGVANALSYLHHDCSPP 892
+C LVY+F++ GSL+ +L++ +A V +W R + G A L++LHH+CSP
Sbjct: 945 YCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNWSTRRKIAIGSARGLAFLHHNCSPH 1004
Query: 893 IIHRDISSKNVLLNLDYEAHVSDFGTAKFLKP-GLH-SWTQFAGTFGYAAPELAQTMEVN 950
IIHRD+ S NVLL+ + EA VSDFG A+ + H S + AGT GY PE Q+ +
Sbjct: 1005 IIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGYVPPEYYQSFRCS 1064
Query: 951 EKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNML----------LTDVLDQRP 1000
K DVYS+GV+ LE + GK P D SP NN++ ++DV D
Sbjct: 1065 TKGDVYSYGVVLLELLTGKRPTD------SPDFGD--NNLVGWVKQHAKLRISDVFDPEL 1116
Query: 1001 QQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKM 1038
+ ++ E++ ++A ACL RP+M QV M
Sbjct: 1117 MKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVMAM 1154
Score = 214 bits (546), Expect = 8e-55
Identities = 165/512 (32%), Positives = 244/512 (47%), Gaps = 34/512 (6%)
Query: 97 IDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAI 156
+DI N G I + + +L +N F G IP L LQ+L ++ K G I
Sbjct: 250 LDISGNKLSGDFSRAISTCTELKLLNISSNQFVGPIPP--LPLKSLQYLSLAENKFTGEI 307
Query: 157 PKSI-GNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGF-LTN 214
P + G L+ L L GN++ G +PP G + L LA+ +N G +P + +
Sbjct: 308 PDFLSGACDTLTGLDLSGNHFYGA-VPPFFGSCSLLESLALSSNNFSGELPMDTLLKMRG 366
Query: 215 LAYIDLSKNSLSGGIPETIGNLS-KLDTLVLSNNTKMSGPIPHSLWN--MSSLTVLYFDN 271
L +DLS N SG +PE++ NLS L TL LS+N SGPI +L ++L LY N
Sbjct: 367 LKVLDLSFNEFSGELPESLTNLSASLLTLDLSSNN-FSGPILPNLCQNPKNTLQELYLQN 425
Query: 272 IGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIG 331
G +G IP ++ N L L L N+LSG+IPS++G L L L L N L G IP +
Sbjct: 426 NGFTGKIPPTLSNCSELVSLHLSFNYLSGTIPSSLGSLSKLRDLKLWLNMLEGEIPQELM 485
Query: 332 NLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSE 391
+ L+ L + N+LTG IP+ + N L ++ N+L G IP + + N +S
Sbjct: 486 YVKTLETLILDFNDLTGEIPSGLSNCTNLNWISLSNNRLTGEIPKWIGRLENLAILKLSN 545
Query: 392 NDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFG 451
N F G++P+++ SL L+ + N F G IP ++ S G IA +F
Sbjct: 546 NSFSGNIPAELGDCRSLIWLDLNTNLFNGTIPAAMFKQS-------------GKIAANFI 592
Query: 452 VYPKLQYLDLSDNK-----------FHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIG 500
+ Y+ K F G S + I++ G F
Sbjct: 593 AGKRYVYIKNDGMKKECHGAGNLLEFQGIRSEQLNRLSTRNPCNITSRVYGGHTSPTFDN 652
Query: 501 LTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGG 560
+ L +S N L+G +P E+ G M LF L + +N S +IP E+G L+ L LDL
Sbjct: 653 NGSMMFLDMSYNMLSGYIPKEI-GSMPYLFILNLGHNDISGSIPDEVGDLRGLNILDLSS 711
Query: 561 NELSGKIPKELVELPNLRMLNLSRNKIEGIIP 592
N+L G+IP+ + L L ++LS N + G IP
Sbjct: 712 NKLDGRIPQAMSALTMLTEIDLSNNNLSGPIP 743
Score = 196 bits (499), Expect = 2e-49
Identities = 142/450 (31%), Positives = 210/450 (46%), Gaps = 28/450 (6%)
Query: 72 IGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGS 131
+ LA G + + L +D+ N FYG +P G+ S + L +N F G
Sbjct: 296 LSLAENKFTGEIPDFLSGACDTLTGLDLSGNHFYGAVPPFFGSCSLLESLALSSNNFSGE 355
Query: 132 IPQE-MCTLTGLQFLDISFCKLNGAIPKSIGNLT-NLSYLILGGNNWSGGPIPPEI--GK 187
+P + + + GL+ LD+SF + +G +P+S+ NL+ +L L L NN+S GPI P +
Sbjct: 356 LPMDTLLKMRGLKVLDLSFNEFSGELPESLTNLSASLLTLDLSSNNFS-GPILPNLCQNP 414
Query: 188 LNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNN 247
N L L +Q + G IP + + L + LS N LSG IP ++G+LSKL L L N
Sbjct: 415 KNTLQELYLQNNGFTGKIPPTLSNCSELVSLHLSFNYLSGTIPSSLGSLSKLRDLKLWLN 474
Query: 248 TKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIG 307
+ G IP L + +L L D L+G IP + N NL ++L N L+G IP IG
Sbjct: 475 -MLEGEIPQELMYVKTLETLILDFNDLTGEIPSGLSNCTNLNWISLSNNRLTGEIPKWIG 533
Query: 308 DLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVAT 367
L+NL L L +N+ SG IPA +G+ +L L + N GTIPA++
Sbjct: 534 RLENLAILKLSNNSFSGNIPAELGDCRSLIWLDLNTNLFNGTIPAAMFKQSGKIAANFIA 593
Query: 368 NKLHGRIPNG-----LYNITNWISFV-----------------VSENDFVGHLPSQICSG 405
K + I N + N + F ++ + GH +
Sbjct: 594 GKRYVYIKNDGMKKECHGAGNLLEFQGIRSEQLNRLSTRNPCNITSRVYGGHTSPTFDNN 653
Query: 406 GSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNK 465
GS+ L+ +N +G IP + + + + L N I G I + G L LDLS NK
Sbjct: 654 GSMMFLDMSYNMLSGYIPKEIGSMPYLFILNLGHNDISGSIPDEVGDLRGLNILDLSSNK 713
Query: 466 FHGQISPNWGKSLNLQTFIISNNNISGVIP 495
G+I L +SNNN+SG IP
Sbjct: 714 LDGRIPQAMSALTMLTEIDLSNNNLSGPIP 743
Score = 145 bits (367), Expect = 4e-34
Identities = 107/363 (29%), Positives = 168/363 (45%), Gaps = 23/363 (6%)
Query: 34 DSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPN 93
+S + S +LL+ ++ N P + + N + + L N G G + T S+
Sbjct: 383 ESLTNLSASLLTLDLSSNNFSGPILPNLCQNPKNTLQELYLQNNGFTGKIPP-TLSNCSE 441
Query: 94 LLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLN 153
L+ + + N GTIP+ +G+LS + L N +G IPQE+ + L+ L + F L
Sbjct: 442 LVSLHLSFNYLSGTIPSSLGSLSKLRDLKLWLNMLEGEIPQELMYVKTLETLILDFNDLT 501
Query: 154 GAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLT 213
G IP + N TNL+++ L NN G IP IG+L NL L + ++ G+IP E+G
Sbjct: 502 GEIPSGLSNCTNLNWISL-SNNRLTGEIPKWIGRLENLAILKLSNNSFSGNIPAELGDCR 560
Query: 214 NLAYIDLSKNSLSGGIPETIGNLS---------------------KLDTLVLSNNTKMSG 252
+L ++DL+ N +G IP + S K + N + G
Sbjct: 561 SLIWLDLNTNLFNGTIPAAMFKQSGKIAANFIAGKRYVYIKNDGMKKECHGAGNLLEFQG 620
Query: 253 PIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNL 312
L +S+ + G + N ++ L + N LSG IP IG + L
Sbjct: 621 IRSEQLNRLSTRNPCNITSRVYGGHTSPTFDNNGSMMFLDMSYNMLSGYIPKEIGSMPYL 680
Query: 313 IKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHG 372
L LG N++SG IP +G+L L +L + N L G IP ++ L LT +++ N L G
Sbjct: 681 FILNLGHNDISGSIPDEVGDLRGLNILDLSSNKLDGRIPQAMSALTMLTEIDLSNNNLSG 740
Query: 373 RIP 375
IP
Sbjct: 741 PIP 743
Score = 94.7 bits (234), Expect = 1e-18
Identities = 100/368 (27%), Positives = 164/368 (44%), Gaps = 70/368 (19%)
Query: 374 IPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPT--SLKTCSS 431
+ + L ++T S +S + G + CS SL L+ N +GP+ T SL +CS
Sbjct: 91 VSSSLLSLTGLESLFLSNSHINGSVSGFKCSA-SLTSLDLSRNSLSGPVTTLTSLGSCSG 149
Query: 432 IERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLN-LQTFIISNNNI 490
++ + + N ++ F G++S G LN L+ +S N+I
Sbjct: 150 LKFLNVSSNTLD----------------------FPGKVSG--GLKLNSLEVLDLSANSI 185
Query: 491 SGVIPLDFI---GLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEI 547
SG + ++ G +L L +S N+++G + + ++ L +S+N+FS IP +
Sbjct: 186 SGANVVGWVLSDGCGELKHLAISGNKISGDVDVSRCVNLEFL---DVSSNNFSTGIPF-L 241
Query: 548 GLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIP--------------- 592
G LQ LD+ GN+LSG + + L++LN+S N+ G IP
Sbjct: 242 GDCSALQHLDISGNKLSGDFSRAISTCTELKLLNISSNQFVGPIPPLPLKSLQYLSLAEN 301
Query: 593 ------IKFDSG----LESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNF- 641
F SG L LDLSGN G +P L L LS N SG +P +
Sbjct: 302 KFTGEIPDFLSGACDTLTGLDLSGNHFYGAVPPFFGSCSLLESLALSSNNFSGELPMDTL 361
Query: 642 --GRNLVFVNISDNQLEGPLPKIPAFLSASFESLK-NNNHLCGNIRGLDPCATSHSRKRK 698
R L +++S N+ G LP+ LSAS +L ++N+ G P + + K
Sbjct: 362 LKMRGLKVLDLSFNEFSGELPESLTNLSASLLTLDLSSNNFSG------PILPNLCQNPK 415
Query: 699 NVLRPVFI 706
N L+ +++
Sbjct: 416 NTLQELYL 423
Score = 76.6 bits (187), Expect = 3e-13
Identities = 49/133 (36%), Positives = 79/133 (58%), Gaps = 4/133 (3%)
Query: 57 KWRGIKCDKSNFISTIGLANLGLK--GTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGN 114
+++GI+ ++ N +ST N+ + G S TF + +++ +D+ N G IP +IG+
Sbjct: 617 EFQGIRSEQLNRLSTRNPCNITSRVYGGHTSPTFDNNGSMMFLDMSYNMLSGYIPKEIGS 676
Query: 115 LSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGN 174
+ + IL +N GSIP E+ L GL LD+S KL+G IP+++ LT L+ + L N
Sbjct: 677 MPYLFILNLGHNDISGSIPDEVGDLRGLNILDLSSNKLDGRIPQAMSALTMLTEIDLSNN 736
Query: 175 NWSGGPIPPEIGK 187
N S GPI PE+G+
Sbjct: 737 NLS-GPI-PEMGQ 747
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 385 bits (989), Expect = e-106
Identities = 286/968 (29%), Positives = 472/968 (48%), Gaps = 129/968 (13%)
Query: 175 NWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIG 234
NW+G I ++ L + L G + + +G L + ++LS+N + IP +I
Sbjct: 64 NWTG--ITCNSNNTGRVIRLELGNKKLSGKLSESLGKLDEIRVLNLSRNFIKDSIPLSIF 121
Query: 235 NLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSI-QNLVNLKELAL 293
NL L TL LS+N +SG IP S+ N+ +L + +GS+P I N ++ + L
Sbjct: 122 NLKNLQTLDLSSND-LSGGIPTSI-NLPALQSFDLSSNKFNGSLPSHICHNSTQIRVVKL 179
Query: 294 DINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPAS 353
+N+ +G+ S G L L LG N+L+G IP + +L L +L +QEN L+G++
Sbjct: 180 AVNYFAGNFTSGFGKCVLLEHLCLGMNDLTGNIPEDLFHLKRLNLLGIQENRLSGSLSRE 239
Query: 354 IGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNA 413
I NL L +V+ N G IP+ + F+ N F+G +P + + SL LLN
Sbjct: 240 IRNLSSLVRLDVSWNLFSGEIPDVFDELPQLKFFLGQTNGFIGGIPKSLANSPSLNLLNL 299
Query: 414 DHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPN 473
+N +G + + ++ + L N+ G + ++ +L+ ++L+ N FHGQ+ +
Sbjct: 300 RNNSLSGRLMLNCTAMIALNSLDLGTNRFNGRLPENLPDCKRLKNVNLARNTFHGQVPES 359
Query: 474 WGKSLNLQTFIISNNNISG-----------------VIPLDFIG----------LTKLGV 506
+ +L F +SN++++ V+ L+F G KL V
Sbjct: 360 FKNFESLSYFSLSNSSLANISSALGILQHCKNLTTLVLTLNFHGEALPDDSSLHFEKLKV 419
Query: 507 LHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGK 566
L +++ +LTG +P L L L +S N + IPS IG + L LDL N +G+
Sbjct: 420 LVVANCRLTGSMP-RWLSSSNELQLLDLSWNRLTGAIPSWIGDFKALFYLDLSNNSFTGE 478
Query: 567 IPKELVELPNLRMLNLS------------------------------------RNKIEGI 590
IPK L +L +L N+S N + G
Sbjct: 479 IPKSLTKLESLTSRNISVNEPSPDFPFFMKRNESARALQYNQIFGFPPTIELGHNNLSGP 538
Query: 591 IPIKFDS--GLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRNLVFV 648
I +F + L DL N L G+IP+ L+ + L L+LS+N LSG+IP + + L F+
Sbjct: 539 IWEEFGNLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSGSIPVSL-QQLSFL 597
Query: 649 ---NISDNQLEGPLP---KIPAFLSASFESLKNNNHLCGNIR-----GLDPCATSHSRKR 697
+++ N L G +P + F ++SFES NHLCG R G + SR+
Sbjct: 598 SKFSVAYNNLSGVIPSGGQFQTFPNSSFES----NHLCGEHRFPCSEGTESALIKRSRRS 653
Query: 698 K--NVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEESQTE-----------EVQRGV 744
+ ++ + IA G+V L+ + +L+ + R+ + + E E+ +
Sbjct: 654 RGGDIGMAIGIAFGSVFLLTLL--SLIVLRARRRSGEVDPEIEESESMNRKELGEIGSKL 711
Query: 745 LFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEE 804
+ S+D ++ +++++++T +FD ++G G G VYKA L +G VA+KKL +
Sbjct: 712 VVLFQSNDKELSYDDLLDSTNSFDQANIIGCGGFGMVYKATLPDGKKVAIKKLSGDCGQ- 770
Query: 805 MSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILN--NDTQ 862
+ F +E+ETL+ +H N++ L GFC + L+Y ++E GSLD L+ ND
Sbjct: 771 ----IEREFEAEVETLSRAQHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDGP 826
Query: 863 AVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFL 922
A+ W+ R+ + +G A L YLH C P I+HRDI S N+LL+ ++ +H++DFG A+ +
Sbjct: 827 AL-LKWKTRLRIAQGAAKGLLYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARLM 885
Query: 923 KP-GLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSP 981
P H T GT GY PE Q K DVYSFGV+ LE + K P D+
Sbjct: 886 SPYETHVSTDLVGTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTDKRPVDM------- 938
Query: 982 STRPMANNMLLTDVL----DQRPQQVMEPI------DEEVILIARLAFACLSQNPRLRPS 1031
+P L++ V+ + R +V +P+ D+E+ + +A CLS+NP+ RP+
Sbjct: 939 -CKPKGCRDLISWVVKMKHESRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPT 997
Query: 1032 MGQVCKML 1039
Q+ L
Sbjct: 998 TQQLVSWL 1005
Score = 64.3 bits (155), Expect = 2e-09
Identities = 51/183 (27%), Positives = 83/183 (44%), Gaps = 8/183 (4%)
Query: 97 IDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAI 156
I++ +N+ G I + GNL + + K N GSIP + +T L+ LD+S +L+G+I
Sbjct: 528 IELGHNNLSGPIWEEFGNLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSGSI 587
Query: 157 PKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLA 216
P S+ L+ LS + NN SG + P G+ + + + ++L G T A
Sbjct: 588 PVSLQQLSFLSKFSVAYNNLSG--VIPSGGQFQTFPNSSFESNHLCGEHRFPCSEGTESA 645
Query: 217 YIDLSKNSLSGGIPETIG------NLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFD 270
I S+ S G I IG L L +L++ + SG + + S+
Sbjct: 646 LIKRSRRSRGGDIGMAIGIAFGSVFLLTLLSLIVLRARRRSGEVDPEIEESESMNRKELG 705
Query: 271 NIG 273
IG
Sbjct: 706 EIG 708
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated receptor
kinase 1 precursor (EC 2.7.1.37) (BRI1-associated
receptor kinase 1) (Somatic embryogenesis receptor-like
kinase 3)
Length = 615
Score = 222 bits (566), Expect = 4e-57
Identities = 166/509 (32%), Positives = 245/509 (47%), Gaps = 43/509 (8%)
Query: 556 LDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDSGLE--SLDLSGNFLKGNI 613
+DLG LSG++ +L +LPNL+ L L N I G IP + + E SLDL N L G I
Sbjct: 73 VDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGPI 132
Query: 614 PTGLADLVRLSKLNLSHNMLSGTIPQNFGRNLVF--VNISDNQLEGPLP---KIPAFLSA 668
P+ L L +L L L++N LSG IP++ L +++S+N L G +P F
Sbjct: 133 PSTLGRLKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIPVNGSFSLFTPI 192
Query: 669 SFESLKNNNHLCGNIRGLDPCATSHSRKRKNVLRPVFIALGAVILVLCVVGALMYIMCGR 728
SF + K + P S + + + + + A +L V A+ R
Sbjct: 193 SFANTKLTPLPASPPPPISPTPPSPAGSNR-ITGAIAGGVAAGAALLFAVPAIALAWWRR 251
Query: 729 KKPNEE------SQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVY 782
KKP + + EV G L + + A+ NF +K ++G G G VY
Sbjct: 252 KKPQDHFFDVPAEEDPEVHLGQL-------KRFSLRELQVASDNFSNKNILGRGGFGKVY 304
Query: 783 KAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSF 842
K L++G +VAVK+L EE + F +E+E ++ HRN+++L GFC
Sbjct: 305 KGRLADGTLVAVKRLK----EERTQGGELQFQTEVEMISMAVHRNLLRLRGFCMTPTERL 360
Query: 843 LVYKFLEGGSLDQILNNDTQAVA-FDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSK 901
LVY ++ GS+ L ++ DW KR + G A L+YLH C P IIHRD+ +
Sbjct: 361 LVYPYMANGSVASCLRERPESQPPLDWPKRQRIALGSARGLAYLHDHCDPKIIHRDVKAA 420
Query: 902 NVLLNLDYEAHVSDFGTAKFLK-PGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGV 960
N+LL+ ++EA V DFG AK + H T GT G+ APE T + +EK DV+ +GV
Sbjct: 421 NILLDEEFEAVVGDFGLAKLMDYKDTHVTTAVRGTIGHIAPEYLSTGKSSEKTDVFGYGV 480
Query: 961 LALETIMGKHPGDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPI----------DEE 1010
+ LE I G+ DL L MLL V ++ +E + DEE
Sbjct: 481 MLLELITGQRAFDLARLANDDDV------MLLDWVKGLLKEKKLEALVDVDLQGNYKDEE 534
Query: 1011 VILIARLAFACLSQNPRLRPSMGQVCKML 1039
V + ++A C +P RP M +V +ML
Sbjct: 535 VEQLIQVALLCTQSSPMERPKMSEVVRML 563
Score = 87.0 bits (214), Expect = 2e-16
Identities = 50/127 (39%), Positives = 78/127 (61%), Gaps = 1/127 (0%)
Query: 262 SSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNN 321
+S+T + N LSG + + L NL+ L L N+++G+IP +G+L L+ L L NN
Sbjct: 68 NSVTRVDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNN 127
Query: 322 LSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIP-NGLYN 380
LSGPIP+++G L L+ L + N+L+G IP S+ + L V +++ N L G IP NG ++
Sbjct: 128 LSGPIPSTLGRLKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIPVNGSFS 187
Query: 381 ITNWISF 387
+ ISF
Sbjct: 188 LFTPISF 194
Score = 85.5 bits (210), Expect = 7e-16
Identities = 52/130 (40%), Positives = 73/130 (56%), Gaps = 1/130 (0%)
Query: 241 TLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSG 300
T V N +SG + L + +L L + ++G+IP+ + NL L L L +N+LSG
Sbjct: 71 TRVDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSG 130
Query: 301 SIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWL 360
IPST+G LK L L L +N+LSG IP S+ ++ LQVL + N LTG IP + G+
Sbjct: 131 PIPSTLGRLKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIPVN-GSFSLF 189
Query: 361 TVFEVATNKL 370
T A KL
Sbjct: 190 TPISFANTKL 199
Score = 84.0 bits (206), Expect = 2e-15
Identities = 57/167 (34%), Positives = 90/167 (53%), Gaps = 4/167 (2%)
Query: 189 NNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNT 248
N++ + + +NL G + ++G L NL Y++L N+++G IPE +GNL++L +L L N
Sbjct: 68 NSVTRVDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLN- 126
Query: 249 KMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGD 308
+SGPIP +L + L L +N LSG IP S+ ++ L+ L L N L+G IP G
Sbjct: 127 NLSGPIPSTLGRLKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIPVN-GS 185
Query: 309 LKNLIKLYLGSNNLSGPIPASIGNLINLQVLS-VQENNLTGTIPASI 354
+ + L+ P+PAS I+ S N +TG I +
Sbjct: 186 FSLFTPISFANTKLT-PLPASPPPPISPTPPSPAGSNRITGAIAGGV 231
Score = 83.6 bits (205), Expect = 3e-15
Identities = 47/107 (43%), Positives = 68/107 (62%), Gaps = 1/107 (0%)
Query: 173 GNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPET 232
GN G + ++G+L NL +L + +N+ G+IP+++G LT L +DL N+LSG IP T
Sbjct: 76 GNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGPIPST 135
Query: 233 IGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIP 279
+G L KL L L+NN+ +SG IP SL + +L VL N L+G IP
Sbjct: 136 LGRLKKLRFLRLNNNS-LSGEIPRSLTAVLTLQVLDLSNNPLTGDIP 181
Score = 76.3 bits (186), Expect = 4e-13
Identities = 69/257 (26%), Positives = 110/257 (41%), Gaps = 57/257 (22%)
Query: 2 MVLPTLIMILCVLP-TLSVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRG 60
+++P ++ VL L V+ ++E AL K+S D ++ L S PC W
Sbjct: 5 LMIPCFFWLILVLDLVLRVSGNAEGD-ALSALKNSLADPNKVLQSWDATLVTPCT--WFH 61
Query: 61 IKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISI 120
+ C+ N ++ + L N L G L Q+G L N+
Sbjct: 62 VTCNSDNSVTRVDLGNANLSGQL-------------------------VMQLGQLPNLQY 96
Query: 121 LTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGP 180
L +N G+IP+++ LT L LD+ L+G IP ++G L L +L
Sbjct: 97 LELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGPIPSTLGRLKKLRFL----------- 145
Query: 181 IPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLD 240
+LNN ++L G IP+ + + L +DLS N L+G IP + L
Sbjct: 146 ------RLNN--------NSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIP--VNGSFSLF 189
Query: 241 TLVLSNNTKMSGPIPHS 257
T + NTK++ P+P S
Sbjct: 190 TPISFANTKLT-PLPAS 205
Score = 73.6 bits (179), Expect = 3e-12
Identities = 39/115 (33%), Positives = 64/115 (54%)
Query: 311 NLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKL 370
++ ++ LG+ NLSG + +G L NLQ L + NN+TGTIP +GNL L ++ N L
Sbjct: 69 SVTRVDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNL 128
Query: 371 HGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTS 425
G IP+ L + ++ N G +P + + +L++L+ +N TG IP +
Sbjct: 129 SGPIPSTLGRLKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIPVN 183
Score = 72.4 bits (176), Expect = 6e-12
Identities = 40/103 (38%), Positives = 55/103 (52%)
Query: 297 HLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGN 356
+LSG + +G L NL L L SNN++G IP +GNL L L + NNL+G IP+++G
Sbjct: 79 NLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGPIPSTLGR 138
Query: 357 LKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLP 399
LK L + N L G IP L + +S N G +P
Sbjct: 139 LKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIP 181
Score = 59.3 bits (142), Expect = 5e-08
Identities = 59/212 (27%), Positives = 90/212 (41%), Gaps = 42/212 (19%)
Query: 396 GHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPK 455
G L Q+ +L+ L N TG IP L + + + L +N + G I G K
Sbjct: 82 GQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGPIPSTLGRLKK 141
Query: 456 LQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLD---------FIGLTKLGV 506
L++L L++N G+I + L LQ +SNN ++G IP++ TKL
Sbjct: 142 LRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIPVNGSFSLFTPISFANTKLTP 201
Query: 507 LHLS--------------SNQLTGKLPMEVLGGMKSLFDL----------KISNNHFSDN 542
L S SN++TG + V G LF + K +HF D
Sbjct: 202 LPASPPPPISPTPPSPAGSNRITGAIAGGVAAGAALLFAVPAIALAWWRRKKPQDHFFD- 260
Query: 543 IPSE------IGLLQR--LQELDLGGNELSGK 566
+P+E +G L+R L+EL + + S K
Sbjct: 261 VPAEEDPEVHLGQLKRFSLRELQVASDNFSNK 292
Score = 57.4 bits (137), Expect = 2e-07
Identities = 39/127 (30%), Positives = 61/127 (47%), Gaps = 3/127 (2%)
Query: 404 SGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSD 463
S S+ ++ + +G + L +++ + L N I G I + G +L LDL
Sbjct: 66 SDNSVTRVDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYL 125
Query: 464 NKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVL 523
N G I G+ L+ ++NN++SG IP + L VL LS+N LTG +P +
Sbjct: 126 NNLSGPIPSTLGRLKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIP---V 182
Query: 524 GGMKSLF 530
G SLF
Sbjct: 183 NGSFSLF 189
>TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor
Length = 674
Score = 207 bits (528), Expect = 9e-53
Identities = 183/610 (30%), Positives = 291/610 (47%), Gaps = 86/610 (14%)
Query: 471 SPNW-------GKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVL 523
SP W SL+L + + + N++G +P + + L + L+ N L+G +P+E L
Sbjct: 85 SPQWTNTSLFNDSSLHLLSLQLPSANLTGSLPREIGEFSMLQSVFLNINSLSGSIPLE-L 143
Query: 524 GGMKSLFDLKISNNHFSDNIPSEI-GLLQRLQELDLGGNELSGKIPKELV---ELPNLRM 579
G SL D+ +S N + +P I L +L + GN LSG +P+ + NL++
Sbjct: 144 GYTSSLSDVDLSGNALAGVLPPSIWNLCDKLVSFKIHGNNLSGVLPEPALPNSTCGNLQV 203
Query: 580 LNLSRNKIEGIIP--IKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTI 637
L+L NK G P I G++SLDLS N +G +P GL ++ L LNLSHN SG +
Sbjct: 204 LDLGGNKFSGEFPEFITRFKGVKSLDLSSNVFEGLVPEGLG-VLELESLNLSHNNFSGML 262
Query: 638 PQNFGRNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHSRKR 697
P +FG + F + SFE N+ LCG L PC S SR
Sbjct: 263 P-DFGES-------------------KFGAESFEG--NSPSLCG--LPLKPCLGS-SRLS 297
Query: 698 KNVLRPVFIAL--GAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFSIWSHDG-- 753
+ + I L GAV++ ++G Y+ ++K + ES+ + + I +G
Sbjct: 298 PGAVAGLVIGLMSGAVVVASLLIG---YLQNKKRKSSIESEDDLEEGDEEDEIGEKEGGE 354
Query: 754 ----------KMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDE 803
+ ++++ AT +K S G VYKA+LS+G +A++ L
Sbjct: 355 GKLVVFQGGENLTLDDVLNATGQVMEK-----TSYGTVYKAKLSDGGNIALRLL-----R 404
Query: 804 EMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSK-FSFLVYKFLEGGSLDQILN-NDT 861
E +C S + I L I+H N++ L F + L+Y +L SL +L+ +
Sbjct: 405 EGTCKDRSSCLPVIRQLGRIRHENLVPLRAFYQGKRGEKLLIYDYLPNISLHDLLHESKP 464
Query: 862 QAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKF 921
+ A +W +R + G+A L+YLH PIIH +I SKNVL++ + A +++FG K
Sbjct: 465 RKPALNWARRHKIALGIARGLAYLHTGQEVPIIHGNIRSKNVLVDDFFFARLTEFGLDKI 524
Query: 922 LKPGL-HSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPG-------- 972
+ + A + GY APEL + + N + DVY+FG+L LE +MGK PG
Sbjct: 525 MVQAVADEIVSQAKSDGYKAPELHKMKKCNPRSDVYAFGILLLEILMGKKPGKSGRNGNE 584
Query: 973 --DLISLFLSPSTRPMANNMLLTDVLD-QRPQQVMEPIDEEVILIARLAFACLSQNPRLR 1029
DL SL + +V D + + + P++E ++ +LA C + +R
Sbjct: 585 FVDLPSL-----VKAAVLEETTMEVFDLEAMKGIRSPMEEGLVHALKLAMGCCAPVTTVR 639
Query: 1030 PSMGQVCKML 1039
PSM +V K L
Sbjct: 640 PSMEEVVKQL 649
Score = 89.7 bits (221), Expect = 4e-17
Identities = 60/166 (36%), Positives = 91/166 (54%), Gaps = 6/166 (3%)
Query: 190 NLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTK 249
+LL L + +NL GS+P+EIG + L + L+ NSLSG IP +G S L + LS N
Sbjct: 100 HLLSLQLPSANLTGSLPREIGEFSMLQSVFLNINSLSGSIPLELGYTSSLSDVDLSGNA- 158
Query: 250 MSGPIPHSLWNMSSLTVLY-FDNIGLSGSIPD-SIQNLV--NLKELALDINHLSGSIPST 305
++G +P S+WN+ V + LSG +P+ ++ N NL+ L L N SG P
Sbjct: 159 LAGVLPPSIWNLCDKLVSFKIHGNNLSGVLPEPALPNSTCGNLQVLDLGGNKFSGEFPEF 218
Query: 306 IGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIP 351
I K + L L SN G +P +G ++ L+ L++ NN +G +P
Sbjct: 219 ITRFKGVKSLDLSSNVFEGLVPEGLG-VLELESLNLSHNNFSGMLP 263
Score = 88.6 bits (218), Expect = 8e-17
Identities = 79/243 (32%), Positives = 123/243 (50%), Gaps = 10/243 (4%)
Query: 18 SVAEDSEAKLALLKWKDSFDDQSQTLL-STWKNNTNPCKPKWRGIKCDKSNFISTIGLAN 76
S++ S+ KL L K K S S++LL S+W ++ C+ WRG+K SN S + ++
Sbjct: 26 SLSGSSDVKLLLGKIKSSLQGNSESLLLSSWNSSVPVCQ--WRGVKWVFSNG-SPLQCSD 82
Query: 77 LGL-KGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQE 135
L + T SL S +LL + + + + G++P +IG S + + N GSIP E
Sbjct: 83 LSSPQWTNTSLFNDSSLHLLSLQLPSANLTGSLPREIGEFSMLQSVFLNINSLSGSIPLE 142
Query: 136 MCTLTGLQFLDISFCKLNGAIPKSIGNLTN-LSYLILGGNNWSGGPIPPEI--GKLNNLL 192
+ + L +D+S L G +P SI NL + L + GNN SG P + NL
Sbjct: 143 LGYTSSLSDVDLSGNALAGVLPPSIWNLCDKLVSFKIHGNNLSGVLPEPALPNSTCGNLQ 202
Query: 193 HLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSG 252
L + + G P+ I + +DLS N G +PE +G L +L++L LS+N SG
Sbjct: 203 VLDLGGNKFSGEFPEFITRFKGVKSLDLSSNVFEGLVPEGLGVL-ELESLNLSHN-NFSG 260
Query: 253 PIP 255
+P
Sbjct: 261 MLP 263
Score = 85.1 bits (209), Expect = 9e-16
Identities = 80/282 (28%), Positives = 135/282 (47%), Gaps = 34/282 (12%)
Query: 286 VNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENN 345
++L L L +L+GS+P IG+ L ++L N+LSG IP +G +L + + N
Sbjct: 99 LHLLSLQLPSANLTGSLPREIGEFSMLQSVFLNINSLSGSIPLELGYTSSLSDVDLSGNA 158
Query: 346 LTGTIPASIGNL-KWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICS 404
L G +P SI NL L F++ N L G +P LP+ C
Sbjct: 159 LAGVLPPSIWNLCDKLVSFKIHGNNLSGVLPEPA-------------------LPNSTC- 198
Query: 405 GGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDN 464
G+L++L+ N+F+G P + ++ + L N EG + + GV +L+ L+LS N
Sbjct: 199 -GNLQVLDLGGNKFSGEFPEFITRFKGVKSLDLSSNVFEGLVPEGLGVL-ELESLNLSHN 256
Query: 465 KFHGQISPNWGKS-LNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVL 523
F G + P++G+S ++F ++ ++ G +PL LG LS + G L + ++
Sbjct: 257 NFSGML-PDFGESKFGAESFEGNSPSLCG-LPLK----PCLGSSRLSPGAVAG-LVIGLM 309
Query: 524 GG---MKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNE 562
G + SL + N +I SE L + +E ++G E
Sbjct: 310 SGAVVVASLLIGYLQNKKRKSSIESEDDLEEGDEEDEIGEKE 351
Score = 81.3 bits (199), Expect = 1e-14
Identities = 63/178 (35%), Positives = 91/178 (50%), Gaps = 17/178 (9%)
Query: 257 SLWNMSSLTVLYFD--NIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIK 314
SL+N SSL +L + L+GS+P I L+ + L+IN LSGSIP +G +L
Sbjct: 92 SLFNDSSLHLLSLQLPSANLTGSLPREIGEFSMLQSVFLNINSLSGSIPLELGYTSSLSD 151
Query: 315 LYLGSNNLSGPIPASIGNLIN-LQVLSVQENNLTGTIP------ASIGNLKWLTVFEVAT 367
+ L N L+G +P SI NL + L + NNL+G +P ++ GNL+ V ++
Sbjct: 152 VDLSGNALAGVLPPSIWNLCDKLVSFKIHGNNLSGVLPEPALPNSTCGNLQ---VLDLGG 208
Query: 368 NKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRL--LNADHNRFTGPIP 423
NK G P + S +S N F G +P + G L L LN HN F+G +P
Sbjct: 209 NKFSGEFPEFITRFKGVKSLDLSSNVFEGLVPEGL---GVLELESLNLSHNNFSGMLP 263
Score = 76.6 bits (187), Expect = 3e-13
Identities = 54/153 (35%), Positives = 82/153 (53%), Gaps = 6/153 (3%)
Query: 179 GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNL-S 237
G +P EIG+ + L + + ++L GSIP E+G+ ++L+ +DLS N+L+G +P +I NL
Sbjct: 113 GSLPREIGEFSMLQSVFLNINSLSGSIPLELGYTSSLSDVDLSGNALAGVLPPSIWNLCD 172
Query: 238 KLDTLVLSNNTKMSGPIPHSLWNMS---SLTVLYFDNIGLSGSIPDSIQNLVNLKELALD 294
KL + + N +SG +P S +L VL SG P+ I +K L L
Sbjct: 173 KLVSFKIHGN-NLSGVLPEPALPNSTCGNLQVLDLGGNKFSGEFPEFITRFKGVKSLDLS 231
Query: 295 INHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIP 327
N G +P +G L+ L L L NN SG +P
Sbjct: 232 SNVFEGLVPEGLGVLE-LESLNLSHNNFSGMLP 263
>SIRK_ARATH (O64483) Senescence-induced receptor-like serine/threonine
kinase precursor (FLG22-induced receptor-like kinase 1)
Length = 876
Score = 170 bits (430), Expect = 2e-41
Identities = 135/470 (28%), Positives = 219/470 (45%), Gaps = 54/470 (11%)
Query: 575 PNLRMLNLSRNKIEGIIPIKFDS--GLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNM 632
P + LN+S +++ G I F + + LDLSGN L G IP LA+L L++LN+ N
Sbjct: 414 PRVVSLNISFSELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLTELNVEGNK 473
Query: 633 LSGTIPQNFGRNLVFVNISDNQLEGPLPKIPAFLSASFESLK--NNNHLCGNIRGLDPCA 690
L+G +PQ + + G L SL+ N LC + D C+
Sbjct: 474 LTGIVPQR---------LHERSKNGSL------------SLRFGRNPDLCLS----DSCS 508
Query: 691 TSHSRKRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFSIWS 750
+ + + + P+ + +I+VL AL R K ++ T + G L +
Sbjct: 509 NTKKKNKNGYIIPLVVV--GIIVVLLTALALFR----RFKKKQQRGTLGERNGPLKTAKR 562
Query: 751 HDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSS 810
+ + ++ T NF+ ++G G G VY ++ G VAVK L E S
Sbjct: 563 Y---FKYSEVVNITNNFER--VIGKGGFGKVYHGVIN-GEQVAVKVL-----SEESAQGY 611
Query: 811 KSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEK 870
K F +E++ L + H N+ L G+C+ L+Y+++ +L L + WE+
Sbjct: 612 KEFRAEVDLLMRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGKRSFI-LSWEE 670
Query: 871 RVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAK--FLKPGLHS 928
R+ + A L YLH+ C PPI+HRD+ N+LLN +A ++DFG ++ ++
Sbjct: 671 RLKISLDAAQGLEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSGQI 730
Query: 929 WTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGK---HPGDLISLFLSPSTRP 985
T AG+ GY PE T ++NEK DVYS GV+ LE I G+ + +S R
Sbjct: 731 STVVAGSIGYLDPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSKTEKVHISDHVRS 790
Query: 986 MANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQV 1035
+ N + ++DQR ++ + ++ +A AC RP+M QV
Sbjct: 791 ILANGDIRGIVDQRLRERYDV--GSAWKMSEIALACTEHTSAQRPTMSQV 838
Score = 51.2 bits (121), Expect = 1e-05
Identities = 29/77 (37%), Positives = 41/77 (52%)
Query: 532 LKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGII 591
L IS + I L +++LDL GN L+G+IP L LPNL LN+ NK+ GI+
Sbjct: 419 LNISFSELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLTELNVEGNKLTGIV 478
Query: 592 PIKFDSGLESLDLSGNF 608
P + ++ LS F
Sbjct: 479 PQRLHERSKNGSLSLRF 495
Score = 49.7 bits (117), Expect = 4e-05
Identities = 31/76 (40%), Positives = 44/76 (57%), Gaps = 4/76 (5%)
Query: 280 DSIQ--NLVNLKELALDIN--HLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLIN 335
D IQ N N + ++L+I+ L G I +L ++ KL L N L+G IPA + NL N
Sbjct: 404 DCIQSDNTTNPRVVSLNISFSELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPN 463
Query: 336 LQVLSVQENNLTGTIP 351
L L+V+ N LTG +P
Sbjct: 464 LTELNVEGNKLTGIVP 479
Score = 47.0 bits (110), Expect = 3e-04
Identities = 25/68 (36%), Positives = 37/68 (53%)
Query: 312 LIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLH 371
++ L + + L G I + NL +++ L + N LTG IPA + NL LT V NKL
Sbjct: 416 VVSLNISFSELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLTELNVEGNKLT 475
Query: 372 GRIPNGLY 379
G +P L+
Sbjct: 476 GIVPQRLH 483
Score = 45.8 bits (107), Expect = 6e-04
Identities = 27/93 (29%), Positives = 47/93 (50%), Gaps = 9/93 (9%)
Query: 52 NPCKP---KWRGIKCDKSNFISTIGLANLG-----LKGTLHSLTFSSFPNLLMIDIRNNS 103
+PC P W GI C +S+ + + +L L+G + FS+ ++ +D+ N+
Sbjct: 391 DPCVPVDYSWEGIDCIQSDNTTNPRVVSLNISFSELRGQIDP-AFSNLTSIRKLDLSGNT 449
Query: 104 FYGTIPAQIGNLSNISILTFKNNYFDGSIPQEM 136
G IPA + NL N++ L + N G +PQ +
Sbjct: 450 LTGEIPAFLANLPNLTELNVEGNKLTGIVPQRL 482
Score = 42.4 bits (98), Expect = 0.007
Identities = 22/77 (28%), Positives = 40/77 (51%)
Query: 446 IAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLG 505
I D P++ L++S ++ GQI P + +++ +S N ++G IP L L
Sbjct: 406 IQSDNTTNPRVVSLNISFSELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLT 465
Query: 506 VLHLSSNQLTGKLPMEV 522
L++ N+LTG +P +
Sbjct: 466 ELNVEGNKLTGIVPQRL 482
Score = 41.6 bits (96), Expect = 0.011
Identities = 20/54 (37%), Positives = 33/54 (61%)
Query: 274 LSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIP 327
L G I + NL ++++L L N L+G IP+ + +L NL +L + N L+G +P
Sbjct: 426 LRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLTELNVEGNKLTGIVP 479
Score = 41.6 bits (96), Expect = 0.011
Identities = 32/101 (31%), Positives = 45/101 (43%), Gaps = 14/101 (13%)
Query: 175 NWSGGPIPP-----------EIGKLNN--LLHLAIQKSNLVGSIPQEIGFLTNLAYIDLS 221
NW G P P + N ++ L I S L G I LT++ +DLS
Sbjct: 387 NWQGDPCVPVDYSWEGIDCIQSDNTTNPRVVSLNISFSELRGQIDPAFSNLTSIRKLDLS 446
Query: 222 KNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMS 262
N+L+G IP + NL L L + N K++G +P L S
Sbjct: 447 GNTLTGEIPAFLANLPNLTELNVEGN-KLTGIVPQRLHERS 486
Score = 39.3 bits (90), Expect = 0.057
Identities = 29/100 (29%), Positives = 43/100 (43%), Gaps = 25/100 (25%)
Query: 145 LDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGS 204
L+ISF +L G I + NLT++ L L GN L G
Sbjct: 419 LNISFSELRGQIDPAFSNLTSIRKLDLSGNT-------------------------LTGE 453
Query: 205 IPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVL 244
IP + L NL +++ N L+G +P+ + SK +L L
Sbjct: 454 IPAFLANLPNLTELNVEGNKLTGIVPQRLHERSKNGSLSL 493
Score = 38.1 bits (87), Expect = 0.13
Identities = 21/67 (31%), Positives = 39/67 (57%), Gaps = 1/67 (1%)
Query: 507 LHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGK 566
L++S ++L G++ + S+ L +S N + IP+ + L L EL++ GN+L+G
Sbjct: 419 LNISFSELRGQID-PAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLTELNVEGNKLTGI 477
Query: 567 IPKELVE 573
+P+ L E
Sbjct: 478 VPQRLHE 484
Score = 36.2 bits (82), Expect = 0.48
Identities = 24/78 (30%), Positives = 39/78 (49%), Gaps = 2/78 (2%)
Query: 103 SFYGTIPAQIGNLSNISILTFKNNYFD--GSIPQEMCTLTGLQFLDISFCKLNGAIPKSI 160
S+ G Q N +N +++ ++ + G I LT ++ LD+S L G IP +
Sbjct: 399 SWEGIDCIQSDNTTNPRVVSLNISFSELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFL 458
Query: 161 GNLTNLSYLILGGNNWSG 178
NL NL+ L + GN +G
Sbjct: 459 ANLPNLTELNVEGNKLTG 476
Score = 36.2 bits (82), Expect = 0.48
Identities = 19/59 (32%), Positives = 33/59 (55%)
Query: 248 TKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTI 306
+++ G I + N++S+ L L+G IP + NL NL EL ++ N L+G +P +
Sbjct: 424 SELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLTELNVEGNKLTGIVPQRL 482
Score = 34.7 bits (78), Expect = 1.4
Identities = 25/100 (25%), Positives = 43/100 (43%), Gaps = 11/100 (11%)
Query: 365 VATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPT 424
++ ++L G+I N+T+ +S N G +P+ + + +L LN + N+ TG +P
Sbjct: 421 ISFSELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLTELNVEGNKLTGIVPQ 480
Query: 425 SLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDN 464
L S G ++ FG P L D N
Sbjct: 481 RLHERSK-----------NGSLSLRFGRNPDLCLSDSCSN 509
Score = 33.1 bits (74), Expect = 4.1
Identities = 19/75 (25%), Positives = 33/75 (43%)
Query: 435 ITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVI 494
+ + +++ G I F ++ LDLS N G+I NL + N ++G++
Sbjct: 419 LNISFSELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLTELNVEGNKLTGIV 478
Query: 495 PLDFIGLTKLGVLHL 509
P +K G L L
Sbjct: 479 PQRLHERSKNGSLSL 493
Score = 32.0 bits (71), Expect = 9.1
Identities = 28/122 (22%), Positives = 55/122 (44%), Gaps = 16/122 (13%)
Query: 385 ISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEG 444
+S +S ++ G + + S+R L+ N TG IP L ++ + +E N++ G
Sbjct: 417 VSLNISFSELRGQIDPAFSNLTSIRKLDLSGNTLTGEIPAFLANLPNLTELNVEGNKLTG 476
Query: 445 DIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNN----NISG-VIPLDFI 499
+ Q L + +G +S +G++ +L +N N +G +IPL +
Sbjct: 477 IVPQ-----------RLHERSKNGSLSLRFGRNPDLCLSDSCSNTKKKNKNGYIIPLVVV 525
Query: 500 GL 501
G+
Sbjct: 526 GI 527
>TMM_ARATH (Q9SSD1) TOO MANY MOUTHS protein precursor (TMM)
Length = 496
Score = 157 bits (397), Expect = 1e-37
Identities = 88/236 (37%), Positives = 138/236 (58%), Gaps = 4/236 (1%)
Query: 93 NLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKL 152
+L + +R N F G IP ++GNL+N+ +L N+ +GSIP +GL+ LD+S +L
Sbjct: 160 SLQTLVLRENGFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGLRSLDLSGNRL 219
Query: 153 NGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFL 212
G+IP + L LS L L N +G P+PP + +L+ + + ++ + G IP+ I L
Sbjct: 220 TGSIPGFV--LPALSVLDLNQNLLTG-PVPPTLTSCGSLIKIDLSRNRVTGPIPESINRL 276
Query: 213 TNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLW-NMSSLTVLYFDN 271
L +DLS N LSG P ++ L+ L L+L NTK S IP + + + +L +L N
Sbjct: 277 NQLVLLDLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKGLKNLMILVLSN 336
Query: 272 IGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIP 327
+ GSIP S+ L +L+ L L+ N+L+G IP D+K+L +L L N+L+GP+P
Sbjct: 337 TNIQGSIPKSLTRLNSLRVLHLEGNNLTGEIPLEFRDVKHLSELRLNDNSLTGPVP 392
Score = 126 bits (316), Expect = 4e-28
Identities = 92/294 (31%), Positives = 136/294 (45%), Gaps = 51/294 (17%)
Query: 262 SSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNN 321
SSL L G G IPD + NL NLK L L NHL+GSIP + L L L N
Sbjct: 159 SSLQTLVLRENGFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGLRSLDLSGNR 218
Query: 322 LSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNI 381
L+G IP + L L VL + +N LTG +P ++ + L +++ N++ G IP + +
Sbjct: 219 LTGSIPGFV--LPALSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRNRVTGPIPESINRL 276
Query: 382 TNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQ 441
L LL+ +NR +GP P+SL+ +S++ + L+ N
Sbjct: 277 ------------------------NQLVLLDLSYNRLSGPFPSSLQGLNSLQALMLKGN- 311
Query: 442 IEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSL-NLQTFIISNNNISGVIPLDFIG 500
KF I N K L NL ++SN NI G IP
Sbjct: 312 ----------------------TKFSTTIPENAFKGLKNLMILVLSNTNIQGSIPKSLTR 349
Query: 501 LTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQ 554
L L VLHL N LTG++P+E +K L +L++++N + +P E + R++
Sbjct: 350 LNSLRVLHLEGNNLTGEIPLE-FRDVKHLSELRLNDNSLTGPVPFERDTVWRMR 402
Score = 123 bits (309), Expect = 2e-27
Identities = 85/274 (31%), Positives = 132/274 (48%), Gaps = 28/274 (10%)
Query: 326 IPASIGNL-INLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNW 384
IPA +G L +LQ L ++EN G IP +GNL L V ++ N L+G IP +
Sbjct: 150 IPAFLGRLGSSLQTLVLRENGFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGL 209
Query: 385 ISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEG 444
S +S N G +P + +L +L+ + N TGP+P +L +C S+ +I L N++ G
Sbjct: 210 RSLDLSGNRLTGSIPGFVLP--ALSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRNRVTG 267
Query: 445 DIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKL 504
I + +L LDLS N+ +SG P GL L
Sbjct: 268 PIPESINRLNQLVLLDLSYNR------------------------LSGPFPSSLQGLNSL 303
Query: 505 GVLHLSSN-QLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNEL 563
L L N + + +P G+K+L L +SN + +IP + L L+ L L GN L
Sbjct: 304 QALMLKGNTKFSTTIPENAFKGLKNLMILVLSNTNIQGSIPKSLTRLNSLRVLHLEGNNL 363
Query: 564 SGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDS 597
+G+IP E ++ +L L L+ N + G +P + D+
Sbjct: 364 TGEIPLEFRDVKHLSELRLNDNSLTGPVPFERDT 397
Score = 122 bits (307), Expect = 4e-27
Identities = 83/257 (32%), Positives = 136/257 (52%), Gaps = 6/257 (2%)
Query: 181 IPPEIGKLNNLLH-LAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKL 239
IP +G+L + L L ++++ +G IP E+G LTNL +DL KN L+G IP + S L
Sbjct: 150 IPAFLGRLGSSLQTLVLRENGFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGL 209
Query: 240 DTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLS 299
+L LS N +++G IP + + +L+VL + L+G +P ++ + +L ++ L N ++
Sbjct: 210 RSLDLSGN-RLTGSIPGFV--LPALSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRNRVT 266
Query: 300 GSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQEN-NLTGTIPA-SIGNL 357
G IP +I L L+ L L N LSGP P+S+ L +LQ L ++ N + TIP + L
Sbjct: 267 GPIPESINRLNQLVLLDLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKGL 326
Query: 358 KWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNR 417
K L + ++ + G IP L + + + N+ G +P + L L + N
Sbjct: 327 KNLMILVLSNTNIQGSIPKSLTRLNSLRVLHLEGNNLTGEIPLEFRDVKHLSELRLNDNS 386
Query: 418 FTGPIPTSLKTCSSIER 434
TGP+P T + R
Sbjct: 387 LTGPVPFERDTVWRMRR 403
Score = 120 bits (301), Expect = 2e-26
Identities = 87/247 (35%), Positives = 128/247 (51%), Gaps = 7/247 (2%)
Query: 156 IPKSIGNL-TNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTN 214
IP +G L ++L L+L N + G PIP E+G L NL L + K++L GSIP +
Sbjct: 150 IPAFLGRLGSSLQTLVLRENGFLG-PIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSG 208
Query: 215 LAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGL 274
L +DLS N L+G IP + L L L L+ N ++GP+P +L + SL + +
Sbjct: 209 LRSLDLSGNRLTGSIPGFV--LPALSVLDLNQNL-LTGPVPPTLTSCGSLIKIDLSRNRV 265
Query: 275 SGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYL-GSNNLSGPIPA-SIGN 332
+G IP+SI L L L L N LSG PS++ L +L L L G+ S IP +
Sbjct: 266 TGPIPESINRLNQLVLLDLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKG 325
Query: 333 LINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSEN 392
L NL +L + N+ G+IP S+ L L V + N L G IP ++ + +++N
Sbjct: 326 LKNLMILVLSNTNIQGSIPKSLTRLNSLRVLHLEGNNLTGEIPLEFRDVKHLSELRLNDN 385
Query: 393 DFVGHLP 399
G +P
Sbjct: 386 SLTGPVP 392
Score = 107 bits (267), Expect = 2e-22
Identities = 85/247 (34%), Positives = 122/247 (48%), Gaps = 9/247 (3%)
Query: 405 GGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDN 464
G SL+ L N F GPIP L ++++ + L N + G I F + L+ LDLS N
Sbjct: 158 GSSLQTLVLRENGFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGLRSLDLSGN 217
Query: 465 KFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLG 524
+ G I P + L ++ N ++G +P L + LS N++TG +P E +
Sbjct: 218 RLTGSI-PGFVLPA-LSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRNRVTGPIP-ESIN 274
Query: 525 GMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGN-ELSGKIPKELVE-LPNLRMLNL 582
+ L L +S N S PS + L LQ L L GN + S IP+ + L NL +L L
Sbjct: 275 RLNQLVLLDLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKGLKNLMILVL 334
Query: 583 SRNKIEGIIPIKFD--SGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQN 640
S I+G IP + L L L GN L G IP D+ LS+L L+ N L+G +P
Sbjct: 335 SNTNIQGSIPKSLTRLNSLRVLHLEGNNLTGEIPLEFRDVKHLSELRLNDNSLTGPVP-- 392
Query: 641 FGRNLVF 647
F R+ V+
Sbjct: 393 FERDTVW 399
Score = 100 bits (249), Expect = 2e-20
Identities = 80/232 (34%), Positives = 109/232 (46%), Gaps = 27/232 (11%)
Query: 456 LQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLT 515
LQ L L +N F G I G NL+ + N+++G IPL F + L L LS N+LT
Sbjct: 161 LQTLVLRENGFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGLRSLDLSGNRLT 220
Query: 516 GKLPMEVLGGMK---------------------SLFDLKISNNHFSDNIPSEIGLLQRLQ 554
G +P VL + SL + +S N + IP I L +L
Sbjct: 221 GSIPGFVLPALSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRNRVTGPIPESINRLNQLV 280
Query: 555 ELDLGGNELSGKIPKELVELPNLRMLNLSRN-KIEGIIPIKFDSGLES---LDLSGNFLK 610
LDL N LSG P L L +L+ L L N K IP GL++ L LS ++
Sbjct: 281 LLDLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKGLKNLMILVLSNTNIQ 340
Query: 611 GNIPTGLADLVRLSKLNLSHNMLSGTIPQNFG--RNLVFVNISDNQLEGPLP 660
G+IP L L L L+L N L+G IP F ++L + ++DN L GP+P
Sbjct: 341 GSIPKSLTRLNSLRVLHLEGNNLTGEIPLEFRDVKHLSELRLNDNSLTGPVP 392
Score = 95.1 bits (235), Expect = 9e-19
Identities = 65/208 (31%), Positives = 107/208 (51%), Gaps = 8/208 (3%)
Query: 73 GLANLGLKGTLHSLTFSSF--PNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDG 130
GL +L L G + + F P L ++D+ N G +P + + ++ + N G
Sbjct: 208 GLRSLDLSGNRLTGSIPGFVLPALSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRNRVTG 267
Query: 131 SIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGK-LN 189
IP+ + L L LD+S+ +L+G P S+ L +L L+L GN IP K L
Sbjct: 268 PIPESINRLNQLVLLDLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKGLK 327
Query: 190 NLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTK 249
NL+ L + +N+ GSIP+ + L +L + L N+L+G IP ++ L L L++N+
Sbjct: 328 NLMILVLSNTNIQGSIPKSLTRLNSLRVLHLEGNNLTGEIPLEFRDVKHLSELRLNDNS- 386
Query: 250 MSGPIP---HSLWNMSSLTVLYFDNIGL 274
++GP+P ++W M LY +N GL
Sbjct: 387 LTGPVPFERDTVWRMRRKLRLY-NNAGL 413
Score = 87.8 bits (216), Expect = 1e-16
Identities = 58/151 (38%), Positives = 80/151 (52%), Gaps = 6/151 (3%)
Query: 516 GKLPMEV---LGGM-KSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKEL 571
G+ P + LG + SL L + N F IP E+G L L+ LDL N L+G IP
Sbjct: 144 GRAPQRIPAFLGRLGSSLQTLVLRENGFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSF 203
Query: 572 VELPNLRMLNLSRNKIEGIIPIKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHN 631
LR L+LS N++ G IP L LDL+ N L G +P L L K++LS N
Sbjct: 204 NRFSGLRSLDLSGNRLTGSIPGFVLPALSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRN 263
Query: 632 MLSGTIPQNFGR--NLVFVNISDNQLEGPLP 660
++G IP++ R LV +++S N+L GP P
Sbjct: 264 RVTGPIPESINRLNQLVLLDLSYNRLSGPFP 294
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precursor
(EC 2.7.1.37)
Length = 817
Score = 157 bits (396), Expect = 2e-37
Identities = 102/349 (29%), Positives = 172/349 (49%), Gaps = 28/349 (8%)
Query: 705 FIALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFSIWSHDGKMMFENIIEAT 764
FIA V+ V + A +++ +P+E +E+ + + S+ + + +++AT
Sbjct: 478 FIAAFFVVEVSFISFAWFFVLKRELRPSELWASEKGYKAMT----SNFRRYSYRELVKAT 533
Query: 765 ANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIK 824
F K +G G G VYK L + VAVKKL V + F +E+ + I
Sbjct: 534 RKF--KVELGRGESGTVYKGVLEDDRHVAVKKLENVRQ------GKEVFQAELSVIGRIN 585
Query: 825 HRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSY 884
H N++++ GFCS LV +++E GSL IL ++ + DWE R N+ GVA L+Y
Sbjct: 586 HMNLVRIWGFCSEGSHRLLVSEYVENGSLANILFSEGGNILLDWEGRFNIALGVAKGLAY 645
Query: 885 LHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG--LHSWTQFAGTFGYAAPE 942
LHH+C +IH D+ +N+LL+ +E ++DFG K L G + + GT GY APE
Sbjct: 646 LHHECLEWVIHCDVKPENILLDQAFEPKITDFGLVKLLNRGGSTQNVSHVRGTLGYIAPE 705
Query: 943 LAQTMEVNEKCDVYSFGVLALETIMGKHPGDLIS------LFLSPSTRPMANNM------ 990
++ + K DVYS+GV+ LE + G +L+ L R ++ +
Sbjct: 706 WVSSLPITAKVDVYSYGVVLLELLTGTRVSELVGGTDEVHSMLRKLVRMLSAKLEGEEQS 765
Query: 991 LLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
+ LD + + + + ++ +LA +CL ++ RP+M + L
Sbjct: 766 WIDGYLDSKLNRPVNYVQARTLI--KLAVSCLEEDRSKRPTMEHAVQTL 812
>D100_ARATH (Q00874) DNA-damage-repair/toleration protein DRT100
precursor
Length = 372
Score = 153 bits (386), Expect = 3e-36
Identities = 107/325 (32%), Positives = 161/325 (48%), Gaps = 15/325 (4%)
Query: 28 ALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSNFISTIGLANLGLKGTLHSLT 87
AL +K S + + + +TW NT+ CK +W GI CD + T ++ L+G
Sbjct: 34 ALNAFKSSLSEPNLGIFNTWSENTDCCK-EWYGISCDPDSGRVT----DISLRGESEDAI 88
Query: 88 FSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKN-NYFDGSIPQEMCTLTGLQFLD 146
F R+ G+I + +L+ ++ L + G IP + +L L+ LD
Sbjct: 89 FQKAG-------RSGYMSGSIDPAVCDLTALTSLVLADWKGITGEIPPCITSLASLRILD 141
Query: 147 ISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIP 206
++ K+ G IP IG L+ L+ L L N S G IP + L L HL + ++ + G IP
Sbjct: 142 LAGNKITGEIPAEIGKLSKLAVLNLAENQMS-GEIPASLTSLIELKHLELTENGITGVIP 200
Query: 207 QEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTV 266
+ G L L+ + L +N L+G IPE+I + +L L LS N + GPIP + NM L++
Sbjct: 201 ADFGSLKMLSRVLLGRNELTGSIPESISGMERLADLDLSKN-HIEGPIPEWMGNMKVLSL 259
Query: 267 LYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPI 326
L D L+G IP S+ + L L N L G+IP G L+ L L N+LSG I
Sbjct: 260 LNLDCNSLTGPIPGSLLSNSGLDVANLSRNALEGTIPDVFGSKTYLVSLDLSHNSLSGRI 319
Query: 327 PASIGNLINLQVLSVQENNLTGTIP 351
P S+ + + L + N L G IP
Sbjct: 320 PDSLSSAKFVGHLDISHNKLCGRIP 344
Score = 152 bits (384), Expect = 5e-36
Identities = 103/270 (38%), Positives = 146/270 (53%), Gaps = 7/270 (2%)
Query: 396 GHLPSQICSGGSLR-LLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYP 454
G + +C +L L+ AD TG IP + + +S+ + L N+I G+I + G
Sbjct: 100 GSIDPAVCDLTALTSLVLADWKGITGEIPPCITSLASLRILDLAGNKITGEIPAEIGKLS 159
Query: 455 KLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQL 514
KL L+L++N+ G+I + + L+ ++ N I+GVIP DF L L + L N+L
Sbjct: 160 KLAVLNLAENQMSGEIPASLTSLIELKHLELTENGITGVIPADFGSLKMLSRVLLGRNEL 219
Query: 515 TGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVEL 574
TG +P E + GM+ L DL +S NH IP +G ++ L L+L N L+G IP L+
Sbjct: 220 TGSIP-ESISGMERLADLDLSKNHIEGPIPEWMGNMKVLSLLNLDCNSLTGPIPGSLLSN 278
Query: 575 PNLRMLNLSRNKIEGIIPIKFDSG--LESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNM 632
L + NLSRN +EG IP F S L SLDLS N L G IP L+ + L++SHN
Sbjct: 279 SGLDVANLSRNALEGTIPDVFGSKTYLVSLDLSHNSLSGRIPDSLSSAKFVGHLDISHNK 338
Query: 633 LSGTIPQNFG-RNLVFVNISDNQ--LEGPL 659
L G IP F +L + SDNQ GPL
Sbjct: 339 LCGRIPTGFPFDHLEATSFSDNQCLCGGPL 368
Score = 146 bits (368), Expect = 3e-34
Identities = 94/250 (37%), Positives = 135/250 (53%), Gaps = 1/250 (0%)
Query: 250 MSGPIPHSLWNMSSLTVLYF-DNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGD 308
MSG I ++ ++++LT L D G++G IP I +L +L+ L L N ++G IP+ IG
Sbjct: 98 MSGSIDPAVCDLTALTSLVLADWKGITGEIPPCITSLASLRILDLAGNKITGEIPAEIGK 157
Query: 309 LKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATN 368
L L L L N +SG IPAS+ +LI L+ L + EN +TG IPA G+LK L+ + N
Sbjct: 158 LSKLAVLNLAENQMSGEIPASLTSLIELKHLELTENGITGVIPADFGSLKMLSRVLLGRN 217
Query: 369 KLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKT 428
+L G IP + + +S+N G +P + + L LLN D N TGPIP SL +
Sbjct: 218 ELTGSIPESISGMERLADLDLSKNHIEGPIPEWMGNMKVLSLLNLDCNSLTGPIPGSLLS 277
Query: 429 CSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNN 488
S ++ L N +EG I FG L LDLS N G+I + + + IS+N
Sbjct: 278 NSGLDVANLSRNALEGTIPDVFGSKTYLVSLDLSHNSLSGRIPDSLSSAKFVGHLDISHN 337
Query: 489 NISGVIPLDF 498
+ G IP F
Sbjct: 338 KLCGRIPTGF 347
Score = 130 bits (328), Expect = 1e-29
Identities = 94/301 (31%), Positives = 148/301 (48%), Gaps = 13/301 (4%)
Query: 176 WSGGPIPPEIGKLNNL----------LHLAIQKSNLVGSIPQEIGFLTNLAYIDLSK-NS 224
W G P+ G++ ++ A + + GSI + LT L + L+
Sbjct: 63 WYGISCDPDSGRVTDISLRGESEDAIFQKAGRSGYMSGSIDPAVCDLTALTSLVLADWKG 122
Query: 225 LSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQN 284
++G IP I +L+ L L L+ N K++G IP + +S L VL +SG IP S+ +
Sbjct: 123 ITGEIPPCITSLASLRILDLAGN-KITGEIPAEIGKLSKLAVLNLAENQMSGEIPASLTS 181
Query: 285 LVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQEN 344
L+ LK L L N ++G IP+ G LK L ++ LG N L+G IP SI + L L + +N
Sbjct: 182 LIELKHLELTENGITGVIPADFGSLKMLSRVLLGRNELTGSIPESISGMERLADLDLSKN 241
Query: 345 NLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICS 404
++ G IP +GN+K L++ + N L G IP L + + +S N G +P S
Sbjct: 242 HIEGPIPEWMGNMKVLSLLNLDCNSLTGPIPGSLLSNSGLDVANLSRNALEGTIPDVFGS 301
Query: 405 GGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDN 464
L L+ HN +G IP SL + + + + N++ G I F + L+ SDN
Sbjct: 302 KTYLVSLDLSHNSLSGRIPDSLSSAKFVGHLDISHNKLCGRIPTGF-PFDHLEATSFSDN 360
Query: 465 K 465
+
Sbjct: 361 Q 361
Score = 120 bits (301), Expect = 2e-26
Identities = 86/274 (31%), Positives = 131/274 (47%), Gaps = 28/274 (10%)
Query: 297 HLSGSIPSTIGDLKNLIKLYLGS-NNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIG 355
++SGSI + DL L L L ++G IP I +L +L++L + N +TG IPA IG
Sbjct: 97 YMSGSIDPAVCDLTALTSLVLADWKGITGEIPPCITSLASLRILDLAGNKITGEIPAEIG 156
Query: 356 NLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADH 415
L L V +A N++ G IP L ++ ++EN G +P+ GSL++L+
Sbjct: 157 KLSKLAVLNLAENQMSGEIPASLTSLIELKHLELTENGITGVIPADF---GSLKMLS--- 210
Query: 416 NRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNW- 474
R+ L N++ G I + +L LDLS N G I P W
Sbjct: 211 ------------------RVLLGRNELTGSIPESISGMERLADLDLSKNHIEGPI-PEWM 251
Query: 475 GKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKI 534
G L + N+++G IP + + L V +LS N L G +P +V G L L +
Sbjct: 252 GNMKVLSLLNLDCNSLTGPIPGSLLSNSGLDVANLSRNALEGTIP-DVFGSKTYLVSLDL 310
Query: 535 SNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIP 568
S+N S IP + + + LD+ N+L G+IP
Sbjct: 311 SHNSLSGRIPDSLSSAKFVGHLDISHNKLCGRIP 344
Score = 80.5 bits (197), Expect = 2e-14
Identities = 47/123 (38%), Positives = 74/123 (59%), Gaps = 4/123 (3%)
Query: 543 IPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDSGLE-- 600
IP I L L+ LDL GN+++G+IP E+ +L L +LNL+ N++ G IP S +E
Sbjct: 127 IPPCITSLASLRILDLAGNKITGEIPAEIGKLSKLAVLNLAENQMSGEIPASLTSLIELK 186
Query: 601 SLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFG--RNLVFVNISDNQLEGP 658
L+L+ N + G IP L LS++ L N L+G+IP++ L +++S N +EGP
Sbjct: 187 HLELTENGITGVIPADFGSLKMLSRVLLGRNELTGSIPESISGMERLADLDLSKNHIEGP 246
Query: 659 LPK 661
+P+
Sbjct: 247 IPE 249
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase receptor
precursor (EC 2.7.1.37) (S-receptor kinase) (SRK)
Length = 849
Score = 146 bits (369), Expect = 3e-34
Identities = 110/380 (28%), Positives = 181/380 (46%), Gaps = 39/380 (10%)
Query: 690 ATSHSRKRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPN-------EESQTEEVQR 742
A ++KR + + + +G +L+L ++ L R K + + +Q +
Sbjct: 434 AADIAKKRNASGKIISLTVGVSVLLLLIMFCLWKRKQKRAKASAISIANTQRNQNLPMNE 493
Query: 743 GVLFSIWSHDGKMMFEN----------IIEATANFDDKYLVGVGSQGNVYKAELSEGLVV 792
VL S G+ FE +++AT NF +G G G VYK L +G +
Sbjct: 494 MVLSSKREFSGEYKFEELELPLIEMETVVKATENFSSCNKLGQGGFGIVYKGRLLDGKEI 553
Query: 793 AVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGS 852
AVK+L + S + FM+E+ + ++H N++++ G C L+Y++LE S
Sbjct: 554 AVKRL-----SKTSVQGTDEFMNEVTLIARLQHINLVQVLGCCIEGDEKMLIYEYLENLS 608
Query: 853 LDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAH 912
LD L T+ +W +R ++ GVA L YLH D IIHRD+ N+LL+ +
Sbjct: 609 LDSYLFGKTRRSKLNWNERFDITNGVARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPK 668
Query: 913 VSDFGTAK-FLKPGLHSWT-QFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGK- 969
+SDFG A+ F + + T + GT+GY +PE A +EK DV+SFGV+ LE + GK
Sbjct: 669 ISDFGMARIFERDETEANTMKVVGTYGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKK 728
Query: 970 --------HPGDLISLFLSPSTRPMANNM---LLTDVLDQRPQQVMEPIDEEVILIARLA 1018
+ DL+S S A + ++ D L +P + +P +EV+ ++
Sbjct: 729 NRGFYNLDYENDLLSYVWSRWKEGRALEIVDPVIVDSLSSQP-SIFQP--QEVLKCIQIG 785
Query: 1019 FACLSQNPRLRPSMGQVCKM 1038
C+ + RP+M V M
Sbjct: 786 LLCVQELAEHRPAMSSVVWM 805
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 144 bits (364), Expect = 1e-33
Identities = 108/341 (31%), Positives = 177/341 (51%), Gaps = 56/341 (16%)
Query: 733 EESQTEEVQRGVLFSIWSHDGKMM---------FENIIEATANFDDKYLVGVGSQGNVYK 783
++SQ ++ G++ S GK++ F ++ AT NF ++G G G VY+
Sbjct: 47 QQSQFSDISTGII----SDSGKLLESPNLKVYNFLDLKTATKNFKPDSMLGQGGFGKVYR 102
Query: 784 ----------AELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHG 833
+ + G++VA+K+L+ E + F+ + SE+ L + HRN++KL G
Sbjct: 103 GWVDATTLAPSRVGSGMIVAIKRLN---SESVQGFAE--WRSEVNFLGMLSHRNLVKLLG 157
Query: 834 FCSHSKFSFLVYKFLEGGSLDQIL--NNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSP 891
+C K LVY+F+ GSL+ L ND F W+ R+ +V G A L++LH
Sbjct: 158 YCREDKELLLVYEFMPKGSLESHLFRRNDP----FPWDLRIKIVIGAARGLAFLH-SLQR 212
Query: 892 PIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG---LHSWTQFAGTFGYAAPELAQTME 948
+I+RD + N+LL+ +Y+A +SDFG AK L P H T+ GT+GYAAPE T
Sbjct: 213 EVIYRDFKASNILLDSNYDAKLSDFGLAK-LGPADEKSHVTTRIMGTYGYAAPEYMATGH 271
Query: 949 VNEKCDVYSFGVLALETIMG------KHPGDLISL--FLSPSTRPMANNMLLTDVLDQ-- 998
+ K DV++FGV+ LE + G K P SL +L P ++N + ++D+
Sbjct: 272 LYVKSDVFAFGVVLLEIMTGLTAHNTKRPRGQESLVDWLRPE---LSNKHRVKQIMDKGI 328
Query: 999 RPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
+ Q + E +AR+ +C+ +P+ RP M +V ++L
Sbjct: 329 KGQYTTKVATE----MARITLSCIEPDPKNRPHMKEVVEVL 365
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 144 bits (363), Expect = 1e-33
Identities = 84/217 (38%), Positives = 119/217 (54%), Gaps = 9/217 (4%)
Query: 757 FENIIEATANFDDKYLVGVGSQGNVYKAEL-SEGLVVAVKKLHLVTDEEMSCFSSKSFMS 815
F + AT NF +G G G VYK L S G VVAVK+L + ++ F+
Sbjct: 76 FRELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQL-----DRNGLQGNREFLV 130
Query: 816 EIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNN-DTQAVAFDWEKRVNV 874
E+ L+ + H N++ L G+C+ LVY+F+ GSL+ L++ A DW R+ +
Sbjct: 131 EVLMLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKI 190
Query: 875 VKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG--LHSWTQF 932
G A L +LH +PP+I+RD S N+LL+ + +SDFG AK G H T+
Sbjct: 191 AAGAAKGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTRV 250
Query: 933 AGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGK 969
GT+GY APE A T ++ K DVYSFGV+ LE I G+
Sbjct: 251 MGTYGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGR 287
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 144 bits (362), Expect = 2e-33
Identities = 102/304 (33%), Positives = 152/304 (49%), Gaps = 32/304 (10%)
Query: 757 FENIIEATANFDDKYLVGVGSQGNVYKAELSE----------GLVVAVKKLHLVTDEEMS 806
F + AT NF ++G G G+V+K + E G+V+AVKKL+ +
Sbjct: 59 FAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTASKPGTGVVIAVKKLN-----QDG 113
Query: 807 CFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQ-ILNNDTQAVA 865
+ +++E+ L H N++KL G+C + LVY+F+ GSL+ + +
Sbjct: 114 WQGHQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGSYFQP 173
Query: 866 FDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG 925
W R+ V G A L++LH+ +I+RD + N+LL+ +Y A +SDFG AK G
Sbjct: 174 LSWTLRLKVALGAAKGLAFLHN-AETSVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPTG 232
Query: 926 --LHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKH-------PGDLIS 976
H T+ GT+GYAAPE T + K DVYS+GV+ LE + G+ PG+
Sbjct: 233 DKSHVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVLSGRRAVDKNRPPGE--- 289
Query: 977 LFLSPSTRP-MANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQV 1035
L RP +AN L V+D R Q EE +A LA CL+ +LRP+M +V
Sbjct: 290 QKLVEWARPLLANKRKLFRVIDNRLQDQYSM--EEACKVATLALRCLTFEIKLRPNMNEV 347
Query: 1036 CKML 1039
L
Sbjct: 348 VSHL 351
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 143 bits (360), Expect = 3e-33
Identities = 81/217 (37%), Positives = 120/217 (54%), Gaps = 9/217 (4%)
Query: 757 FENIIEATANFDDKYLVGVGSQGNVYKAELSE-GLVVAVKKLHLVTDEEMSCFSSKSFMS 815
F+ + EAT NF +G G G V+K + + VVA+K+L + + F+
Sbjct: 93 FQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQL-----DRNGVQGIREFVV 147
Query: 816 EIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNN-DTQAVAFDWEKRVNV 874
E+ TL+ H N++KL GFC+ LVY+++ GSL+ L+ + DW R+ +
Sbjct: 148 EVLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRMKI 207
Query: 875 VKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG--LHSWTQF 932
G A L YLH +PP+I+RD+ N+LL DY+ +SDFG AK G H T+
Sbjct: 208 AAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVSTRV 267
Query: 933 AGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGK 969
GT+GY AP+ A T ++ K D+YSFGV+ LE I G+
Sbjct: 268 MGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGR 304
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 127,027,435
Number of Sequences: 164201
Number of extensions: 5598390
Number of successful extensions: 22119
Number of sequences better than 10.0: 1858
Number of HSP's better than 10.0 without gapping: 816
Number of HSP's successfully gapped in prelim test: 1042
Number of HSP's that attempted gapping in prelim test: 13637
Number of HSP's gapped (non-prelim): 3799
length of query: 1067
length of database: 59,974,054
effective HSP length: 121
effective length of query: 946
effective length of database: 40,105,733
effective search space: 37940023418
effective search space used: 37940023418
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC133779.11


