

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133571.7 + phase: 0
(79 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
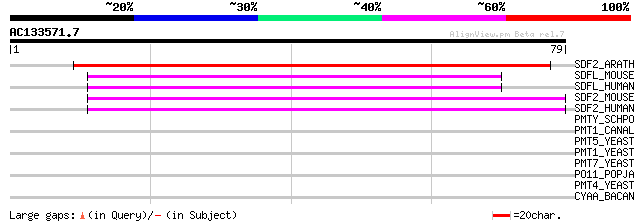
Score E
Sequences producing significant alignments: (bits) Value
SDF2_ARATH (Q93ZE8) Stromal cell-derived factor 2-like protein p... 129 2e-30
SDFL_MOUSE (Q9ESP1) Stromal cell-derived factor 2-like protein 1... 46 2e-05
SDFL_HUMAN (Q9HCN8) Stromal cell-derived factor 2-like protein 1... 44 7e-05
SDF2_MOUSE (Q9DCT5) Stromal cell-derived factor 2 precursor (SDF-2) 44 7e-05
SDF2_HUMAN (Q99470) Stromal cell-derived factor 2 precursor (SDF-2) 44 7e-05
PMTY_SCHPO (O42933) Probable dolichyl-phosphate-mannose--protein... 32 0.29
PMT1_CANAL (O74189) Dolichyl-phosphate-mannose--protein mannosyl... 32 0.29
PMT5_YEAST (P52867) Dolichyl-phosphate-mannose--protein mannosyl... 31 0.65
PMT1_YEAST (P33775) Dolichyl-phosphate-mannose--protein mannosyl... 30 1.1
PMT7_YEAST (Q06644) Dolichyl-phosphate-mannose--protein mannosyl... 29 2.5
PO11_POPJA (Q03276) Retrovirus-related Pol polyprotein from type... 27 7.2
PMT4_YEAST (P46971) Dolichyl-phosphate-mannose--protein mannosyl... 27 9.4
CYAA_BACAN (P40136) Calmodulin-sensitive adenylate cyclase precu... 27 9.4
>SDF2_ARATH (Q93ZE8) Stromal cell-derived factor 2-like protein
precursor (SDF2-like protein)
Length = 218
Score = 129 bits (323), Expect = 2e-30
Identities = 57/68 (83%), Positives = 63/68 (91%)
Query: 10 EGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 69
EGSGKTWKQDQR RLQHIDTSGY +SHDKKY RIAGGQQEVCG+REK+ADN+WLAAEGVY
Sbjct: 151 EGSGKTWKQDQRVRLQHIDTSGYLHSHDKKYQRIAGGQQEVCGIREKKADNIWLAAEGVY 210
Query: 70 LPVTESKQ 77
LP+ ES +
Sbjct: 211 LPLNESSK 218
>SDFL_MOUSE (Q9ESP1) Stromal cell-derived factor 2-like protein 1
precursor (SDF2 like protein 1)
Length = 221
Score = 45.8 bits (107), Expect = 2e-05
Identities = 20/59 (33%), Positives = 32/59 (53%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
SG+ W+++ R QH+ TS + ++Y GQ EV G+ A N W A EG+++
Sbjct: 150 SGQHWEREASVRFQHVGTSVFLSVTGEQYGNPIRGQHEVHGMPSANAHNTWKAMEGIFI 208
>SDFL_HUMAN (Q9HCN8) Stromal cell-derived factor 2-like protein 1
precursor (SDF2 like protein 1) (PWP1-interacting
protein 8) (UNQ1941/PRO4424)
Length = 221
Score = 43.9 bits (102), Expect = 7e-05
Identities = 19/59 (32%), Positives = 31/59 (52%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYL 70
SG+ W+++ R QH+ TS + ++Y GQ EV G+ N W A EG+++
Sbjct: 150 SGQHWEREAAVRFQHVGTSVFLSVTGEQYGSPIRGQHEVHGMPSANTHNTWKAMEGIFI 208
>SDF2_MOUSE (Q9DCT5) Stromal cell-derived factor 2 precursor (SDF-2)
Length = 211
Score = 43.9 bits (102), Expect = 7e-05
Identities = 21/68 (30%), Positives = 35/68 (50%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYLP 71
+G W +D R +H T ++Y R GQ+EV G+ + +N W A EG+++
Sbjct: 138 NGPYWVRDGEVRFKHSSTDVLLSVTGEQYGRPISGQKEVHGMAQPSQNNYWKAMEGIFMK 197
Query: 72 VTESKQAE 79
+E +AE
Sbjct: 198 PSELLRAE 205
>SDF2_HUMAN (Q99470) Stromal cell-derived factor 2 precursor (SDF-2)
Length = 211
Score = 43.9 bits (102), Expect = 7e-05
Identities = 21/68 (30%), Positives = 35/68 (50%)
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYLP 71
+G W +D R +H T ++Y R GQ+EV G+ + +N W A EG+++
Sbjct: 138 NGPYWVRDGEVRFKHSSTEVLLSVTGEQYGRPISGQKEVHGMAQPSQNNYWKAMEGIFMK 197
Query: 72 VTESKQAE 79
+E +AE
Sbjct: 198 PSELLKAE 205
>PMTY_SCHPO (O42933) Probable dolichyl-phosphate-mannose--protein
mannosyltransferase C16C6.09 (EC 2.4.1.109)
Length = 778
Score = 32.0 bits (71), Expect = 0.29
Identities = 14/46 (30%), Positives = 26/46 (56%), Gaps = 6/46 (13%)
Query: 24 LQHIDTSGYFYSHDKKY------SRIAGGQQEVCGVREKRADNVWL 63
++H+ T+ + +SH +KY RI+ G Q+V G + +N W+
Sbjct: 347 IKHMGTNAFLHSHPEKYPIPYDDGRISSGGQQVTGYQFDDENNYWM 392
>PMT1_CANAL (O74189) Dolichyl-phosphate-mannose--protein
mannosyltransferase 1 (EC 2.4.1.109)
Length = 877
Score = 32.0 bits (71), Expect = 0.29
Identities = 22/58 (37%), Positives = 28/58 (47%), Gaps = 3/58 (5%)
Query: 22 FRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVRE-KRADNVWL--AAEGVYLPVTESK 76
FRL+H T Y +S + K GQQEV + KRA W E LP +E+K
Sbjct: 482 FRLRHAMTGHYLFSSEVKLPEWGFGQQEVTSASQGKRALTHWYIETNENSILPPSEAK 539
>PMT5_YEAST (P52867) Dolichyl-phosphate-mannose--protein
mannosyltransferase 5 (EC 2.4.1.109)
Length = 743
Score = 30.8 bits (68), Expect = 0.65
Identities = 14/32 (43%), Positives = 19/32 (58%)
Query: 19 DQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEV 50
+ +FRL+H T Y +SH +K GQQEV
Sbjct: 461 ETKFRLKHYLTGCYLFSHPEKLPEWGFGQQEV 492
>PMT1_YEAST (P33775) Dolichyl-phosphate-mannose--protein
mannosyltransferase 1 (EC 2.4.1.109)
Length = 817
Score = 30.0 bits (66), Expect = 1.1
Identities = 16/41 (39%), Positives = 20/41 (48%)
Query: 19 DQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRAD 59
D +FRL+H T Y +SH+ K QQEV R D
Sbjct: 465 DTKFRLRHAMTGCYLFSHEVKLPAWGFEQQEVTCASSGRHD 505
>PMT7_YEAST (Q06644) Dolichyl-phosphate-mannose--protein
mannosyltransferase 7 (EC 2.4.1.109)
Length = 662
Score = 28.9 bits (63), Expect = 2.5
Identities = 16/41 (39%), Positives = 19/41 (46%), Gaps = 1/41 (2%)
Query: 24 LQHIDTS-GYFYSHDKKYSRIAGGQQEVCGVREKRADNVWL 63
L+H+D+ GY SHD Y Q E ADN WL
Sbjct: 300 LRHLDSMVGYLASHDISYPSDVDEQLVALSFEEFAADNEWL 340
>PO11_POPJA (Q03276) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 245
Score = 27.3 bits (59), Expect = 7.2
Identities = 12/39 (30%), Positives = 21/39 (53%)
Query: 24 LQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVW 62
+Q + GYF ++ +++ G CG E+ AD+VW
Sbjct: 150 VQLVTGHGYFKAYYRRFKLREGSGLCECGEAEETADHVW 188
>PMT4_YEAST (P46971) Dolichyl-phosphate-mannose--protein
mannosyltransferase 4 (EC 2.4.1.109)
Length = 762
Score = 26.9 bits (58), Expect = 9.4
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 2/51 (3%)
Query: 22 FRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRAD--NVWLAAEGVYL 70
FRL H+DTS ++H+ + G QQ+ +K D N W+ E V L
Sbjct: 474 FRLFHVDTSVALWTHNDELLPDWGFQQQEINGNKKVIDPSNNWVVDEIVNL 524
>CYAA_BACAN (P40136) Calmodulin-sensitive adenylate cyclase
precursor (EC 4.6.1.1) (ATP pyrophosphate-lyase)
(Adenylyl cyclase) (Edema factor) (EF) (Anthrax edema
toxin adenylate cyclase component)
Length = 800
Score = 26.9 bits (58), Expect = 9.4
Identities = 11/35 (31%), Positives = 19/35 (53%)
Query: 14 KTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQ 48
K W+ RF ++I Y Y ++ Y++IA G +
Sbjct: 606 KNWEMTGRFIEKNITGKDYLYYFNRSYNKIAPGNK 640
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.131 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,759,970
Number of Sequences: 164201
Number of extensions: 375215
Number of successful extensions: 534
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 520
Number of HSP's gapped (non-prelim): 17
length of query: 79
length of database: 59,974,054
effective HSP length: 55
effective length of query: 24
effective length of database: 50,942,999
effective search space: 1222631976
effective search space used: 1222631976
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC133571.7


