

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
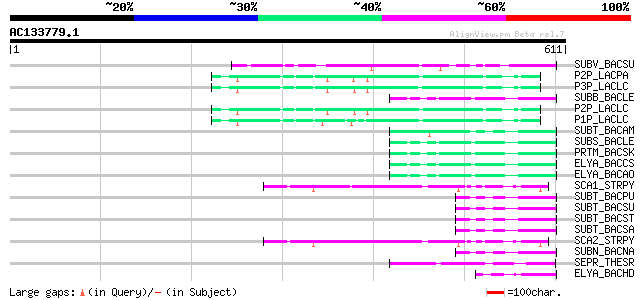
Query= AC133779.1 + phase: 0 /pseudo
(611 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SUBV_BACSU (P29141) Minor extracellular protease vpr precursor (... 101 7e-21
P2P_LACPA (Q02470) PII-type proteinase precursor (EC 3.4.21.96) ... 59 5e-08
P3P_LACLC (P15292) PIII-type proteinase precursor (EC 3.4.21.96)... 58 8e-08
SUBB_BACLE (P29599) Subtilisin BL (EC 3.4.21.62) (Alkaline prote... 55 7e-07
P2P_LACLC (P15293) PII-type proteinase precursor (EC 3.4.21.96) ... 55 7e-07
P1P_LACLC (P16271) PI-type proteinase precursor (EC 3.4.21.-) (W... 53 3e-06
SUBT_BACAM (P00782) Subtilisin BPN' precursor (EC 3.4.21.62) (Su... 52 3e-06
SUBS_BACLE (P29600) Subtilisin Savinase (EC 3.4.21.62) (Alkaline... 51 8e-06
PRTM_BACSK (Q99405) M-protease (EC 3.4.21.-) 51 8e-06
ELYA_BACCS (P41362) Alkaline protease precursor (EC 3.4.21.-) 51 8e-06
ELYA_BACAO (P27693) Alkaline protease precursor (EC 3.4.21.-) 51 8e-06
SCA1_STRPY (P15926) C5a peptidase precursor (EC 3.4.21.-) (SCP) 50 2e-05
SUBT_BACPU (P07518) Subtilisin (EC 3.4.21.62) (Alkaline mesenter... 49 4e-05
SUBT_BACSU (P04189) Subtilisin E precursor (EC 3.4.21.62) 49 5e-05
SUBT_BACST (P29142) Subtilisin J precursor (EC 3.4.21.62) 49 5e-05
SUBT_BACSA (P00783) Subtilisin amylosacchariticus precursor (EC ... 49 5e-05
SCA2_STRPY (P58099) C5a peptidase precursor (EC 3.4.21.-) (SCP) 49 5e-05
SUBN_BACNA (P35835) Subtilisin NAT precursor (EC 3.4.21.62) 48 9e-05
SEPR_THESR (P80146) Extracellular serine proteinase precursor (E... 45 7e-04
ELYA_BACHD (P41363) Thermostable alkaline protease precursor (EC... 45 7e-04
>SUBV_BACSU (P29141) Minor extracellular protease vpr precursor (EC
3.4.21.-)
Length = 806
Score = 101 bits (251), Expect = 7e-21
Identities = 103/366 (28%), Positives = 164/366 (44%), Gaps = 51/366 (13%)
Query: 245 NGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNP 304
NGTIKG +P + + Y+ L +V+A +++A+ DGAD++++S G N NP
Sbjct: 244 NGTIKGVAPDATLLAYRV---LGPGGSGTTENVIAGVERAVQDGADVMNLSLGNSLN-NP 299
Query: 305 EVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMT 364
+ + + A++ ++ V S GN GP +TV + R+ SV
Sbjct: 300 DWATSTALD----WAMSEGVVAVTSNGNSGPNG----------WTVGSPGTSREAISVGA 345
Query: 365 INNKTLTGASLFVNLPPNQDFLIIISTDAKFAN---VTDVDAQFCRPGTLDPSKVNGKVV 421
A F + + D K N V V+A + + GKV
Sbjct: 346 TQLPLNEYAVTFGSYSSAKVMGYNKEDDVKALNNKEVELVEAGIGEAKDFEGKDLTGKVA 405
Query: 422 ACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHV-VSTINYY---DARSI 477
R G I + + A AGA+G+++ N + G+ P + V TI + +
Sbjct: 406 VVKR-GSIAFVDKADNAKKAGAIGMVVYNN--LSGEIEANVPGMSVPTIKLSLEDGEKLV 462
Query: 478 TTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPG 537
+ K E T T +++ + AL + +A FSSRGP + +++KPD++APG
Sbjct: 463 SALKAGE-------TKTTFKLTVSKALGEQ-----VADFSSRGP-VMDTWMIKPDISAPG 509
Query: 538 VNILAAYSLLASVSNLVT-DNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIK 596
VNI VS + T D + + +QGTSM+ PH+ G +IK P WS IK
Sbjct: 510 VNI---------VSTIPTHDPDHPYGYGSKQGTSMASPHIAGAVAVIKQAKPKWSVEQIK 560
Query: 597 SAIMTT 602
+AIM T
Sbjct: 561 AAIMNT 566
>P2P_LACPA (Q02470) PII-type proteinase precursor (EC 3.4.21.96)
(Lactocepin) (Cell wall-associated serine proteinase)
(LP151)
Length = 1902
Score = 58.5 bits (140), Expect = 5e-08
Identities = 88/384 (22%), Positives = 151/384 (38%), Gaps = 53/384 (13%)
Query: 223 GTHTLSTAGGNFVQNATIFGIGNGTIK---GGSPRSRVATYKACWSLTDVVDCFGADVLA 279
G H G N G G+ K G +P +++ K + A +++
Sbjct: 282 GMHVAGIIGAN--------GTGDDPTKSVVGVAPEAQLLAMKVFTNSDTSATTGSATLVS 333
Query: 280 AIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGS 339
AI+ + GAD++++S G ++ + + EI+ +A V SAGN G T GS
Sbjct: 334 AIEDSAKIGADVLNMSLGS--DSGNQTLEDPEIA-AVQNANESGTAAVISAGNSG-TSGS 389
Query: 340 VTNVAPWVF-------TVAASTLDRDFSSVMTINNKTLTGASLFV----NLPPNQDFLII 388
T + V R ++V + N + ++ + +L + + +
Sbjct: 390 ATQGVNKDYYGLQDNEMVGTPGTSRGATTVASAENTDVISQAVTITDGKDLQLGPETIQL 449
Query: 389 ISTD-------AKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSA 441
S D KF V D + D + +A + G++N + + A +A
Sbjct: 450 SSNDFTGSFDQKKFYVVKDASGDLSKGAAADYTADAKGKIAIVKRGELNFADKQKYAQAA 509
Query: 442 GAVGVIMRNQPEVDG-KTLLAEPHVVSTINYYDARSITTPKGSEITPEDIKTNATIRMSP 500
GA G+I+ N DG T L + +T + S T K + + ++++
Sbjct: 510 GAAGLIIVNN---DGTATPLTSIRLTTTFPTFGLSSKTGQKLVDWVTAHPDDSLGVKIAL 566
Query: 501 ANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRG 560
N + M+ F+S GP V KPD+TAPG NI + T N G
Sbjct: 567 TLLPNQKYTEDKMSDFTSYGP--VSNLSFKPDITAPGGNIWS------------TQNNNG 612
Query: 561 FPFNIQQGTSMSCPHVVGTAGLIK 584
+ GTSM+ P + G+ L+K
Sbjct: 613 --YTNMSGTSMASPFIAGSQALLK 634
>P3P_LACLC (P15292) PIII-type proteinase precursor (EC 3.4.21.96)
(Lactocepin) (Cell wall-associated serine proteinase)
Length = 1902
Score = 57.8 bits (138), Expect = 8e-08
Identities = 88/384 (22%), Positives = 152/384 (38%), Gaps = 53/384 (13%)
Query: 223 GTHTLSTAGGNFVQNATIFGIGNGTIK---GGSPRSRVATYKACWSLTDVVDCFGADVLA 279
G H G N G G+ K G +P +++ K + A V++
Sbjct: 282 GMHVAGIIGAN--------GTGDDPAKSVVGVAPEAQLLAMKVFSNSDTSAKTGSATVVS 333
Query: 280 AIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGS 339
AI+ + GAD++++S G N+ + + E++ +A V SAGN G T GS
Sbjct: 334 AIEDSAKIGADVLNMSLGS--NSGNQTLEDPELA-AVQNANESGTAAVISAGNSG-TSGS 389
Query: 340 VTNVAPWVF-------TVAASTLDRDFSSVMTINNKTLTGASLFVN----LPPNQDFLII 388
T + V + R ++V + N + ++ + L + + +
Sbjct: 390 ATEGVNKDYYGLQDNEMVGSPGTSRGATTVASAENTDVITQAVTITDGTGLQLGPETIQL 449
Query: 389 ISTD-------AKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSA 441
S D KF V D + D + +A + G+ + + + A +A
Sbjct: 450 SSHDFTGSFDQKKFYIVKDASGNLSKGALADYTADAKGKIAIVKRGEFSFDDKQKYAQAA 509
Query: 442 GAVGVIMRNQPEVDGK-TLLAEPHVVSTINYYDARSITTPKGSEITPEDIKTNATIRMSP 500
GA G+I+ N DG T + + +T + S+T K + + ++++
Sbjct: 510 GAAGLIIVN---TDGTATPMTSIALTTTFPTFGLSSVTGQKLVDWVTAHPDDSLGVKITL 566
Query: 501 ANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRG 560
A N + M+ F+S GP V KPD+TAPG NI + T N G
Sbjct: 567 AMLPNQKYTEDKMSDFTSYGP--VSNLSFKPDITAPGGNIWS------------TQNNNG 612
Query: 561 FPFNIQQGTSMSCPHVVGTAGLIK 584
+ GTSM+ P + G+ L+K
Sbjct: 613 --YTNMSGTSMASPFIAGSQALLK 634
>SUBB_BACLE (P29599) Subtilisin BL (EC 3.4.21.62) (Alkaline
protease)
Length = 269
Score = 54.7 bits (130), Expect = 7e-07
Identities = 51/184 (27%), Positives = 76/184 (40%), Gaps = 28/184 (15%)
Query: 419 KVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSIT 478
KV+ D G I+SIA+G E AG G+ + N L P +T+ A +
Sbjct: 92 KVLGADGRGAISSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLE--QAVNSA 140
Query: 479 TPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGV 538
T +G + + A+ PA N V A+ + Y D+ APGV
Sbjct: 141 TSRGVLVVAASGNSGASSISYPARYANAMA---VGATDQNNNRASFSQYGAGLDIVAPGV 197
Query: 539 NILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSA 598
N+ + Y G + GTSM+ PHV G A L+K +P+WS I++
Sbjct: 198 NVQSTYP--------------GSTYASLNGTSMATPHVAGAAALVKQKNPSWSNVQIRNH 243
Query: 599 IMTT 602
+ T
Sbjct: 244 LKNT 247
>P2P_LACLC (P15293) PII-type proteinase precursor (EC 3.4.21.96)
(Lactocepin) (Cell wall-associated serine proteinase)
(LP151)
Length = 1902
Score = 54.7 bits (130), Expect = 7e-07
Identities = 85/384 (22%), Positives = 150/384 (38%), Gaps = 53/384 (13%)
Query: 223 GTHTLSTAGGNFVQNATIFGIGNGTIK---GGSPRSRVATYKACWSLTDVVDCFGADVLA 279
G H G N G G+ K G +P +++ K + A +++
Sbjct: 282 GMHVAGIIGAN--------GTGDDPAKSVVGVAPEAQLLAMKVFTNSDTSATTGSATLVS 333
Query: 280 AIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGS 339
AI+ + GAD++++S G ++ + + E++ +A V SAGN G T GS
Sbjct: 334 AIEDSAKIGADVLNMSLGS--DSGNQTLEDPELA-AVQNANESGTAAVISAGNSG-TSGS 389
Query: 340 VTNVAPWVF-------TVAASTLDRDFSSVMTINNKTLTGASLFVN----LPPNQDFLII 388
T + V R ++V + N + ++ + L + + +
Sbjct: 390 ATEGVNKDYYGLQDNEMVGTPGTSRGATTVASAENTDVITQAVTITDGTGLQLGPETIQL 449
Query: 389 ISTD-------AKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSA 441
S D KF V D + D + +A + G++ + + A +A
Sbjct: 450 SSNDFTGSFDQKKFYVVKDASGNLSKGKVADYTADAKGKIAIVKRGELTFADKQKYAQAA 509
Query: 442 GAVGVIMRNQPEVDGK-TLLAEPHVVSTINYYDARSITTPKGSEITPEDIKTNATIRMSP 500
GA G+I+ N DG T + + +T + S+T K + + ++++
Sbjct: 510 GAAGLIIVNN---DGTATPVTSMALTTTFPTFGLSSVTGQKLVDWVAAHPDDSLGVKIAL 566
Query: 501 ANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRG 560
N + M+ F+S GP V KPD+TAPG NI + T N G
Sbjct: 567 TLVPNQKYTEDKMSDFTSYGP--VSNLSFKPDITAPGGNIWS------------TQNNNG 612
Query: 561 FPFNIQQGTSMSCPHVVGTAGLIK 584
+ GTSM+ P + G+ L+K
Sbjct: 613 --YTNMSGTSMASPFIAGSQALLK 634
>P1P_LACLC (P16271) PI-type proteinase precursor (EC 3.4.21.-)
(Wall-associated serine proteinase)
Length = 1902
Score = 52.8 bits (125), Expect = 3e-06
Identities = 89/388 (22%), Positives = 152/388 (38%), Gaps = 61/388 (15%)
Query: 223 GTHTLSTAGGNFVQNATIFGIGNGTIK---GGSPRSRVATYKACWSLTDVVDCFGADVLA 279
G H G N G G+ K G +P +++ K + + +++
Sbjct: 282 GMHVAGIIGAN--------GTGDDPAKSVVGVAPEAQLLAMKVFTNSDTSATTGSSTLVS 333
Query: 280 AIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGS 339
AI+ + GAD++++S G ++ + + E++ +A V SAGN G T GS
Sbjct: 334 AIEDSAKIGADVLNMSLGS--DSGNQTLEDPELA-AVQNANESGTAAVISAGNSG-TSGS 389
Query: 340 VTN-------------------VAPWVFTVAASTLDRDFSSVMTINNKTLTGASL---FV 377
T + TVA++ + +TI + T G L +
Sbjct: 390 ATEGVNKDYYGLQDNEMVGTPGTSRGATTVASAENTDVITQAVTITDGT--GLQLGPGTI 447
Query: 378 NLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQE 437
L N DF KF V D + D + +A + G+++ + +
Sbjct: 448 QLSSN-DFTGSFD-QKKFYVVKDASGNLSKGALADYTADAKGKIAIVKRGELSFDDKQKY 505
Query: 438 ALSAGAVGVIMRNQPEVDGK-TLLAEPHVVSTINYYDARSITTPKGSEITPEDIKTNATI 496
A +AGA G+I+ N DG T + + +T + S+T K + + +
Sbjct: 506 AQAAGAAGLIIVNN---DGTATPVTSMALTTTFPTFGLSSVTGQKLVDWVTAHPDDSLGV 562
Query: 497 RMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTD 556
+++ N + M+ F+S GP V KPD+TAPG NI + T
Sbjct: 563 KIALTLVPNQKYTEDKMSDFTSYGP--VSNLSFKPDITAPGGNIWS------------TQ 608
Query: 557 NRRGFPFNIQQGTSMSCPHVVGTAGLIK 584
N G + GTSM+ P + G+ L+K
Sbjct: 609 NNNG--YTNMSGTSMASPFIAGSQALLK 634
>SUBT_BACAM (P00782) Subtilisin BPN' precursor (EC 3.4.21.62)
(Subtilisin Novo) (Subtilisin DFE) (Alkaline protease)
Length = 382
Score = 52.4 bits (124), Expect = 3e-06
Identities = 54/188 (28%), Positives = 75/188 (39%), Gaps = 32/188 (17%)
Query: 419 KVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLL---AEPHVVSTINYYDAR 475
KV+ D G+ + I G E A + VI + G L + V S + A
Sbjct: 201 KVLGADGSGQYSWIINGIEWAIANNMDVINMSLGGPSGSAALKAAVDKAVASGVVVVAAA 260
Query: 476 SITTPKGSEITPE-DIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVT 534
GS T K + I + ++ N R ASFSS GP DV
Sbjct: 261 GNEGTSGSSSTVGYPGKYPSVIAVGAVDSSNQR------ASFSSVGPEL--------DVM 306
Query: 535 APGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAA 594
APGV+I + G + GTSM+ PHV G A LI + HPNW+
Sbjct: 307 APGVSIQSTLP--------------GNKYGAYNGTSMASPHVAGAAALILSKHPNWTNTQ 352
Query: 595 IKSAIMTT 602
++S++ T
Sbjct: 353 VRSSLENT 360
Score = 37.0 bits (84), Expect = 0.15
Identities = 36/115 (31%), Positives = 54/115 (46%), Gaps = 16/115 (13%)
Query: 277 VLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPT 336
++ I+ AI + D+I++S GG + D+ A+A +++VA+AGNEG T
Sbjct: 214 IINGIEWAIANNMDVINMSLGGPSGSAALKAAVDK-------AVASGVVVVAAAGNEG-T 265
Query: 337 PGSVTNVA-----PWVFTVAA---STLDRDFSSVMTINNKTLTGASLFVNLPPNQ 383
GS + V P V V A S FSSV + G S+ LP N+
Sbjct: 266 SGSSSTVGYPGKYPSVIAVGAVDSSNQRASFSSVGPELDVMAPGVSIQSTLPGNK 320
>SUBS_BACLE (P29600) Subtilisin Savinase (EC 3.4.21.62) (Alkaline
protease)
Length = 269
Score = 51.2 bits (121), Expect = 8e-06
Identities = 49/184 (26%), Positives = 74/184 (39%), Gaps = 28/184 (15%)
Query: 419 KVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSIT 478
KV+ G ++SIA+G E AG G+ + N L P +T+ A +
Sbjct: 92 KVLGASGSGSVSSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLE--QAVNSA 140
Query: 479 TPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGV 538
T +G + + A PA N V A+ + Y D+ APGV
Sbjct: 141 TSRGVLVVAASGNSGAGSISYPARYANAMA---VGATDQNNNRASFSQYGAGLDIVAPGV 197
Query: 539 NILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSA 598
N+ + Y G + GTSM+ PHV G A L+K +P+WS I++
Sbjct: 198 NVQSTYP--------------GSTYASLNGTSMATPHVAGAAALVKQKNPSWSNVQIRNH 243
Query: 599 IMTT 602
+ T
Sbjct: 244 LKNT 247
>PRTM_BACSK (Q99405) M-protease (EC 3.4.21.-)
Length = 269
Score = 51.2 bits (121), Expect = 8e-06
Identities = 49/184 (26%), Positives = 74/184 (39%), Gaps = 28/184 (15%)
Query: 419 KVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSIT 478
KV+ G ++SIA+G E AG G+ + N L P +T+ A +
Sbjct: 92 KVLGASGSGSVSSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLE--QAVNSA 140
Query: 479 TPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGV 538
T +G + + A PA N V A+ + Y D+ APGV
Sbjct: 141 TSRGVLVVAASGNSGAGSISYPARYANAMA---VGATDQNNNRASFSQYGAGLDIVAPGV 197
Query: 539 NILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSA 598
N+ + Y G + GTSM+ PHV G A L+K +P+WS I++
Sbjct: 198 NVQSTYP--------------GSTYASLNGTSMATPHVAGVAALVKQKNPSWSNVQIRNH 243
Query: 599 IMTT 602
+ T
Sbjct: 244 LKNT 247
>ELYA_BACCS (P41362) Alkaline protease precursor (EC 3.4.21.-)
Length = 380
Score = 51.2 bits (121), Expect = 8e-06
Identities = 49/184 (26%), Positives = 74/184 (39%), Gaps = 28/184 (15%)
Query: 419 KVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSIT 478
KV+ G ++SIA+G E AG G+ + N L P +T+ A +
Sbjct: 203 KVLGASGSGSVSSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLE--QAVNSA 251
Query: 479 TPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGV 538
T +G + + A PA N V A+ + Y D+ APGV
Sbjct: 252 TSRGVLVVAASGNSGAGSISYPARYANAMA---VGATDQNNNRASFSQYGAGLDIVAPGV 308
Query: 539 NILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSA 598
N+ + Y G + GTSM+ PHV G A L+K +P+WS I++
Sbjct: 309 NVQSTYP--------------GSTYASLNGTSMATPHVAGAAALVKQKNPSWSNVQIRNH 354
Query: 599 IMTT 602
+ T
Sbjct: 355 LKNT 358
>ELYA_BACAO (P27693) Alkaline protease precursor (EC 3.4.21.-)
Length = 380
Score = 51.2 bits (121), Expect = 8e-06
Identities = 49/184 (26%), Positives = 74/184 (39%), Gaps = 28/184 (15%)
Query: 419 KVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSIT 478
KV+ G ++SIA+G E AG G+ + N L P +T+ A +
Sbjct: 203 KVLGASGSGSVSSIAQGLEW--AGNNGMHVANLS-------LGSPSPSATLE--QAVNSA 251
Query: 479 TPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGV 538
T +G + + A PA N V A+ + Y D+ APGV
Sbjct: 252 TSRGVLVVAASGNSGAGSISYPARYANAMA---VGATDQNNNRASFSQYGAGLDIVAPGV 308
Query: 539 NILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSA 598
N+ + Y G + GTSM+ PHV G A L+K +P+WS I++
Sbjct: 309 NVQSTYP--------------GSTYASLNGTSMATPHVAGAAALVKQKNPSWSNVQIRNH 354
Query: 599 IMTT 602
+ T
Sbjct: 355 LKNT 358
>SCA1_STRPY (P15926) C5a peptidase precursor (EC 3.4.21.-) (SCP)
Length = 1167
Score = 50.1 bits (118), Expect = 2e-05
Identities = 81/327 (24%), Positives = 137/327 (41%), Gaps = 45/327 (13%)
Query: 280 AIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNE----GP 335
AI A+ GA +I++S G + DE +A ++ + +V SAGN+ G
Sbjct: 245 AIRDAVNLGAKVINMSFGNAALAYANL--PDETKKAFDYAKSKGVSIVTSAGNDSSFGGK 302
Query: 336 TPGSVTNVAPWVFTVAASTLDRDFSSVMTINNKTLTGASLFVNLPPNQDFLIIISTDAKF 395
T + + + + D + +K LT ++ V QD + + + +F
Sbjct: 303 TRLPLADHPDYGVVGTPAAADSTLTVASYSPDKQLTETAM-VKTDDQQDKEMPVLSTNRF 361
Query: 396 ANVTDVDAQFCRPGTL--DPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPE 453
D + G D V GK+ +R G I+ + A AGAVGV++ + +
Sbjct: 362 EPNKAYDYAYANRGMKEDDFKDVKGKIALIER-GDIDFKDKVANAKKAGAVGVLIYDNQD 420
Query: 454 VDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDIKT---NATIRMSPANALNGRKPA 510
P + ++ A I+ G + KT NAT ++ P +G K
Sbjct: 421 K------GFPIELPNVDQMPAAFISRKDGLLLKDNPQKTITFNATPKVLPT--ASGTK-- 470
Query: 511 PVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTS 570
++ FSS G +KPD+ APG +IL++ V +N+ + GTS
Sbjct: 471 --LSRFSSWG--LTADGNIKPDIAAPGQDILSS----------VANNK----YAKLSGTS 512
Query: 571 MSCPHVVGTAGLI----KTLHPNWSPA 593
MS P V G GL+ +T +P+ +P+
Sbjct: 513 MSAPLVAGIMGLLQKQYETQYPDMTPS 539
>SUBT_BACPU (P07518) Subtilisin (EC 3.4.21.62) (Alkaline
mesentericopeptidase)
Length = 275
Score = 48.9 bits (115), Expect = 4e-05
Identities = 36/112 (32%), Positives = 49/112 (43%), Gaps = 28/112 (25%)
Query: 491 KTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASV 550
K +TI + N+ N R ASFSS G DV APGV+I +
Sbjct: 170 KYPSTIAVGAVNSANQR------ASFSSAGSEL--------DVMAPGVSIQSTLP----- 210
Query: 551 SNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
G + GTSM+ PHV G A LI + HP W+ A ++ + +T
Sbjct: 211 ---------GGTYGAYNGTSMATPHVAGAAALILSKHPTWTNAQVRDRLEST 253
Score = 33.9 bits (76), Expect = 1.3
Identities = 46/185 (24%), Positives = 77/185 (40%), Gaps = 38/185 (20%)
Query: 209 LPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT-IKGGSPRSRVATYKACWSLT 267
+PS +D GTH T I + N + G +P S + K
Sbjct: 51 VPSETNPYQDGSSHGTHVAGT----------IAALNNSIGVLGVAPSSALYAVK------ 94
Query: 268 DVVDCFGAD----VLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARN 323
V+D G+ ++ I+ AI + D+I++S GG + D+ A++
Sbjct: 95 -VLDSTGSGQYSWIINGIEWAISNNMDVINMSLGGPTGSTALKTVVDK-------AVSSG 146
Query: 324 ILLVASAGNEGPTPGSVTNV---APWVFTVAASTLD-----RDFSSVMTINNKTLTGASL 375
I++ A+AGNEG + GS + V A + T+A ++ FSS + + G S+
Sbjct: 147 IVVAAAAGNEG-SSGSTSTVGYPAKYPSTIAVGAVNSANQRASFSSAGSELDVMAPGVSI 205
Query: 376 FVNLP 380
LP
Sbjct: 206 QSTLP 210
>SUBT_BACSU (P04189) Subtilisin E precursor (EC 3.4.21.62)
Length = 381
Score = 48.5 bits (114), Expect = 5e-05
Identities = 36/112 (32%), Positives = 49/112 (43%), Gaps = 28/112 (25%)
Query: 491 KTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASV 550
K +TI + N+ N R ASFSS G DV APGV+I +
Sbjct: 276 KYPSTIAVGAVNSSNQR------ASFSSAGSEL--------DVMAPGVSIQSTLP----- 316
Query: 551 SNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
G + GTSM+ PHV G A LI + HP W+ A ++ + +T
Sbjct: 317 ---------GGTYGAYNGTSMATPHVAGAAALILSKHPTWTNAQVRDRLEST 359
Score = 33.9 bits (76), Expect = 1.3
Identities = 46/185 (24%), Positives = 77/185 (40%), Gaps = 38/185 (20%)
Query: 209 LPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT-IKGGSPRSRVATYKACWSLT 267
+PS +D GTH T I + N + G SP + + K
Sbjct: 157 VPSETNPYQDGSSHGTHVAGT----------IAALNNSIGVLGVSPSASLYAVK------ 200
Query: 268 DVVDCFGAD----VLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARN 323
V+D G+ ++ I+ AI + D+I++S GG + D+ A++
Sbjct: 201 -VLDSTGSGQYSWIINGIEWAISNNMDVINMSLGGPTGSTALKTVVDK-------AVSSG 252
Query: 324 ILLVASAGNEGPTPGSVTNV---APWVFTVAASTLD-----RDFSSVMTINNKTLTGASL 375
I++ A+AGNEG + GS + V A + T+A ++ FSS + + G S+
Sbjct: 253 IVVAAAAGNEG-SSGSTSTVGYPAKYPSTIAVGAVNSSNQRASFSSAGSELDVMAPGVSI 311
Query: 376 FVNLP 380
LP
Sbjct: 312 QSTLP 316
>SUBT_BACST (P29142) Subtilisin J precursor (EC 3.4.21.62)
Length = 381
Score = 48.5 bits (114), Expect = 5e-05
Identities = 36/112 (32%), Positives = 49/112 (43%), Gaps = 28/112 (25%)
Query: 491 KTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASV 550
K +TI + N+ N R ASFSS G DV APGV+I +
Sbjct: 276 KYPSTIAVGAVNSSNQR------ASFSSAGSEL--------DVMAPGVSIQSTLP----- 316
Query: 551 SNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
G + GTSM+ PHV G A LI + HP W+ A ++ + +T
Sbjct: 317 ---------GGTYGAYNGTSMATPHVAGAAALILSKHPTWTNAQVRDRLEST 359
Score = 33.5 bits (75), Expect = 1.7
Identities = 46/184 (25%), Positives = 73/184 (39%), Gaps = 36/184 (19%)
Query: 209 LPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT-IKGGSPRSRVATYKACWSLT 267
+PS +D GTH T I + N + G SP + + K
Sbjct: 157 VPSETNPYQDGSSHGTHVAGT----------IAALNNSIGVLGVSPSASLYAVK------ 200
Query: 268 DVVDCFGAD----VLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARN 323
V+D G+ ++ I+ AI + D+I++S GG + D+ A++
Sbjct: 201 -VLDSTGSGQYSWIINGIEWAISNNMDVINMSLGGPSGSTALKTVVDK-------AVSSG 252
Query: 324 ILLVASAGNEGPTPGSVTNVAP-------WVFTVAASTLDRDFSSVMTINNKTLTGASLF 376
I++ A+AGNEG + S T P V V +S FSS + + G S+
Sbjct: 253 IVVAAAAGNEGSSGSSSTVGYPAKYPSTIAVGAVNSSNQRASFSSAGSELDVMAPGVSIQ 312
Query: 377 VNLP 380
LP
Sbjct: 313 STLP 316
>SUBT_BACSA (P00783) Subtilisin amylosacchariticus precursor (EC
3.4.21.62)
Length = 381
Score = 48.5 bits (114), Expect = 5e-05
Identities = 36/112 (32%), Positives = 49/112 (43%), Gaps = 28/112 (25%)
Query: 491 KTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASV 550
K +TI + N+ N R ASFSS G DV APGV+I +
Sbjct: 276 KYPSTIAVGAVNSSNQR------ASFSSAGSEL--------DVMAPGVSIQSTLP----- 316
Query: 551 SNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
G + GTSM+ PHV G A LI + HP W+ A ++ + +T
Sbjct: 317 ---------GGTYGAYNGTSMATPHVAGAAALILSKHPTWTNAQVRDRLEST 359
Score = 33.5 bits (75), Expect = 1.7
Identities = 46/184 (25%), Positives = 73/184 (39%), Gaps = 36/184 (19%)
Query: 209 LPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT-IKGGSPRSRVATYKACWSLT 267
+PS +D GTH T I + N + G SP + + K
Sbjct: 157 VPSETNPYQDGSSHGTHVAGT----------IAALNNSIGVLGVSPSASLYAVK------ 200
Query: 268 DVVDCFGAD----VLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARN 323
V+D G+ ++ I+ AI + D+I++S GG + D+ A++
Sbjct: 201 -VLDSTGSGQYSWIINGIEWAISNNMDVINMSLGGPSGSTALKTVVDK-------AVSSG 252
Query: 324 ILLVASAGNEGPTPGSVTNVAP-------WVFTVAASTLDRDFSSVMTINNKTLTGASLF 376
I++ A+AGNEG + S T P V V +S FSS + + G S+
Sbjct: 253 IVVAAAAGNEGSSGSSSTVGYPAKYPSTIAVGAVNSSNQRASFSSAGSELDVMAPGVSIQ 312
Query: 377 VNLP 380
LP
Sbjct: 313 STLP 316
>SCA2_STRPY (P58099) C5a peptidase precursor (EC 3.4.21.-) (SCP)
Length = 1181
Score = 48.5 bits (114), Expect = 5e-05
Identities = 80/327 (24%), Positives = 136/327 (41%), Gaps = 45/327 (13%)
Query: 280 AIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNE----GP 335
AI A+ GA +I++S G + DE +A ++ + +V SAGN+ G
Sbjct: 245 AIIDAVNLGAKVINMSFGNAALAYANL--PDETKKAFDYAKSKGVSIVTSAGNDSSFGGK 302
Query: 336 TPGSVTNVAPWVFTVAASTLDRDFSSVMTINNKTLTGASLFVNLPPNQDFLIIISTDAKF 395
T + + + + D + +K LT + + ++ST+ +F
Sbjct: 303 TRLPLADHPDYGVVGTPAAADSTLTVASYSPDKQLTETATVKTADQQDKEMPVLSTN-RF 361
Query: 396 ANVTDVDAQFCRPGTL--DPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPE 453
D + G D V GK+ +R G I+ + A AGAVGV++ + +
Sbjct: 362 EPNKAYDYAYANRGMKEDDFKDVKGKIALIER-GDIDFKDKIANAKKAGAVGVLIYDNQD 420
Query: 454 VDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDIKT---NATIRMSPANALNGRKPA 510
P + ++ A I+ G + KT NAT ++ P +G K
Sbjct: 421 K------GFPIELPNVDQMPAAFISRKDGLLLKENPQKTITFNATPKVLPT--ASGTK-- 470
Query: 511 PVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTS 570
++ FSS G +KPD+ APG +IL++ V +N+ + GTS
Sbjct: 471 --LSRFSSWG--LTADGNIKPDIAAPGQDILSS----------VANNK----YAKLSGTS 512
Query: 571 MSCPHVVGTAGLI----KTLHPNWSPA 593
MS P V G GL+ +T +P+ +P+
Sbjct: 513 MSAPLVAGIMGLLQKQYETQYPDMTPS 539
>SUBN_BACNA (P35835) Subtilisin NAT precursor (EC 3.4.21.62)
Length = 381
Score = 47.8 bits (112), Expect = 9e-05
Identities = 36/112 (32%), Positives = 49/112 (43%), Gaps = 28/112 (25%)
Query: 491 KTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASV 550
K +TI + N+ N R ASFSS G DV APGV+I +
Sbjct: 276 KYPSTIAVGAVNSSNQR------ASFSSVGSEL--------DVMAPGVSIQSTLP----- 316
Query: 551 SNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
G + GTSM+ PHV G A LI + HP W+ A ++ + +T
Sbjct: 317 ---------GGTYGAYNGTSMATPHVAGAAALILSKHPTWTNAQVRDRLEST 359
Score = 34.3 bits (77), Expect = 0.98
Identities = 46/185 (24%), Positives = 78/185 (41%), Gaps = 38/185 (20%)
Query: 209 LPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT-IKGGSPRSRVATYKACWSLT 267
+PS +D GTH T I + N + G +P + + K
Sbjct: 157 VPSETNPYQDGSSHGTHVAGT----------IAALNNSIGVLGVAPSASLYAVK------ 200
Query: 268 DVVDCFGAD----VLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARN 323
V+D G+ ++ I+ AI + D+I++S GG + D+ A++
Sbjct: 201 -VLDSTGSGQYSWIINGIEWAISNNMDVINMSLGGPTGSTALKTVVDK-------AVSSG 252
Query: 324 ILLVASAGNEGPTPGSVTNV---APWVFTVAASTLD-----RDFSSVMTINNKTLTGASL 375
I++ A+AGNEG + GS + V A + T+A ++ FSSV + + G S+
Sbjct: 253 IVVAAAAGNEG-SSGSTSTVGYPAKYPSTIAVGAVNSSNQRASFSSVGSELDVMAPGVSI 311
Query: 376 FVNLP 380
LP
Sbjct: 312 QSTLP 316
>SEPR_THESR (P80146) Extracellular serine proteinase precursor (EC
3.4.21.-)
Length = 408
Score = 44.7 bits (104), Expect = 7e-04
Identities = 41/182 (22%), Positives = 77/182 (41%), Gaps = 24/182 (13%)
Query: 419 KVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSIT 478
+V+ C+ G +S+ G + ++ V + N G + + V++ IN +T
Sbjct: 227 RVLDCNGSGSNSSVIAGLDWVTQNHVKPAVINMSLGGGASTALDTAVMNAIN----AGVT 282
Query: 479 TPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGV 538
+ D + R++ A + ASFS+ G D+ APG
Sbjct: 283 VVVAAGNDNRDACFYSPARVTAAITVGATTSTDYRASFSNYGRCL--------DLFAPGQ 334
Query: 539 NILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSA 598
+I +A+ ++ +N ++ GTSM+ PHV G A L +P +P+ + SA
Sbjct: 335 SITSAWYTSSTATNTIS------------GTSMATPHVTGAAALYLQWYPTATPSQVASA 382
Query: 599 IM 600
++
Sbjct: 383 LL 384
>ELYA_BACHD (P41363) Thermostable alkaline protease precursor (EC
3.4.21.-)
Length = 361
Score = 44.7 bits (104), Expect = 7e-04
Identities = 30/89 (33%), Positives = 46/89 (50%), Gaps = 22/89 (24%)
Query: 514 ASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSC 573
ASFS+ GP + +++APGVN+ + Y T NR + GTSM+
Sbjct: 273 ASFSTYGP--------EIEISAPGVNVNSTY----------TGNR----YVSLSGTSMAT 310
Query: 574 PHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
PHV G A L+K+ +P+++ I+ I T
Sbjct: 311 PHVAGVAALVKSRYPSYTNNQIRQRINQT 339
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,188,688
Number of Sequences: 164201
Number of extensions: 2992090
Number of successful extensions: 6599
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 32
Number of HSP's that attempted gapping in prelim test: 6510
Number of HSP's gapped (non-prelim): 87
length of query: 611
length of database: 59,974,054
effective HSP length: 116
effective length of query: 495
effective length of database: 40,926,738
effective search space: 20258735310
effective search space used: 20258735310
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 69 (31.2 bits)
Medicago: description of AC133779.1


